Abstract
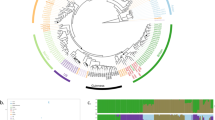
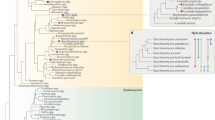
The most common fermented beverage, lager beer, is produced by interspecies hybrids of the brewing yeast Saccharomyces cerevisiae and its wild relative S. eubayanus. Lager-brewing yeasts are not the only example of hybrid vigour or heterosis in yeasts, but the full breadth of interspecies hybrids associated with human fermentations has received less attention. Here we present a comprehensive genomic analysis of 122 Saccharomyces hybrids and introgressed strains. These strains arose from hybridization events between two to four species. Hybrids with S. cerevisiae contributions originated from three lineages of domesticated S. cerevisiae, including the major wine-making lineage and two distinct brewing lineages. In contrast, the undomesticated parents of these interspecies hybrids were all from wild Holarctic or European lineages. Most hybrids have inherited a mitochondrial genome from a parent other than S. cerevisiae, which recent functional studies suggest could confer adaptation to colder temperatures. A subset of hybrids associated with crisp flavour profiles, including both lineages of lager-brewing yeasts, have inherited inactivated S. cerevisiae alleles of critical phenolic off-flavour genes and/or lost functional copies from the wild parent through multiple genetic mechanisms. These complex hybrids shed light on the convergent and divergent evolutionary trajectories of interspecies hybrids and their impact on innovation in lager brewing and other diverse fermentation industries.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
References and accession numbers for the published data used can be found in Supplementary Dataset 16. Short-read data (newly published here) are available through the NCBI SRA database under the BioProject accession number PRJNA522928. Assembled genomes published here are available under GenBank BioProject PRJNA522928.
Code availability
Custom R and Python scripts used for this publication can be found on GitHub (https://github.com/qlangdon/hybrid-ferment-invent).
References
Hornsey, I. S. Alcohol and Its Role in the Evolution of Human Society (RSC Publishing, 2012).
Fay, J. C. & Benavides, J. A. Evidence for domesticated and wild populations of Saccharomyces cerevisiae. PLoS Genet. 1, e5 (2005).
Liti, G., Peruffo, A., James, S. A., Roberts, I. N. & Louis, E. J. Inferences of evolutionary relationships from a population survey of LTR-retrotransposons and telomeric-associated sequences in the Saccharomyces sensu stricto complex. Yeast 22, 177–192 (2005).
Gallone, B. et al. Origins, evolution, domestication and diversity of Saccharomyces beer yeasts. Curr. Opin. Biotechnol. 49, 148–155 (2018).
Legras, J. L. et al. Adaptation of S. cerevisiae to fermented food environments reveals remarkable genome plasticity and the footprints of domestication. Mol. Biol. Evol. 35, 1712–1727 (2018).
Rodríguez, M. E. et al. Saccharomyces uvarum is responsible for the traditional fermentation of apple chicha in Patagonia. FEMS Yeast Res. 17, fow109 (2017).
Barbosa, R. et al. Multiple rounds of artificial selection promote microbe secondary domestication—the case of cachaça yeasts. Genome Biol. Evol. 10, 1939–1955 (2018).
Gallone, B. et al. Domestication and divergence of Saccharomyces cerevisiae beer yeasts. Cell 166, 1397–1410 (2016).
Gonçalves, M. et al. Distinct domestication trajectories in top-fermenting beer yeasts and wine yeasts. Curr. Biol. 26, 1–12 (2016).
Duan, S. F. et al. The origin and adaptive evolution of domesticated populations of yeast from Far East Asia. Nat. Commun. 9, 2690 (2018).
Peter, J. et al. Genome evolution across 1,011 Saccharomyces cerevisiae isolates. Nature 556, 339–344 (2018).
Marsit, S. & Dequin, S. Diversity and adaptive evolution of Saccharomyces wine yeast: a review. FEMS Yeast Res. 15, 1–12 (2015).
Almeida, P., Barbosa, R., Bensasson, D., Gonçalves, P. & Sampaio, J. P. Adaptive divergence in wine yeasts and their wild relatives suggests a prominent role for introgressions and rapid evolution at noncoding sites. Mol. Ecol. 26, 2167–2182 (2017).
