Abstract
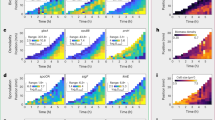
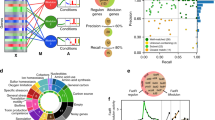
Microbes are exposed to changing environments, to which they can respond by adopting various lifestyles such as swimming, colony formation or dormancy. These lifestyles are often studied in isolation, thereby giving a fragmented view of the life cycle as a whole. Here, we study lifestyles in the context of this whole. We first use machine learning to reconstruct the expression changes underlying life cycle progression in the bacterium Bacillus subtilis, based on hundreds of previously acquired expression profiles. This yields a timeline that reveals the modular organization of the life cycle. By analysing over 380 Bacillales genomes, we then show that life cycle modularity gives rise to mosaic evolution in which life stages such as motility and sporulation are conserved and lost as discrete units. We postulate that this mosaic conservation pattern results from habitat changes that make these life stages obsolete or detrimental. Indeed, when evolving eight distinct Bacillales strains and species under laboratory conditions that favour colony growth, we observe rapid and parallel losses of the sporulation life stage across species, induced by mutations that affect the same global regulator. We conclude that a life cycle perspective is pivotal to understanding the causes and consequences of modularity in both regulation and evolution.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The flow cytometry data are publically available through the FlowRepository, accession No. FR-FCM-ZYVN. The sequencing reads are publically available on the European Nucleotide Archive (ENA) database, accession No. PRJEB32792. The remaining data are included in the Supplementary information or available through public repositories as mentioned in the Supplementary information.
References
Kolenbrander, P. E., Palmer, R. J., Periasamy, S. & Jakubovics, N. S. Oral multispecies biofilm development and the key role of cell–cell distance. Nat. Rev. Microbiol. 8, 471–480 (2010).
Flores, E. & Herrero, A. Compartmentalized function through cell differentiation in filamentous cyanobacteria. Nat. Rev. Microbiol. 8, 39–50 (2010).
McDougald, D., Rice, S. A., Barraud, N., Steinberg, P. D. & Kjelleberg, S. Should we stay or should we go: Mechanisms and ecological consequences for biofilm dispersal. Nat. Rev. Microbiol. 10, 39–50 (2012).
Boutte, C. C. & Crosson, S. Bacterial lifestyle shapes stringent response activation. Trends Microbiol. 21, 174–180 (2013).
Claessen, D., Rozen, D. E., Kuipers, O. P., Søgaard-Andersen, L. & van Wezel, G. P. Bacterial solutions to multicellularity: a tale of biofilms, filaments and fruiting bodies. Nat. Rev. Microbiol. 12, 115–124 (2014).
van Gestel, J., Vlamakis, H. & Kolter, R. Division of labor in biofilms: the ecology of cell differentiation. Microbiol. Spectr. 3, MB-0002–MB-2014 (2015).
Yan, J., Nadell, C. D. & Bassler, B. L. Environmental fluctuation governs selection for plasticity in biofilm production. ISME J. 11, 1569–1577 (2017).
Mhatre, E., Monterrosa, R. G. & Kovács, Á. T. From environmental signals to regulators: modulation of biofilm development in Gram-positive bacteria. J. Basic Microbiol. 54, 616–632 (2014).
Winslow, C. E. A. What do we mean by a bacterial life cycle? Science 81, 314–315 (1935).
O’Toole, G., Kaplan, H. B. & Kolter, R. Biofilm formation as microbial development. Annu. Rev. Microbiol. 54, 49–79 (2000).
Hammerschmidt, K., Rose, C. J., Kerr, B. & Rainey, P. B. Life cycles, fitness decoupling and the evolution of multicellularity. Nature 515, 75–79 (2014).
Stragier, P. & Losick, R. Molecular genetics of sporulation in Bacillus subtilis. Annu. Rev. Genet. 30, 297–341 (1996).
Curtis, P. D. & Brun, Y. V. Getting in the loop: regulation of development in Caulobacter crescentus. Microbiol. Mol. Biol. Rev. 74, 13–41 (2010).
Norman, T. M., Lord, N. D., Paulsson, J. & Losick, R. Memory and modularity in cell-fate decision making. Nature 503, 481–486 (2013).
