Abstract
Developmental time is a key life-history trait with large effects on Darwinian fitness. In many insects, developmental time is currently under strong selection to minimize ecological mismatches in seasonal timing induced by climate change. The genetic basis of responses to such selection, however, is poorly understood. To address this problem, we set up a long-term evolve-and-resequence experiment in the beetle Tribolium castaneum and selected replicate, outbred populations for fast or slow embryonic development. The response to this selection was substantial and embryonic developmental timing of the selection lines started to diverge during dorsal closure. Pooled whole-genome resequencing, gene expression analysis and an RNAi screen pinpoint a 222 bp deletion containing binding sites for Broad and Tramtrack upstream of the ecdysone degrading enzyme Cyp18a1 as a main target of selection. Using CRISPR/Cas9 to reconstruct this allele in the homogenous genetic background of a laboratory strain, we unravel how this single deletion advances the embryonic ecdysone peak inducing dorsal closure and show that this allele accelerates larval development but causes a trade-off with fecundity. Our study uncovers a life-history allele of large effect and reveals the evolvability of developmental time in a natural insect population.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Sequencing reads have been deposited in NCBIs Sequencing Read Archive as BioProject PRJNA942224. All other data are provided as supplementary data files. Source data are provided with this paper.
Code availability
All custom code is available in Zenodo at https://doi.org/10.5281/zenodo.8395048 ref. 89. All other code used is referenced.
References
Dobreva, M. P., Camacho, J. & Abzhanov, A. Time to synchronize our clocks: connecting developmental mechanisms and evolutionary consequences of heterochrony. J. Exp. Zool. B 338, 87–106 (2022).
Gould, S. J. Ontogeny and Phylogeny (Harvard Univ. Press, 1977).
Roff, D. A. The Evolution of Life Histories: Theory and Analysis (Chapman & Hall, 1992).
Stearns, S. C. The Evolution of Life Histories (Oxford Univ. Press, 1992).
Stearns, S. C. Life history evolution: successes, limitations, and prospects. Naturwissenschaften 87, 476–486 (2000).
Stearns, S. C., Kaiser, M. & Kawecki, T. J. The differential genetic and environmental canalization of fitness components in Drosophila melanogaster. J. Evol. Biol. 8, 539–557 (1995).
Stearns, S. C. & Kawecki, T. J. Fitness sensitivity and the canalization of life-history traits. Evolution 48, 1438–1450 (1994).
Kharouba, H. M. et al. Global shifts in the phenological synchrony of species interactions over recent decades. Proc Natl Acad. Sci. USA 115, 5211–5216 (2018).
Samplonius, J. M. et al. Strengthening the evidence base for temperature-mediated phenological asynchrony and its impacts. Nat. Ecol. Evol. 5, 155–164 (2021).
Thackeray, S. J. et al. Phenological sensitivity to climate across taxa and trophic levels. Nature 535, 241–245 (2016).
Singer, M. C. & Parmesan, C. Phenological asynchrony between herbivorous insects and their hosts: signal of climate change or pre-existing adaptive strategy? Phil. Trans. R. Soc. B 365, 3161–3176 (2010).
Visser, M. E. & Gienapp, P. Evolutionary and demographic consequences of phenological mismatches. Nat. Ecol. Evol. 3, 879–885 (2019).
van Asch, M., Salis, L., Holleman, L. J. M., van Lith, B. & Visser, M. E. Evolutionary response of the egg hatching date of a herbivorous insect under climate change. Nat. Clim. Change 3, 244–248 (2013).
Franks, S. J. & Hoffmann, A. A. Genetics of climate change adaptation. Annu. Rev. Genet. 46, 185–208 (2012).
Renner, S. S. & Zohner, C. M. Climate change and phenological mismatch in trophic interactions among plants, insects, and vertebrates. Annu. Rev. Ecol. Evol. Syst. 49, 165–182 (2018).