Hittinger, C. T., Steele, J. L. & Ryder, D. S. Diverse yeasts for diverse fermented beverages and foods. Curr. Opin. Biotechnol. 49, 199–206 (2018).
Gibson, B. & Liti, G. Saccharomyces pastorianus: genomic insights inspiring innovation for industry. Yeast 32, 17–27 (2015).
Libkind, D. et al. Microbe domestication and the identification of the wild genetic stock of lager-brewing yeast. Proc. Natl Acad. Sci. USA 108, 14539–14544 (2011).
Baker, E. P. et al. Mitochondrial DNA and temperature tolerance in lager yeasts. Sci. Adv. 5, eaav1869 (2019).
Baker, E. P. & Hittinger, C. T. Evolution of a novel chimeric maltotriose transporter in Saccharomyces eubayanus from parent proteins unable to perform this function. PLoS Genet. 15, e1007786 (2019).
Hebly, M. et al. S. cerevisiae × S. eubayanus interspecific hybrid, the best of both worlds and beyond. FEMS Yeast Res. 15, 1–14 (2015).
Gibson, B. R., Storgårds, E., Krogerus, K. & Vidgren, V. Comparative physiology and fermentation performance of Saaz and Frohberg lager yeast strains and the parental species Saccharomyces eubayanus. Yeast 30, 255–266 (2013).
Gorter de Vries, A. R. et al. Laboratory evolution of a Saccharomyces cerevisiae x S. eubayanus hybrid under simulated lager-brewing conditions. Front. Genet. 10, 242 (2019).
Monerawela, C. & Bond, U. Brewing up a storm: the genomes of lager yeasts and how they evolved. Biotechnol. Adv. 35, 512–519 (2017).
Peris, D., Pérez-Torrado, R., Hittinger, C. T., Barrio, E. & Querol, A. On the origins and industrial applications of Saccharomyces cerevisiae × Saccharomyces kudriavzevii hybrids. Yeast 35, 51–69 (2018).
Nguyen, H. V. & Boekhout, T. Characterization of Saccharomyces uvarum (Beijerinck, 1898) and related hybrids: assessment of molecular markers that predict the parent and hybrid genomes and a proposal to name yeast hybrids. FEMS Yeast Res. 17, 1–19 (2017).
Nguyen, H. V., Legras, J. L., Neuvéglise, C. & Gaillardin, C. Deciphering the hybridisation history leading to the lager lineage based on the mosaic genomes of Saccharomyces bayanus strains NBRC1948 and CBS380 T. PLoS ONE 6, e25821 (2011).
Almeida, P. et al. A Gondwanan imprint on global diversity and domestication of wine and cider yeast Saccharomyces uvarum. Nat. Commun. 5, 4044 (2014).
Dunn, B. & Sherlock, G. Reconstruction of the genome origins and evolution of the hybrid lager yeast Saccharomyces pastorianus. Genome Res. 18, 1610–1623 (2008).
Hittinger, C. T. Saccharomyces diversity and evolution: a budding model genus. Trends Genet. 29, 309–317 (2013).
Boynton, P. J. & Greig, D. The ecology and evolution of non-domesticated Saccharomyces species. Yeast 31, 449–462 (2014).
Hittinger, C. T. et al. Remarkably ancient balanced polymorphisms in a multi-locus gene network. Nature 464, 54–58 (2010).
Sampaio, J. P. & Gonçalves, P. Natural populations of Saccharomyces kudriavzevii in Portugal are associated with oak bark and are sympatric with S. cerevisiae and S. paradoxus. Appl. Environ. Microbiol. 74, 2144–2152 (2008).
Peris, D. et al. Complex ancestries of lager-brewing hybrids were shaped by standing variation in the wild yeast Saccharomyces eubayanus. PLoS Genet. 12, e1006155 (2016).
Salvadó, Z., Arroyo-López, F. N., Barrio, E., Querol, A. & Guillamón, J. M. Quantifying the individual effects of ethanol and temperature on the fitness advantage of Saccharomyces cerevisiae. Food Microbiol. 28, 1155–1161 (2011).
Gonçalves, P., Valério, E., Correia, C., de Almeida, J. M. G. C. F. & Sampaio, J. P. Evidence for divergent evolution of growth temperature preference in sympatric Saccharomyces species. PLoS ONE 6, e20739 (2011).
