Abstract
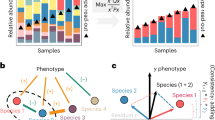
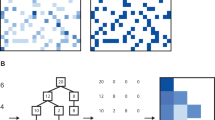
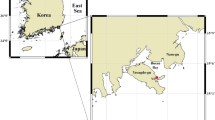
Previous studies suggested that microbial communities can harbour keystone species whose removal can cause a dramatic shift in microbiome structure and functioning. Yet, an efficient method to systematically identify keystone species in microbial communities is still lacking. Here we propose a data-driven keystone species identification (DKI) framework based on deep learning to resolve this challenge. Our key idea is to implicitly learn the assembly rules of microbial communities from a particular habitat by training a deep-learning model using microbiome samples collected from this habitat. The well-trained deep-learning model enables us to quantify the community-specific keystoneness of each species in any microbiome sample from this habitat by conducting a thought experiment on species removal. We systematically validated this DKI framework using synthetic data and applied DKI to analyse real data. We found that those taxa with high median keystoneness across different communities display strong community specificity. The presented DKI framework demonstrates the power of machine learning in tackling a fundamental problem in community ecology, paving the way for the data-driven management of complex microbial communities.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Gut microbiome data were collected from the curatedMetagenomicData32 database. Oral microbiome data are available at the CNGB Sequence Archive (CNSA) of the China National GeneBank DataBase (CNGBdb) (CNSA CNP0000687 for the 4D-SZ cohort and CNP0001221 for the Yunnan cohort). Coral and soil microbiome data were collected from Qiita40 (IDs 10895 and 2104). Data supporting our findings are provided at https://github.com/spxuw/DKI.
Code availability
The code used in this work is available at https://github.com/spxuw/DKI.
References
Paine, R. T. A note on trophic complexity and community stability. Am. Nat. 103, 91–93 (1969).
Paine, R. T. Food web complexity and species diversity. Am. Nat. 100, 65–75 (1966).
Cottee-Jones, H. E. W. & Whittaker, R. J. The keystone species concept: a critical appraisal. Front Biogeogr. 4, 117–127 (2012).
Power, M. E. et al. Challenges in the quest for keystones: identifying keystone species is difficult—but essential to understanding how loss of species will affect ecosystems. BioScience 46, 609–620 (1996).
Ze, X., Duncan, S. H., Louis, P. & Flint, H. J. Ruminococcus bromii is a keystone species for the degradation of resistant starch in the human colon. ISME J. 6, 1535–1543 (2012).
Banerjee, S. et al. Network analysis reveals functional redundancy and keystone taxa amongst bacterial and fungal communities during organic matter decomposition in an arable soil. Soil Biol. Biochem. 97, 188–198 (2016).
Trosvik, P. & de Muinck, E. J. Ecology of bacteria in the human gastrointestinal tract—identification of keystone and foundation taxa. Microbiome 3, 44 (2015).
Xun, W. et al. Specialized metabolic functions of keystone taxa sustain soil microbiome stability. Microbiome 9, 35 (2021).
LeBlanc, N. & Crouch, J. A. Prokaryotic taxa play keystone roles in the soil microbiome associated with woody perennial plants in the genus Buxus. Ecol. Evol. 9, 11102–11111 (2019).
The Human Microbiome Project Consortium Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Franzosa, E. A. et al. Identifying personal microbiomes using metagenomic codes. Proc. Natl Acad. Sci. USA 112, E2930–E2938 (2015).
Vujkovic-Cvijin, I. et al. Host variables confound gut microbiota studies of human disease. Nature 587, 448–454 (2020).
Stein, R. R. et al. Ecological modeling from time-series inference: insight into dynamics and stability of intestinal microbiota. PLoS Comput. Biol. 9, e1003388 (2013).
Fisher, C. K. & Mehta, P. Identifying keystone species in the human gut microbiome from metagenomic timeseries using sparse linear regression. PLoS ONE 9, e102451 (2014).
Bucci, V. et al. MDSINE: Microbial Dynamical Systems INference Engine for microbiome time-series analyses. Genome Biol. 17, 121 (2016).
Berry, D. & Widder, S. Deciphering microbial interactions and detecting keystone species with co-occurrence networks. Front. Microbiol. 5, 219 (2014).
Banerjee, S., Schlaeppi, K. & van der Heijden, M. G. A. Keystone taxa as drivers of microbiome structure and functioning. Nat. Rev. Microbiol. 16, 567–576 (2018).
Garrett, W. S. et al. Enterobacteriaceae act in concert with the gut microbiota to induce spontaneous and maternally transmitted colitis. Cell Host Microbe 8, 292–300 (2010).
Hajishengallis, G. et al. Low-abundance biofilm species orchestrates inflammatory periodontal disease through the commensal microbiota and complement. Cell Host Microbe 10, 497–506 (2011).
Agler, M. T. et al. Microbial hub taxa link host and abiotic factors to plant microbiome variation. PLoS Biol. 14, e1002352 (2016).