Mensch, J. et al. Identifying candidate genes affecting developmental time in Drosophila melanogaster: pervasive pleiotropy and gene-by-environment interaction. BMC Dev. Biol. 8, 78 (2008).
Flatt, T. Life-history evolution and the genetics of fitness components in Drosophila melanogaster. Genetics 214, 3–48 (2020).
Horvath, B., Betancourt, A. J. & Kalinka, A. T. A novel method for quantifying the rate of embryogenesis uncovers considerable genetic variation for the duration of embryonic development in Drosophila melanogaster. BMC Evol. Biol. 16, 200 (2016).
Chippindale, A. K., Alipaz, J. A., Chen, H. W. & Rose, M. R. Experimental evolution of accelerated development in Drosophila.1. Developmental speed and larval survival. Evolution 51, 1536–1551 (1997).
Nunney, L. The response to selection for fast larval development in Drosophila melanogaster and its effect on adult weight: an example of a fitness trade-off. Evolution 50, 1193–1204 (1996).
Prasad, N. G. et al. Evolution of reduced pre-adult viability and larval growth rate in laboratory populations of Drosophila melanogaster selected for shorter development time. Genet. Res. 76, 249–259 (2000).
Sharma, K., Mishra, N. & Shakarad, M. N. Evolution of reduced minimum critical size as a response to selection for rapid pre-adult development in Drosophila melanogaster. R. Soc. Open Sci. 7, 191910 (2020).
Zwaan, B., Bijlsma, R. & Hoekstra, R. F. Artificial selection for developmental time in Drosophila melanogaster in relation to the evolution of aging: direct and correlated responses. Evolution 49, 635–648 (1995).
Fischer, K., Zwaan, B. J. & Brakefield, P. M. Realized correlated responses to artificial selection on pre-adult life-history traits in a butterfly. Heredity 98, 157–164 (2007).
Seslija, D. & Tucic, N. Selection for developmental time in bean weevil (Acanthoscelides obtectus): correlated responses for other life history traits and genetic architecture of line differentiation. Entomol. Exp. Appl. 106, 19–35 (2003).
Soliman, M. H. Directional and stabilizing selection for developmental time and correlated response in reproductive fitness in Tribolium castaneum. Theor. Appl. Genet. 63, 111–116 (1982).
Nascimento, J. C. D., Cruz, I. B. M. D., Monjeló, L. A. & Oliveira, A. K. D. Genetic components affecting embryonic developmental time of Drosophila melanogaster. Genet. Mol. Biol. 25, 157–160 (2002).
Neyfakh, A. A. & Hartl, D. L. Genetic control of the rate of embryonic development: selection for faster development at elevated temperatures. Evolution 47, 1625–1631 (1993).
Marinkovic, D. & Ayala, F. J. Selection for different rates of embryonic development in Drosophila melanogaster and Drosophila simulans. Genetika 18, 205–219 (1986).
Long, A., Liti, G., Luptak, A. & Tenaillon, O. Elucidating the molecular architecture of adaptation via evolve and resequence experiments. Nat. Rev. Genet. 16, 567–582 (2015).
Klingler, M. & Bucher, G. The red flour beetle T. castaneum: elaborate genetic toolkit and unbiased large scale RNAi screening to study insect biology and evolution. Evodevo 13, 14 (2022).
Richards, S. et al. The genome of the model beetle and pest Tribolium castaneum. Nature 452, 949–955 (2008).
Flatt, T. & Heyland, A. Mechanisms of Life History Evolution: The Genetics and Physiology of Life History Traits and Trade-offs (Oxford Univ. Press, 2011).
Roff, D. A. Trade-offs between growth and reproduction: an analysis of the quantitative genetic evidence. J. Evol. Biol. 13, 434–445 (2000).
Lee, Y. et al. Inverse correlation between longevity and developmental rate among wild C. elegans strains. Aging 8, 986–999 (2016).