Li, X. C., Peris, D., Hittinger, C. T., Sia, E. A. & Fay, J. C. Mitochondria-encoded genes contribute to evolution of heat and cold tolerance in yeast. Sci. Adv. 5, eaav1848 (2019).
Ortiz-Tovar, G., Pérez-Torrado, R., Adam, A. C., Barrio, E. & Querol, A. A comparison of the performance of natural hybrids Saccharomyces cerevisiae × Saccharomyces kudriavzevii at low temperatures reveals the crucial role of their S. kudriavzevii genomic contribution. Int. J. Food Microbiol. 274, 12–19 (2018).
Tronchoni, J., Medina, V., Guillamón, J. M., Querol, A. & Pérez-Torrado, R. Transcriptomics of cryophilic Saccharomyces kudriavzevii reveals the key role of gene translation efficiency in cold stress adaptations. BMC Genomics 15, 1–10 (2014).
Huh, K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003).
Chou, J. Y., Hung, Y. S., Lin, K. H., Lee, H. Y. & Leu, J. Y. Multiple molecular mechanisms cause reproductive isolation between three yeast species. PLoS Biol. 8, e1000432 (2010).
Lee, H. Y. et al. Incompatibility of nuclear and mitochondrial genomes causes hybrid sterility between two yeast species. Cell 135, 1065–1073 (2008).
Hou, J. & Schacherer, J. Negative epistasis: a route to intraspecific reproductive isolation in yeast? Curr. Genet. 62, 25–29 (2016).
Novo, M. et al. Eukaryote-to-eukaryote gene transfer events revealed by the genome sequence of the wine yeast Saccharomyces cerevisiae EC1118. Proc. Natl Acad. Sci. USA 106, 16333–16338 (2009).
Ashburner, M. et al. Gene Ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Consortium, T. G. O. The gene ontology resource: 20 years and still GOing strong. Nucleic Acids Res. 47, D330–D338 (2019).
Han, E.-K., Cotty, F., Sottas, C., Jiang, H. & Michels, C. A. Characterization of AGT1 encoding a general alpha-glucoside transporter from Saccharomyces. Mol. Microbiol. 17, 1093–1107 (1995).
Salema-Oom, M., Pinto, V. V., Gonçalves, P. & Spencer-Martins, I. Maltotriose utilization by industrial Saccharomyces strains: characterization of a new member of the alpha-glucoside transporter family. Appl. Environ, Microbiol. 71, 5044–5049 (2005).
Horák, J. Regulations of sugar transporters: insights from yeast. Curr. Genet. 59, 1–31 (2013).
Dietvorst, J., Londesborough, J. & Steensma, H. Y. Maltotriose utilization in lager yeast strains: MTTI encodes a maltotriose transporter. Yeast 22, 775–788 (2005).
Diderich, J. A., Weening, S. M., van den Broek, M., Pronk, J. T. & Daran, J.-M. G. Selection of Pof-Saccharomyces eubayanus variants for the construction of S. cerevisiae × S. eubayanus hybrids with reduced 4-vinyl guaiacol formation. Front. Microbiol. 9, 1640 (2018).
Mukai, N., Masaki, K., Fujii, T., Kawamukai, M. & Iefuji, H. PAD1 and FDC1 are essential for the decarboxylation of phenylacrylic acids in Saccharomyces cerevisiae. J. Biosci. Bioeng. 109, 564–569 (2010).
Shen, X.-X. et al. Tempo and mode of genome evolution in the budding yeast subphylum. Cell 175, 1533–1545 (2018).
Bing, J., Han, P.-J., Liu, W.-Q., Wang, Q.-M. & Bai, F.-Y. Evidence for a Far East Asian origin of lager beer yeast. Curr. Biol. 24, R380–R381 (2014).
Borneman, A. R., Forgan, A. H., Pretorius, I. S. & Chambers, P. J. Comparative genome analysis of a Saccharomyces cerevisiae wine strain. FEMS Yeast Res. 8, 1185–1195 (2008).
Borneman, A. R. et al. Whole-genome comparison reveals novel genetic elements that characterize the genome of industrial strains of Saccharomyces cerevisiae. PLoS Genet. 7, e1001287 (2011).