Banerjee, S., Schlaeppi, K. & van der Heijden, M. G. Reply to ‘Can we predict microbial keystones?’. Nat. Rev. Microbiol. 17, 194–194 (2019).
Röttjers, L. & Faust, K. Can we predict keystones? Nat. Rev. Microbiol. 17, 193 (2019).
Amit, G. & Bashan, A. Top-down identification of keystone taxa in the microbiome. Nat. Commun. 14, 3951 (2023).
Valls, A., Coll, M. & Christensen, V. Keystone species: toward an operational concept for marine biodiversity conservation. Ecol. Monogr. 85, 29–47 (2015).
Tian, L. et al. Deciphering functional redundancy in the human microbiome. Nat. Commun. 11, 6217 (2020).
Gouveia, C., Móréh, Á. & Jordán, F. Combining centrality indices: maximizing the predictability of keystone species in food webs. Ecol. Indic. 126, 107617 (2021).
Kruse, R., Borgelt, C., Braune, C., Mostaghim, S. & Steinbrecher, M. Computational Intelligence: A Methodological Introduction (Springer, 2022).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In Proc. 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 770–778 (IEEE, 2016).
Michel-Mata, S., Wang, X.-W., Liu, Y.-Y. & Angulo, M. T. Predicting microbiome compositions from species assemblages through deep learning. Imeta 1, e3 (2022).
Almeida-Neto, M., Guimaraes, P., Guimaraes, P. R. Jr, Loyola, R. D. & Ulrich, W. A consistent metric for nestedness analysis in ecological systems: reconciling concept and measurement. Oikos 117, 1227–1239 (2008).
Friedman, J., Higgins, L. M. & Gore, J. Community structure follows simple assembly rules in microbial microcosms. Nat. Ecol. Evol. 1, 0109 (2017).
Pasolli, E. et al. Accessible, curated metagenomic data through ExperimentHub. Nat. Methods 14, 1023–1024 (2017).
Tudela, H., Claus, S. P. & Saleh, M. Next generation microbiome research: identification of keystone species in the metabolic regulation of host–gut microbiota interplay. Front. Cell Dev. Biol. 9, 719072 (2021).
Alessandri, G., van Sinderen, D. & Ventura, M. The genus Bifidobacterium: from genomics to functionality of an important component of the mammalian gut microbiota. Comput. Struct. Biotechnol. J. 19, 1472–1487 (2021).
Zhang, Z. et al. Spatial heterogeneity and co-occurrence patterns of human mucosal-associated intestinal microbiota. ISME J. 8, 881–893 (2014).
Leylabadlo, H. E. et al. The critical role of Faecalibacterium prausnitzii in human health: an overview. Microb. Pathog. 149, 104344 (2020).
Gotoh, A., Ojima, M. N. & Katayama, T. Minority species influences microbiota formation: the role of Bifidobacterium with extracellular glycosidases in bifidus flora formation in breastfed infant guts. Microb. Biotechnol. 12, 259–264 (2019).
Bui, T. P. N. et al. Intestinimonas-like bacteria are important butyrate producers that utilize Nε-fructosyllysine and lysine in formula-fed infants and adults. J. Funct. Foods 70, 103974 (2020).
Li, L. et al. Revealing proteome-level functional redundancy in the human gut microbiome using ultra-deep metaproteomics. Nat. Commun. 14, 3428 (2023).
Gonzalez, A. et al. Qiita: rapid, web-enabled microbiome meta-analysis. Nat. Methods 15, 796–798 (2018).
Friedman, J. & Alm, E. J. Inferring correlation networks from genomic survey data. PLoS Comput. Biol. 8, e1002687 (2012).
Acknowledgements
Y.-Y.L. acknowledges funding support from the National Institutes of Health (R01AI141529, R01HD093761, RF1AG067744, UH3OD023268, U19AI095219 and U01HL089856) as well as the Office of the Assistant Secretary of Defense for Health Affairs, through the Traumatic Brain Injury and Psychological Health Research Program (Focused Program Award) under award no. W81XWH-22-S-TBIPH2, endorsed by the Department of Defense. X.-W.W. acknowledges funding support from the National Institutes of Health (K25HL166208). Z.S. acknowledges funding support from the National Institutes of Health (K99HL163519). M.T.A. acknowledges the financial support provided by CONACyT grant No. A1-S-13909 and PAPIIT 104915.
Author information
Authors and Affiliations
Contributions
Y.-Y.L. conceived and designed the project. X.-W.W. performed all the numerical calculations. X.-W.W. and Z.S. analysed real data. X.-W.W. and Y.-Y.L. wrote the manuscript. H.J., S.M.-M., M.T.A., L.D., X.H. and S.T.W. edited the manuscript. All authors approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Ecology & Evolution thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Discussions 1–8 and Figs. 1–14.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, XW., Sun, Z., Jia, H. et al. Identifying keystone species in microbial communities using deep learning. Nat Ecol Evol 8, 22–31 (2024). https://doi.org/10.1038/s41559-023-02250-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-023-02250-2
This article is cited by
-
Data-driven prediction of colonization outcomes for complex microbial communities
Nature Communications (2024)