Panfilio, K. A. Extraembryonic development in insects and the acrobatics of blastokinesis. Dev. Biol. 313, 471–491 (2008).
Schlotterer, C., Tobler, R., Kofler, R. & Nolte, V. Sequencing pools of individuals – mining genome-wide polymorphism data without big funding. Nat. Rev. Genet. 15, 749–763 (2014).
Hoedjes, K. M. et al. Distinct genomic signals of lifespan and life history evolution in response to postponed reproduction and larval diet in Drosophila. Evol. Lett. 3, 598–609 (2019).
Guittard, E. et al. CYP18A1, a key enzyme of Drosophila steroid hormone inactivation, is essential for metamorphosis. Dev. Biol. 349, 35–45 (2011).
Rewitz, K. F., Yamanaka, N. & O’Connor, M. B. Steroid hormone inactivation is required during the juvenile–adult transition in Drosophila. Dev. Cell 19, 895–902 (2010).
Nijhout, H. F. et al. The developmental control of size in insects. Wiley Interdiscip. Rev. Dev. Biol. 3, 113–134 (2014).
Niwa, Y. S. & Niwa, R. Transcriptional regulation of insect steroid hormone biosynthesis and its role in controlling timing of molting and metamorphosis. Dev. Growth Differ. 58, 94–105 (2016).
Kozlova, T. & Thummel, C. S. Essential roles for ecdysone signaling during Drosophila mid-embryonic development. Science 301, 1911–1914 (2003).
Maroy, P., Kaufmann, G. & Dubendorfer, A. Embryonic ecdysteroids of Drosophila melanogaster. J. Insect Physiol. 34, 633–637 (1988).
Yoo, B. et al. 20-hydroxyecdysone (20E) signaling regulates amnioserosa morphogenesis during Drosophila dorsal closure: EcR modulates gene expression in a complex with the AP-1 subunit, Jun. Biol. Open 10, bio058605 (2021).
Kidokoro, K., Wata, K., Fujiwara, Y. & Takeda, M. Effects of juvenile hormone analogs and 20-hydroxyecdysone on diapause termination in eggs of Locusta migratoria and Oxya yezoensis. J. Insect Physiol. 52, 473–479 (2006).
Sarrazin, A. F., Peel, A. D. & Averof, M. A segmentation clock with two-segment periodicity in insects. Science 336, 338–341 (2012).
Grant, C. E., Bailey, T. L. & Noble, W. S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011).
Karim, F. D., Guild, G. M. & Thummel, C. S. The Drosophila Broad-Complex plays a key role in controlling ecdysone-regulated gene expression at the onset of metamorphosis. Development 118, 977–988 (1993).
Pagans, S., Ortiz-Lombardia, M., Espinas, M. L., Bernues, J. & Azorin, F. The Drosophila transcription factor tramtrack (TTK) interacts with Trithorax-like (GAGA) and represses GAGA-mediated activation. Nucleic Acids Res. 30, 4406–4413 (2002).
Yu, Y., Yussa, M., Song, J. B., Hirsch, J. & Pick, L. A double interaction screen identifies positive and negative ftz gene regulators and Ftz-interacting proteins. Mech. Dev. 83, 95–105 (1999).
Sun, J., Smith, L., Armento, A. & Deng, W. M. Regulation of the endocycle/gene amplification switch by Notch and ecdysone signaling. J. Cell Biol. 182, 885–896 (2008).
Zuin, J. et al. Nonlinear control of transcription through enhancer-promoter interactions. Nature 604, 571–577 (2022).
Teleman, A. A., Chen, Y. W. & Cohen, S. M. Drosophila melted modulates FOXO and TOR activity. Dev. Cell 9, 271–281 (2005).
Baker, K. D. & Thummel, C. S. Diabetic larvae and obese flies–emerging studies of metabolism in Drosophila. Cell Metab. 6, 257–266 (2007).