Russell, J. R., Cabeen, M. T., Wiggins, P. A., Paulsson, J. & Losick, R. Noise in a phosphorelay drives stochastic entry into sporulation in Bacillus subtilis. EMBO J. 36, 2856–2869 (2017). e201796988.
Kearns, D. B. A field guide to bacterial swarming motility. Nat. Rev. Microbiol. 8, 634–644 (2010).
Grau, R. R. et al. A duo of potassium-responsive histidine kinases govern the multicellular destiny of Bacillus subtilis. mBio 6, e00581 (2015).
Cairns, L. S., Hobley, L. & Stanley-Wall, N. R. Biofilm formation by Bacillus subtilis: new insights into regulatory strategies and assembly mechanisms. Mol. Microbiol. 93, 587–598 (2014).
Mielich-Süss, B. & Lopez, D. Molecular mechanisms involved in Bacillus subtilis biofilm formation. Environ. Microbiol. 17, 555–565 (2015).
van Gestel, J., Vlamakis, H. & Kolter, R. From cell differentiation to cell collectives: Bacillus subtilis uses division of labor to migrate. PLoS Biol. 13, e1002141 (2015).
Beauregard, P. B., Chai, Y., Vlamakis, H., Losick, R. & Kolter, R. Bacillus subtilis biofilm induction by plant polysaccharides. Proc. Natl Acad. Sci. USA 110, E1621–E1630 (2013).
Higgins, D. & Dworkin, J. Recent progress in Bacillus subtilis sporulation. FEMS Microbiol. Rev. 36, 131–148 (2012).
Setlow, P. Spore germination. Curr. Opin. Microbiol. 6, 550–556 (2003).
Smits, W. K., Kuipers, O. P. & Veening, J. W. Phenotypic variation in bacteria: the role of feedback regulation. Nat. Rev. Microbiol. 4, 259–271 (2006).
Bonneau, R. et al. The Inferelator: an algorithm for learning parsimonious regulatory networks from systems-biology data sets de novo. Genome Biol. 7, R36 (2006).
Aguilar, C., Vlamakis, H., Losick, R. & Kolter, R. Thinking about Bacillus subtilis as a multicellular organism. Curr. Opin. Microbiol. 10, 638–643 (2007).
Veening, J. W., Smits, W. K. & Kuipers, O. P. Bistability, epigenetics, and bet-hedging in bacteria. Annu. Rev. Microbiol. 62, 193–210 (2008).
Sierro, N., Makita, Y., de Hoon, M. & Nakai, K. DBTBS: a database of transcriptional regulation in Bacillus subtilis containing upstream intergenic conservation information. Nucleic Acids Res. 36, D93–D96 (2008).
Fadda, A. et al. Inferring the transcriptional network of Bacillus subtilis. Mol. Biosyst. 5, 1840–1852 (2009).
Kobayashi, K. Gradual activation of the response regulator DegU controls serial expression of genes for flagellum formation and biofilm formation in Bacillus subtilis. Mol. Microbiol. 66, 395–409 (2007).
Lopez, D. & Kolter, R. Extracellular signals that define distinct and coexisting cell fates in Bacillus subtilis. FEMS Microbiol. Rev. 34, 134–149 (2010).
Freyre-González, J. A. et al. Lessons from the modular organization of the transcriptional regulatory network of Bacillus subtilis. BMC Syst. Biol. 7, 127 (2013).
Leyn, S. A. et al. Genomic reconstruction of the transcriptional regulatory network in Bacillus subtilis. J. Bacteriol. 195, 2463–2473 (2013).
Arrieta‐Ortiz, M. L. et al. An experimentally supported model of the Bacillus subtilis global transcriptional regulatory network. Mol. Syst. Biol. 11, 839 (2015).
Michna, R. H., Zhu, B., Mäder, U. & Stülke, J. SubtiWiki 2.0—an integrated database for the model organism Bacillus subtilis. Nucleic Acids Res. 44, D654–D662 (2016).
Lee, T. I. et al. Transcriptional regulatory networks in Saccharomyces cerevisiae. Science 298, 799–804 (2002).
Sorrells, T. R. & Johnson, A. D. Making sense of transcription networks. Cell 161, 714–723 (2015).
Fang, X. et al. Global transcriptional regulatory network for Escherichia coli robustly connects gene expression to transcription factor activities. Proc. Natl Acad. Sci. USA 114, 10286–10291 (2017).