Borneman, A. R., Forgan, A. H., Kolouchova, R., Fraser, J. A. & Schmidt, S. A. Whole genome comparison reveals high levels of inbreeding and strain redundancy across the spectrum of commercial wine strains of Saccharomyces cerevisiae. G3 6, 957–971 (2016).
Dunn, B., Richter, C., Kvitek, D. J., Pugh, T. & Sherlock, G. Analysis of the Saccharomyces cerevisiae pan-genome reveals a pool of copy number variants distributed in diverse yeast strains from differing industrial environments. Genome Res. 22, 908–924 (2012).
Gayevskiy, V. & Goddard, M. R. Saccharomyces eubayanus and Saccharomyces arboricola reside in North Island native New Zealand forests. Environ. Microbiol. 18, 1137–1147 (2016).
Gayevskiy, V., Lee, S. & Goddard, M. R. European derived Saccharomyces cerevisiae colonisation of New Zealand vineyards aided by humans. FEMS Yeast Res. 16, 1–12 (2016).
Hewitt, S. K., Donaldson, I. J., Lovell, S. C. & Delneri, D. Sequencing and characterisation of rearrangements in three S. pastorianus strains reveals the presence of chimeric genes and gives evidence of breakpoint reuse. PLoS ONE 9, e92203 (2014).
Hose, J. et al. Dosage compensation can buffer copynumber variation in wild yeast. eLife 4, 1–28 (2015).
Krogerus, K., Preiss, R. & Gibson, B. A unique Saccharomyces cerevisiae × Saccharomyces uvarum hybrid isolated from Norwegian farmhouse beer: characterization and reconstruction. Front. Microbiol. 9, 1–15 (2018).
Okuno, M. et al. Next-generation sequencing analysis of lager brewing yeast strains reveals the evolutionary history of interspecies hybridization. DNA Res. 1, 1–14 (2016).
Scannell, D. R. et al. The awesome power of yeast evolutionary genetics: new genome sequences and strain resources for the Saccharomyces sensu stricto genus. G3 1, 11–25 (2011).
Skelly, D. A. et al. Integrative phenomics reveals insight into the structure of phenotypic diversity in budding yeast. Genome Res. 23, 1496–1504 (2013).
Strope, P. K. et al. The 100-genomes strains, an S. cerevisiae resource that illuminates its natural phenotypic and genotypic variation and emergence as an opportunistic pathogen. Genome Res. 125, 762–774 (2015).
van den Broek, M. et al. Chromosomal copy number variation in Saccharomyces pastorianus is evidence for extensive genome dynamics in industrial lager brewing strains. Appl. Environ. Microbiol. 81, 6253–6267 (2015).
Yue, J. X. et al. Contrasting evolutionary genome dynamics between domesticated and wild yeasts. Nat. Genet. 49, 913–924 (2017).
Zheng, D. Q. et al. Genome sequencing and genetic breeding of a bioethanol Saccharomyces cerevisiae strain YJS329. BMC Genomics 13, 479 (2012).
Bergström, A. et al. A high-definition view of functional genetic variation from natural yeast genomes. Mol. Biol. Evol. 31, 872–888 (2014).
Akao, T. et al. Whole-genome sequencing of sake yeast Saccharomyces cerevisiae Kyokai no. 7. DNA Res. 18, 423–434 (2011).
Almeida, P. et al. A population genomics insight into the Mediterranean origins of wine yeast domestication. Mol. Ecol. 24, 5412–5427 (2015).
Baker, E. et al. The genome sequence of Saccharomyces eubayanus and the domestication of lager-brewing yeasts. Mol. Biol. Evol. 32, 2818–2831 (2015).
Langdon, Q. K., Peris, D., Kyle, B. & Hittinger, C. T. sppIDer: a species identification tool to investigate hybrid genomes with high-throughput sequencing. Mol. Biol. Evol. 35, 2835–2849 (2018).
Liti, G. et al. Population genomics of domestic and wild yeasts. Nature 458, 337–341 (2009).
Liti, G. et al. High quality de novo sequencing and assembly of the Saccharomyces arboricolus genome. BMC Genomics 14, 69 (2013).
Peris, D. et al. Biotechnology for biofuels hybridization and adaptive evolution of diverse Saccharomyces species for cellulosic biofuel production. Biotechnol. Biofuels 10, 1–19 (2017).