Noma, K. et al. RITS acts in cis to promote RNA interference-mediated transcriptional and post-transcriptional silencing. Nat. Genet. 36, 1174–1180 (2004).
Franssen, S. U., Nolte, V., Tobler, R. & Schlotterer, C. Patterns of linkage disequilibrium and long range hitchhiking in evolving experimental Drosophila melanogaster populations. Mol. Biol. Evol. 32, 495–509 (2015).
Horn, T., Narov, K. D. & Panfilio, K. A. Persistent parental RNAi in the beetle Tribolium castaneum involves maternal transmission of long double-stranded RNA (Advanced Genetics 3/03). Adv. Genet. 3, 2270031 (2022).
Dermauw, W., Van Leeuwen, T. & Feyereisen, R. Diversity and evolution of the P450 family in arthropods. Insect Biochem. Mol. Biol. 127, 103490 (2020).
Rewitz, K. F., O’Connor, M. B. & Gilbert, L. I. Molecular evolution of the insect Halloween family of cytochrome P450s: phylogeny, gene organization and functional conservation. Insect Biochem. Mol. Biol. 37, 741–753 (2007).
Zhu, F., Moural, T. W., Shah, K. & Palli, S. R. Integrated analysis of cytochrome P450 gene superfamily in the red flour beetle, Tribolium castaneum. BMC Genomics 14, 174 (2013).
Zera, A. J., Zhao, Z. & Kaliseck, K. Hormones in the field: evolutionary endocrinology of juvenile hormone and ecdysteroids in field populations of the wing-dimorphic cricket Gryllus firmus. Physiol. Biochem. Zool. 80, 592–606 (2007).
Oostra, V. et al. Translating environmental gradients into discontinuous reaction norms via hormone signalling in a polyphenic butterfly. Proc. Biol. Sci. 278, 789–797 (2011).
Oostra, V. et al. Ecdysteroid hormones link the juvenile environment to alternative adult life histories in a seasonal insect. Am. Nat. 184, E79–E92 (2014).
Zijlstra, W. G., Steigenga, M. J., Koch, P. B., Zwaan, B. J. & Brakefield, P. M. Butterfly selected lines explore the hormonal basis of interactions between life histories and morphology. Am. Nat. 163, E76–E87 (2004).
Zera, A. J. & Bottsford, J. The endocrine-genetic basis of life-history variation: the relationship between the ecdysteroid titer and morph-specific reproduction in the wing-polymorphic cricket Gryllus firmus. Evolution 55, 538–549 (2001).
Zera, A. J., Harshman, L. G. & Williams, T. D. Evolutionary endocrinology: the developing synthesis between endocrinology and evolutionary genetics. Annu. Rev. Ecol. Evol. Syst. 38, 793–817 (2007).
Pointer, M. D., Gage, M. J. G. & Spurgin, L. G. Tribolium beetles as a model system in evolution and ecology. Heredity 126, 869–883 (2021).
Charlesworth, B. Causes of natural variation in fitness: evidence from studies of Drosophila populations. Proc. Natl Acad. Sci. USA 112, 1662–1669 (2015).
Charlesworth, B. & Edwards, A. W. A century of variance. Significance 15, 20–25 (2018).
Dittmar, E. L., Oakley, C. G., Conner, J. K., Gould, B. A. & Schemske, D. W. Factors influencing the effect size distribution of adaptive substitutions. Proc. Biol. Sci. 283, 20153065 (2016).
O’Connor, L. J. et al. Extreme polygenicity of complex traits is explained by negative selection. Am. J. Hum. Genet. 105, 456–476 (2019).
Pray, L. A., Goodnight, C. J., Stevens, L., Schwartz, J. M. & Yan, G. Y. The effect of population size on effective population size: an empirical study in the red flour beetle Tribolium castaneum. Genet. Res. 68, 151–155 (1996).