Santos-Zavaleta, A. et al. RegulonDB v 10.5: tackling challenges to unify classic and high throughput knowledge of gene regulation in E. coli K-12. Nucleic Acids Res. 47, D212–D220 (2019).
Haldenwang, W. G. The sigma factors of Bacillus subtilis. Microbiol. Rev. 59, 1–30 (1995).
Gruber, T. M. & Gross, C. A. Multiple sigma subunits and the partitioning of bacterial transcription space. Annu. Rev. Microbiol. 57, 441–466 (2003).
Nicolas, P. et al. Condition-dependent transcriptome reveals high-level regulatory architecture in Bacillus subtilis. Science 335, 1103–1106 (2012).
Feklístov, A., Sharon, B. D., Darst, S. A. & Gross, C. A. Bacterial sigma factors: a historical, structural, and genomic perspective. Annu. Rev. Microbiol. 68, 357–376 (2014).
Veening, J. W. et al. Bet-hedging and epigenetic inheritance in bacterial cell development. Proc. Natl Acad. Sci. USA 105, 4393–4398 (2008).
Marlow, V. L. et al. The prevalence and origin of exoprotease-producing cells in the Bacillus subtilis biofilm. Microbiology 160, 56–66 (2014).
Vilain, S., Luo, Y., Hildreth, M. B. & Brozel, V. S. Analysis of the life cycle of the soil saprophyte Bacillus cereus in liquid soil extract and in soil. Appl. Environ. Microbiol. 72, 4970–4977 (2006).
Otto, A. et al. Systems-wide temporal proteomic profiling in glucose-starved Bacillus subtilis. Nat. Commun. 1, 137 (2010).
Omony, J., de Jong, A., Krawczyk, A. O., Eijlander, R. T. & Kuipers, O. P. Dynamic sporulation gene co-expression networks for Bacillus subtilis 168 and the food-borne isolate Bacillus amyloliquefaciens: a transcriptomic model. Microb. Genom. 4, 1–13 (2018).
Bendall, S. C. et al. Single-cell trajectory detection uncovers progression and regulatory coordination in human B cell development. Cell 157, 714–725 (2014).
Anavy, L. et al. BLIND ordering of large-scale transcriptomic developmental timecourses. Development 141, 1161–1166 (2014).
Haghverdi, L., Büttner, M., Wolf, F. A., Buettner, F. & Theis, F. J. Diffusion pseudotime robustly reconstructs lineage branching. Nat. Methods 13, 845–848 (2016).
Levin, M. et al. The mid-developmental transition and the evolution of animal body plans. Nature 531, 637–641 (2016).
Setty, M. et al. Wishbone identifies bifurcating developmental trajectories from single-cell data. Nat. Biotechnol. 34, 637–645 (2016).
Marioni, J. C. & Arendt, D. How single-cell genomics is changing evolutionary and developmental biology. Annu. Rev. Cell Dev. Biol. 33, 537–553 (2017).
Garrity, L. F. & Ordal, G. W. Chemotaxis in Bacillus subtilis: how bacteria monitor environmental signals. Pharmacol. Ther. 68, 87–104 (1995).
McKenney, P. T., Driks, A. & Eichenberger, P. The Bacillus subtilis endospore: assembly and functions of the multilayered coat. Nat. Rev. Microbiol. 11, 33–44 (2013).
Vlamakis, H., Chai, Y., Beauregard, P., Losick, R. & Kolter, R. Sticking together: building a biofilm the Bacillus subtilis way. Nat. Rev. Microbiol. 11, 157–168 (2013).
Kramer, M. A. Nonlinear principal component analysis using autoassociative neural networks. AIChE J. 37, 233–243 (1991).
Scholz, M., Fraunholz, M. & Selbig, J. in Principal Manifolds for Data Visualization and Dimension Reduction 44–67 (Springer, 2008).
Scholz, M. Validation of nonlinear PCA. Neural Process. Lett. 36, 21–30 (2012).
Verhamme, D. T., Kiley, T. B. & Stanley-Wall, N. R. DegU co-ordinates multicellular behaviour exhibited by Bacillus subtilis. Mol. Microbiol. 65, 554–568 (2007).