Teytelman, L. et al. Impact of chromatin tructures on DNA processing for genomic analyses. PLoS ONE 4, e6700 (2009).
Suzuki, R. & Shimodaira, H. Pvclust: an R package for assessing the uncertainty in hierarchical clustering. Bioinformatics 22, 1540–1542 (2006).
McKenna, A. et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Zhou, X. et al. In silico whole genome sequencer and analyzer (iWGS): a computational pipeline to guide the design and analysis of de novo genome sequencing studies. G3 6, 3655–3662 (2016).
Layer, R. M., Chiang, C., Quinlan, A. R. & Hall, I. M. LUMPY: a probabilistic framework for structural variant discovery. Genome Biol. 15, R84 (2014).
Foury, F., Roganti, T., Lecrenier, N. & Purnelle, B. The complete sequence of the mitochondrial genome of Saccharomyces cerevisiae. FEBS Lett. 440, 325–331 (1998).
Sulo, P. et al. The evolutionary history of Saccharomyces species inferred from completed mitochondrial genomes and revision in the ‘yeast mitochondrial genetic code’. DNA Res. 24, 571–583 (2017).
Peris, D. et al. Molecular phylogenetics and evolution mitochondrial introgression suggests extensive ancestral hybridization events among Saccharomyces species. Mol. Phylogenet. Evol. 108, 49–60 (2017).
Johnson, M. G. et al. HybPiper: extracting coding sequence and introns for phylogenetics from high-throughput sequencing reads using target enrichment. Appl. Plant Sci. https://doi.org/10.3732/apps.1600016 (2016).
Kearse, M. et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649 (2012).
Leigh, J. W. & Bryant, D. POPART: full-feature software for haplotype network construction. Methods Ecol. Evol. 6, 1110–1116 (2015).
Clement, M., Posada, D. & Crandall, K. A. TCS: a computer program to estimate gene genealogies. Mol. Ecol. 9, 1657–1659 (2000).
Librado, P. & Rozas, J. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452 (2009).
Walther, A., Hesselbart, A. & Wendland, J. Genome sequence of Saccharomyces carlsbergensis, the world’s first pure culture lager yeast. G3 4, 783–793 (2014).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Csardi, G. & Nepusz, T. The igraph software package for complex network research. InterJournal 1695, 1–9 (2006).
Opulente, D. A. et al. Factors driving metabolic diversity in the budding yeast subphylum. BMC Biol. 16, 1–15 (2018).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 44, W242–W245 (2016).
Jombart, T. adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24, 1403–1405 (2008).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer-Verlag, 2009).
Pfeifer, B., Wittelsbürger, U., Ramos-Onsins, S. E. & Lercher, M. J. PopGenome: an efficient Swiss army knife for population genomic analyses in R. Mol. Biol. Evol. 31, 1929–1936 (2014).
Acknowledgements
We thank K.J. Verstrepen for coordinating publication with their study; A.B. Hulfachor and M. Bontrager for preparing a subset of Illumina libraries; the University of Wisconsin Biotechnology Center DNA Sequencing Facility for providing Illumina sequencing facilities and services; M.-A. Lachance, A. Kinart, D.T. Doering, R. Thiel and D. Carey for strains; and M. Langdon, A.B. Hulfachor and K. Sylvester for collecting fermentation samples and/or isolating strains. This material is based upon work supported by the National Science Foundation (grant nos. DEB-1253634 to C.T.H. and DGE-1256259 (Graduate Research Fellowship to Q.K.L.), the USDA National Institute of Food and Agriculture Hatch Project (nos. 1003258 and 1020204 to C.T.H.) and in part by the DOE Great Lakes Bioenergy Research Center (DOE BER Office of Science nos. DE-SC0018409 and DE-FC02-07ER64494). Q.K.L. was also supported by the Predoctoral Training Program in Genetics, funded by the National Institutes of Health (grant no. 5T32GM007133). D.P. is a Marie Sklodowska-Curie fellow of the European Union’s Horizon 2020 research and innovation programme (grant no. 747775). E.P.B. was supported by a Louis and Elsa Thomsen Wisconsin Distinguished Graduate Fellowship. U.B. is funded by Horizon 2020 MSCA-ITN grant no. 764364. D.L. was supported by CONICET (grant no. PIP 392), FONCyT (grant no. PICT 3677) and Universidad Nacional del Comahue (grant no. B199). C.T.H. is a Pew Scholar in the Biomedical Sciences, Vilas Faculty Early Career Investigator and H.I. Romnes Faculty Fellow, supported by the Pew Charitable Trusts, Vilas Trust Estate and Office of the Vice Chancellor for Research and Graduate Education with funding from the Wisconsin Alumni Research Foundation, respectively.