Therneau, T. M., Grambsch, P. M. & Pankratz, V. S. Penalized survival models and frailty. J. Comput. Graph. Stat. 12, 156–175 (2003).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Koboldt, D. C. et al. VarScan: variant detection in massively parallel sequencing of individual and pooled samples. Bioinformatics 25, 2283–2285 (2009).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w(1118); iso-2; iso-3. Fly 6, 80–92 (2012).
Herndon, N. et al. Enhanced genome assembly and a new official gene set for Tribolium castaneum. BMC Genomics 21, 47 (2020).
Knaus, B. J. & Grunwald, N. J. VCFR: a package to manipulate and visualize variant call format data in R. Mol. Ecol. Resour. 17, 44–53 (2017).
Lord, J. C., Hartzer, K., Toutges, M. & Oppert, B. Evaluation of quantitative PCR reference genes for gene expression studies in Tribolium castaneum after fungal challenge. J. Microbiol. Methods 80, 219–221 (2010).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408 (2001).
van der Zee, M., Stockhammer, O., von Levetzow, C., Nunes da Fonseca, R. & Roth, S. Sog/Chordin is required for ventral-to-dorsal Dpp/BMP transport and head formation in a short germ insect. Proc. Natl Acad. Sci. USA 103, 16307–16312 (2006).
Jacobs, C. G., Rezende, G. L., Lamers, G. E. & van der Zee, M. The extraembryonic serosa protects the insect egg against desiccation. Proc. R. Soc. B 280, 20131082 (2013).
Tautz, D. & Pfeifle, C. A non-radioactive in situ hybridization method for the localization of specific RNAs in Drosophila embryos reveals translational control of the segmentation gene hunchback. Chromosoma 98, 81–85 (1989).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Cheng, S. et al. Rcode for ChengNEE2023. Zenodo https://doi.org/10.5281/zenodo.8395048 (2023).
Gilles, A. F., Schinko, J. B. & Averof, M. Efficient CRISPR-mediated gene targeting and transgene replacement in the beetle Tribolium castaneum. Development 142, 2832–2839 (2015).
Berghammer, A. J., Weber, M., Trauner, J. & Klingler, M. Red flour beetle (Tribolium) germline transformation and insertional mutagenesis. Cold Spring Harb. Protoc. 2009, pdb prot5259 (2009).
Castro-Mondragon, J. A. et al. JASPAR 2022: the 9th release of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 50, D165–D173 (2022).
Handel, K., Grunfelder, C. G., Roth, S. & Sander, K. Tribolium embryogenesis: a SEM study of cell shapes and movements from blastoderm to serosal closure. Dev. Genes Evol. 210, 167–179 (2000).
Hilbrant, M., Horn, T., Koelzer, S. & Panfilio, K. A. The beetle amnion and serosa functionally interact as apposed epithelia. Elife 5, e13834 (2016).
Acknowledgements
We thank R. Jacobs for building the selection machine; K. Koops for taking care of the beetles; G. Lamers and J. Willemse for help with confocal live imaging; and M. Averof and B. Zwaan for experimental advice. M. Averof, P. Brakefield, M. Richardson and B. Zwaan substantially improved the manuscript. S.C. and D.C. were supported by the Chinese Scholarship Council (scholarship 201808210285 to S.C. and 201906280500 to D.C.).
Author information
Authors and Affiliations
Contributions
J.v.d.H. and M.v.d.Z. share last authorship of this work. S.C. analysed and performed developmental staging, qPCR, RNAi, genotyping, ecdysone measurements, microscopy and CRISPR/Cas9 gene editing. E.A.M.P. contributed to qPCR and data analysis. J.v.d.S. contributed to RNAi and genotyping. K.M.B., F.V. and J.S.B. collected life-history data. A.H. performed in situ hybridization. C.G.C.J. performed and analysed the artificial selection process. D.C. developed and analysed the dynamic model of Cyp18a1 and ecdysone regulation. R.M.H.M. supervised this mathematical modelling. J.v.d.H. analysed and visualized the pooled resequencing data and life-history data, and developed the population genetic model. C.G.C.J. and M.v.d.Z. conceived the study. S.C., C.G.C.J., J.v.d.H. and M.v.d.Z. designed the experiments. M.v.d.Z. wrote the paper with substantial input from J.v.d.H., and feedback from S.C., C.G.C.J., E.A.M.P. and R.M.H.M.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Ecology & Evolution thanks Sheng Li, Karl Gotthard, Enoch Ng’oma and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
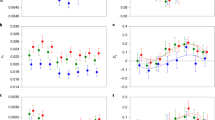
Extended Data Fig. 1 Life history traits of the selection lines.