Fujita, M., González-Pastor, J. E. & Losick, R. High- and low-threshold genes in the Spo0A regulon of Bacillus subtilis. J. Bacteriol. 187, 1357–1368 (2005).
Verhamme, D. T., Murray, E. J. & Stanley-Wall, N. R. DegU and Spo0A jointly control transcription of two loci required for complex colony development by Bacillus subtilis. J. Bacteriol. 191, 100–108 (2009).
Branda, S. S., González-Pastor, J. E., Ben-Yehuda, S., Losick, R. & Kolter, R. Fruiting body formation by Bacillus subtilis. Proc. Natl Acad. Sci. USA 98, 11621–11626 (2001).
Branda, S. S., Chu, F., Kearns, D. B., Losick, R. & Kolter, R. A major protein component of the Bacillus subtilis biofilm matrix. Mol. Microbiol. 59, 1229–1238 (2006).
Galperin, M. Y. Genome diversity of spore-forming firmicutes. Microbiol. Spectr. 1, 1–15 (2013).
Kunst, F. et al. The complete genome sequence of the Gram-positive bacterium Bacillus subtilis. Nature 390, 249–256 (1997).
Barbe, V. et al. From a consortium sequence to a unified sequence: the Bacillus subtilis 168 reference genome a decade later. Microbiology 155, 1758–1775 (2009).
de Hoon, M. J. L., Eichenberger, P. & Vitkup, D. Hierarchical evolution of the bacterial sporulation network. Curr. Biol. 20, R735–R745 (2010).
Wolf, Y. I. & Koonin, E. V. A tight link between orthologs and bidirectional best hits in bacterial and archaeal genomes. Genome Biol. Evol. 4, 1286–1294 (2012).
Gabaldón, T. & Koonin, E. V. Functional and evolutionary implications of gene orthology. Nat. Rev. Genet. 14, 360–366 (2013).
Galperin, M. Y. et al. Genomic determinants of sporulation in Bacilli and Clostridia: towards the minimal set of sporulation-specific genes. Environ. Microbiol. 14, 2870–2890 (2012).
Abecasis, A. B. et al. A genomic signature and the identification of new sporulation genes. J. Bacteriol. 195, 2101–2115 (2013).
Moran, N. A. Microbial minimalism: genome reduction in bacterial pathogens. Cell 108, 583–586 (2002).
Makarova, K. et al. Comparative genomics of the lactic acid bacteria. Proc. Natl Acad. Sci. USA 103, 15611–15616 (2006).
Wolf, Y. I. & Koonin, E. V. Genome reduction as the dominant mode of evolution. BioEssays 35, 829–837 (2013).
Albalat, R. & Cañestro, C. Evolution by gene loss. Nat. Rev. Genet. 17, 379–391 (2016).
Duar, R. M. et al. Lifestyles in transition: evolution and natural history of the genus Lactobacillus. FEMS Microbiol. Rev. 41, S27–S48 (2017).
Sokurenko, E. V., Hasty, D. L. & Dykhuizen, D. E. Pathoadaptive mutations: gene loss and variation in bacterial pathogens. Trends Microbiol. 7, 191–195 (1999).
Maughan, H. et al. The population genetics of phenotypic deterioration in experimental populations of Bacillus subtilis. Evol. Int. J. Org. Evol. 60, 686–695 (2006).
Maughan, H., Masel, J., Birky, C. W. & Nicholson, W. L. The roles of mutation accumulation and selection in loss of sporulation in experimental populations of Bacillus subtilis. Genetics 177, 937–948 (2007).
Maughan, H., Birky, C. W. & Nicholson, W. L. Transcriptome divergence and the loss of plasticity in Bacillus subtilis after 6,000 generations of evolution under relaxed selection for sporulation. J. Bacteriol. 191, 428–433 (2009).
Brown, C. T. et al. Whole-genome sequencing and phenotypic analysis of Bacillus subtilis mutants following evolution under conditions of relaxed selection for sporulation. Appl. Environ. Microbiol. 77, 6867–6877 (2011).
Velicer, G. J. & Yu, Y. T. N. Evolution of novel cooperative swarming in the bacterium Myxococcus xanthus. Nature 425, 75–78 (2003).
van Ditmarsch, D. et al. Convergent evolution of hyperswarming leads to impaired biofilm formation in pathogenic bacteria. Cell Rep. 4, 697–708 (2013).