Author information
Authors and Affiliations
Contributions
Q.K.L. performed most analyses with assistance from D.A.O. D.P. and Q.K.L. performed mitochondrial genome analyses and drafted text. E.P.B. and Q.K.L. analysed genes of functional interest and drafted text. Q.K.L., E.P.B. and D.A.O. sequenced genomes. H.-V.N., U.B., P.G. and J.P.S. contributed key strains to study design. Q.K.L., D.P., E.P.B., D.L. and C.T.H. designed the study. Q.K.L. and C.T.H. wrote the manuscript with editorial input from all coauthors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary text and Supplementary Figs. 1–15.
Supplementary Dataset 1
All hybrids and their parent contributions.
Supplementary Dataset 2
PCA analyses. Percent explained by each principal component included in column headers.
Supplementary Dataset 3
Newick formatted file of the S. kudriavzevii phylogeny with major hybrids.
Supplementary Dataset 4
Newick formatted file of the S. kudriavzevii phylogeny with minor hybrids.
Supplementary Dataset 5
Newick formatted file of the S. eubayanus phylogeny with major hybrids.
Supplementary Dataset 6
Newick formatted file of the S. eubayanus phylogeny with minor hybrids.
Supplementary Dataset 7
Newick formatted file of the S. uvarum phylogeny with major hybrids.
Supplementary Dataset 8
Newick formatted file of the S. uvarum phylogeny with minor hybrids.
Supplementary Dataset 9
Newick formatted file of the S. cerevisiae phylogeny with all strains analysed.
Supplementary Dataset 10
Newick formatted file of the S. cerevisiae phylogeny of just the Ale/Beer1 clade.
Supplementary Dataset 11
Results of Fisher’s exact test and Bonferroni correction of mitochondrially localized genes. mtInteracting = nuclear-encoded but mitochondrially localized gene.
Supplementary Dataset 12
Summary of number of 1:1:1:1 orthologues present in each subgenome.
Supplementary Dataset 13
GO term results of genes found in novel regions of the de novo assembled genomes.
Supplementary Dataset 14
Brewing relevant gene summaries. 30“-“ Indicates when HybPiper failed to recover and assemble genes for this group or that these assemblies failed our length and coverage cutoffs.
Supplementary Dataset 15
Metadata for all strains newly sequenced in this study. The “New hybrid” column denotes hybrid genome sequences that are newly published in this study.
Supplementary Dataset 16
Published data accession information.
Supplementary Dataset 17
Haplotype key for mitochondrial genomes, PAD1 and FDC1. Dataset A only includes strains where 15S rRNA could be assembled, while Dataset B has 15S rRNA removed.
Supplementary Dataset 18
Regions used for minor contribution analyses.
Rights and permissions
About this article
Cite this article
Langdon, Q.K., Peris, D., Baker, E.P. et al. Fermentation innovation through complex hybridization of wild and domesticated yeasts. Nat Ecol Evol 3, 1576–1586 (2019). https://doi.org/10.1038/s41559-019-0998-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-019-0998-8
This article is cited by
-
Evolution of loss of heterozygosity patterns in hybrid genomes of Candida yeast pathogens
BMC Biology (2023)
-
Macroevolutionary diversity of traits and genomes in the model yeast genus Saccharomyces
Nature Communications (2023)
-
Selection of Saccharomyces eubayanus strains from Patagonia (Argentina) with brewing potential and performance in the craft beer industry
European Food Research and Technology (2022)
-
Trends in yeast diversity discovery
Fungal Diversity (2022)
-
The International Scientific Association for Probiotics and Prebiotics (ISAPP) consensus statement on fermented foods
Nature Reviews Gastroenterology & Hepatology (2021)