(a) Pupal DT (days from pupation to eclosion; means ± 2 × s.e.m.) of the selection lines. Selection regime had an effect (χ2 = 10.75; df = 2; p = 0.046): pupae from the fast lines develop significantly faster than those of the non-selected lines (χ2 = 8.13; df = 1; p = 0.004). Sex had no effect (χ2 = 0.017; df = 1; p = 0.897). N = 48 for each selection line. b, Total postembryonic developmental time of the selection lines (days from hatching to eclosion; means ± 2 × s.e.m.). Selection regime had an effect (χ2 = 22.98; df = 2; p = 0.00001): fast lines develop significantly faster than the non-selected lines (χ2 = 19.43; df = 1; p = 0.00001). Sex had no effect (χ2 = 0.2647; df = 1; p = 0.61). N = 48 for each selection line. (c) Growth rate defined as adult weight (see e) divided by total postembryonic time (see b) of an individual (mg/day; means ± 2 x s.e.m.). Selection regime had an effect (χ2 = 11.099; df = 2; p = 0.0039), but the fast lines are not significantly different from the slow lines (χ2 = 2.34; df = 1; p = 0.127). Sex had no significant effect (χ2 = 3.42; df = 1; p = 0.065). N = 48 for each selection line. (d) Pupal weight in the selection lines (mg; means ± 2 x s.e.m). Sex had an effect; closed circles are male, open circles are female means (χ2 = 4.24; df = 1; p = 0.039). Selection regime also had an effect (χ2 = 17.84; df = 2; p = 0.00013): pupae from the slow lines are heavier than those of the non-selected lines (χ2 = 13.09; df = 1; p = 0.0003). N = 48 for each selection line. (e) Adult weight in the selection lines (mg; means ± 2 x s.e.m.). Sex had an effect (χ2 = 6.04; df = 1; p = 0.014), closed circles are male, open circles are female means. Selection regime also had an effect (χ2 = 11.08; df = 2; p = 0.039): adults of the slow lines are significantly heavier than adults of the non-selected lines (χ2 = 7.78; df = 1; p = 0.005). N = 48 for each selection line. (f) Life span in the selection lines (weeks as adult; means ± 2 × s.e.m). Selection regime had no effect (χ2 = 4.6, df = 2, p = 0.10). N = 90 for each selection line, see Methods.
Extended Data Fig. 2 Numerical staging table for Tribolium castaneum.
Largely based on93,94. (a) 0 = no nuclei at surface; (b) 1 = undifferentiated blastoderm (equal nuclei at the surface). (c) 2 = differentiated blastoderm (the large polyploid nuclei of the serosa can be distinguished from the more dense, smaller nuclei of the germ rudiment). (d) 3 = gastrulation (amnion and serosa fold over the embryo; serosal window not yet closed). (e) 4 = extending germband (serosa closed). (f) 5 = extended germ band (distance between posterior end and head is small, size bar; limbs are buds). (g) 6 = limbs growing and extending (head and posterior of the germband still close, see size bar; limbs well developing and extending). (h) 7 = retracting germband (dorsal distance between head and posterior end of the embryo increases again). (i) 8 = completely retracted germband. (j) 9 = start of dorsal closure (rupture of the extraembryonic membranes, dorsal organ formation). (k) 10 = dorsal closure in progress (dorsal organ flatter, lateral sides of the embryo have moved towards dorsal). (l) 11 = dorsal closure completed (lateral sides of the embryo dorsally fused). (m) 12 = hatching (still in vitelline membrane). (n) 13 = hatched (out of vitelline membrane). Scalebar in (a) = 200 µm and applies to all panels.