Song, C., Kidarsa, T. A., van de Mortel, J. E., Loper, J. E. & Raaijmakers, J. M. Living on the edge: emergence of spontaneous gac mutations in Pseudomonas protegens during swarming motility. Environ. Microbiol. 18, 3453–3465 (2016).
Friedlander, T., Mayo, A. E., Tlusty, T. & Alon, U. Evolution of bow-tie architectures in biology. PLoS Comput. Biol. 11, e1004055 (2015).
Yan, J. et al. Bow-tie signaling in c-di-GMP: machine learning in a simple biochemical network. PLoS Comput. Biol. 13, e1005677 (2017).
Kitano, H. Biological robustness. Nat. Rev. Genet. 5, 826–837 (2004).
Bonner, J. T. The Evolution of Complexity by Means of Natural Selection (Princeton Univ. Press, 1988).
Kashtan, N. & Alon, U. Spontaneous evolution of modularity and network motifs. Proc. Natl Acad. Sci. USA 102, 13773–13778 (2005).
Wagner, G. P., Pavlicev, M. & Cheverud, J. M. The road to modularity. Nat. Rev. Genet. 8, 921–931 (2007).
Espinosa-Soto, C. & Wagner, A. Specialization can drive the evolution of modularity. PLoS Comput. Biol. 6, e1000719 (2010).
Lande, R. Evolution of phenotypic plasticity and environmental tolerance of a labile quantitative character in a fluctuating environment. J. Evol. Biol. 27, 866–875 (2014).
Huang, B. & Mackem, S. Evolutionary developmental biology: use it or lose it. Nature 511, 34–35 (2014).
Siljestam, M. & Östman, Ö. The combined effects of temporal autocorrelation and the costs of plasticity on the evolution of plasticity. J. Evol. Biol. 30, 1361–1371 (2017).
Mandic-Mulec, I., Stefanic, P. & van Elsas, J. D. Ecology of Bacillaceae. Microbiol. Spectr. 3, TBS-0017–TBS-2013 (2015).
Parter, M., Kashtan, N. & Alon, U. Facilitated variation: how evolution learns from past environments to generalize to new environments. PLoS Comput. Biol. 4, e1000206 (2008).
Riedl, R. A systems-analytical approach to macro-evolutionary phenomena. Q. Rev. Biol. 52, 351–370 (1977).
Wagner, G. P. & Altenberg, L. Perspective: complex adaptations and the evolution of evolvability. Evolution 50, 967–976 (1996).
Schluter, D. Adaptive radiation along genetic lines of least resistance. Evolution 50, 1766–1774 (1996).
Watson, R. A. & Szathmáry, E. How can evolution learn? Trends Ecol. Evol. 31, 147–157 (2016).
Uller, T., Moczek, A. P., Watson, R. A., Brakefield, P. M. & Laland, K. N. Developmental bias and evolution: a regulatory network perspective. Genetics 209, 949–966 (2018).
Mead, D. A. et al. Complete genome sequence of Paenibacillus strain Y4.12MC10, a novel Paenibacillus lautus strain isolated from Obsidian hot spring in Yellowstone National Park. Stand. Genom. Sci. 6, 381–400 (2012).
van Nimwegen, E. in Power Laws, Scale-Free Networks and Genome Biology 236–253 (Springer, 2006).
Cordero, O. X. & Hogeweg, P. Large changes in regulome size herald the main prokaryotic lineages. Trends Genet. 23, 488–493 (2007).
Barka, E. A. et al. Taxonomy, physiology, and natural products of Actinobacteria. Microbiol. Mol. Biol. Rev. 80, 1–43 (2016).
Fall, R., Kearns, D. B. & Nguyen, T. A defined medium to investigate sliding motility in a Bacillus subtilis flagella-less mutant. BMC Microbiol. 6, 31 (2006).
Scholz, M. & Fraunholz, M. J. A computational model of gene expression reveals early transcriptional events at the subtelomeric regions of the malaria parasite, Plasmodium falciparum. Genome Biol. 9, R88 (2008).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Wu, D. et al. A phylogeny-driven genomic encyclopaedia of Bacteria and Archaea. Nature 462, 1056–1060 (2009).
Wu, D., Jospin, G. & Eisen, J. A. Systematic identification of gene families for use as ‘markers’ for phylogenetic and phylogeny-driven ecological studies of Bacteria and Archaea and their major subgroups. PLoS ONE 8, e77033 (2013).