Extended Data Fig. 3 Manhattan plot of SNPs that differ in allele frequency between the Slow and Non-Selected lines.
P values of SNPs, based on a GLM contrasting the slow (n = 2) and the non-selected (n = 2) lines (see Methods), along the 10 chromosomes of Tribolium castaneum. In total, 1258 SNPs differ in frequency significantly (above the red line = q < 0.01, −logep > 33.0538).
Extended Data Fig. 4 Embryonic phenotypes upon Cyp18a1 pRNAi.
(a) Numbers (percentages) of the different phenotypic classes from one fixation analysis (upper 4 rows), in which Cyp18a1 dsRNA injected mothers were allowed to lay eggs for one day, and batches of 25-50 eggs were fixed every subsequent day for DAPI staining and microscopic analysis. Percentages of developmental arrests are indicated with dark grey background, normal development with white background. The total number of developmental arrests corresponded well to the number of unhatched eggs after the pRNAi screen for developmental time (lower two rows). In 20% of the embryos, we did not observe any start of development at all (upper row), compared to 7.8% in the control RNAi. No developmental arrests were observed upon control pRNAi. (b) Control RNAi, normal blastoderm stage. (c) Cyp18a1 RNAi. Developmental arrests at the blastoderm stage were recognized by irregular positioning of aberrant nuclei. (d) Control RNAi. Normal extending germband (fully extended in (d)). (e) Developmental arrests during germband extension were recognized by shortened and irregular germbands. (f) Control RNAi, normal dorsal closure. (g) Cyp18a1 RNAi. Embryos that were arrested during dorsal closure were still open at the dorsal side, but showed otherwise advanced development. Scale bar in (b) = 200 μm and applies to panels b-g.
Extended Data Fig. 5 Allele frequency of F in a natural population is 0.81.
Genotyping PCR (Ethidium bromide staining shown on agarose gel) of one sample of 48 beetles from the (expanded) wild population collected by Rentokil from a bakery in The Netherlands (see Methods). 31 show only a 740 bp band (homozygous F allele), 2 show only a 942 bp band (homozygous S allele), and 14 are heterozygote (both bands + a hybrid band present). Thus the allele frequency of F is 0.81, and S is 0.19. Ladder is GeneRuler 1kb plus DNA ladder (Invitrogen).
Supplementary information
Supplementary Information
Modelling theory.
Supplementary Tables 1–4
Supplementary Tables 1–4.
Supplementary Video 1
Live imaging of a representative GA-1/nGFP (upper) and a representative CRISPR/nGFP (lower) heterozygote embryo. Time since egg lay is indicated in the middle (hh:mm:ss). The completely retracted germband stage (indicated by appearing text) occurs at the same moment. However, the time interval between the completely retracted germband and the start of dorsal closure (indicated by appearing text) is shorter in the CRISPR/nGFP embryo.
Supplementary Data 1
Alignments of cloned Sanger-sequenced PCR products from the selection lines.
Supplementary Data 2
FIMO output. See Methods.
Supplementary Data 3
Aligned sequences of the CRISPR alleles present in our stock.
Source data
Source Data Fig. 1c,d, 2d, 3a,f, 5d,e and Extended Data Figs. 1a–e,f
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cheng, S., Jacobs, C.G.C., Mogollón Pérez, E.A. et al. A life-history allele of large effect shortens developmental time in a wild insect population. Nat Ecol Evol 8, 70–82 (2024). https://doi.org/10.1038/s41559-023-02246-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-023-02246-y