Eddy, S. R. Hidden markov models. Curr. Opin. Struct. Biol. 6, 361–365 (1996).
Finn, R. D., Clements, J. & Eddy, S. R. HMMER web server: interactive sequence similarity searching. Nucleic Acids Res. 39, W29–W37 (2011).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Castresana, J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol. Biol. Evol. 17, 540–552 (2000).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Löytynoja, A. & Goldman, N. An algorithm for progressive multiple alignment of sequences with insertions. Proc. Natl Acad. Sci. USA 102, 10557–10562 (2005).
Löytynoja, A. & Goldman, N. Phylogeny-aware gap placement prevents errors in sequence alignment and evolutionary analysis. Science 320, 1632–1635 (2008).
Talavera, G. & Castresana, J. Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst. Biol. 56, 564–577 (2007).
Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. TrimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Huerta-Cepas, J., Dopazo, J. & Gabaldón, T. ETE: a python environment for tree exploration. BMC Bioinformatics 11, 24 (2010).
Huerta-Cepas, J., Serra, F. & Bork, P. ETE 3: reconstruction, analysis, and visualization of phylogenomic data. Mol. Biol. Evol. 33, 1635–1638 (2016).
Cock, P. J. A. et al. Biopython: freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics 25, 1422–1423 (2009).
Popescu, A. A., Huber, K. T. & Paradis, E. Ape 3.0: new tools for distance-based phylogenetics and evolutionary analysis in R. Bioinformatics 28, 1536–1537 (2012).
Revell, L. J. Phytools: an R package for phylogenetic comparative biology (and other things). Methods Ecol. Evol. 3, 217–223 (2011).
Pateiro-López, B. & Rodrıguez-Casal, A. Generalizing the convex hull of a sample: the R package alphahull. J. Stat. Softw. 34, 1–28 (2010).
VanDerWal, J., Falconi, L., Januchowski, S., Shoo, L. & Storlie, C. SDMTools: tools for processing data associated with species distribution modelling exercises. R version 1 (2014); https://cran.r-project.org/web/packages/SDMTools/index.html
Adler, D. et al. Rgl: 3D visualization using OpenGL. R version 095 (2016); https://cran.r-project.org/web/packages/rgl/index.html
Ross, K. F. A. & Billing, E. The water and solid content of living bacterial spores and vegetative cells as indicated by refractive index measurements. Microbiology 16, 418–425 (1957).
Lee, J. A. et al. MIFlowCyt: the minimum information about a flow cytometry experiment. Cytometry A 73, 926–930 (2008).
Spidlen, J., Breuer, K., Rosenberg, C., Kotecha, N. & Brinkman, R. R. FlowRepository: a resource of annotated flow cytometry datasets associated with peer-reviewed publications. Cytometry A 81, 727–731 (2012).
Barrick, J. E. et al. Identifying structural variation in haploid microbial genomes from short-read resequencing data using breseq. BMC Genom. 15, 1039 (2014).
Deatherage, D. E. & Barrick, J. E. in Engineering and Analyzing Multicellular Systems: Methods and Protocols (eds Sun, L. & Shou, W.) 165–188 (Springer, 2014).
Zhu, B. & Stülke, J. SubtiWiki in 2018: from genes and proteins to functional network annotation of the model organism Bacillus subtilis. Nucleic Acids Res. 46, D743–D748 (2018).
Acknowledgements
J.v.G. thanks M. Olombrada, M. Toll-Riera, N. Lyons and J. Payne for discussions. We thank R. Kolter for providing strains. J.v.G. thanks the Wierenga Rengerink PhD Prize from the University of Groningen, Rubicon Fellowship (No. 2015-2) from the Netherlands Organisation for Scientific Research (NWO), EMBO long-term fellowship (ALTF, No. 1101-2016), Marie Sklodowska-Curie Individual Fellowship (No. 742235), Swiss Federal Institute of Aquatic Science and Technology (Eawag) and ETH Zürich for financial support. A.W. acknowledges support by ERC Advanced Grant No. 739874, Swiss National Science Foundation grant No. 31003A_172887 as well as by the University Priority Research Program in Evolutionary Biology at the University of Zurich. M.A. was supported by Swiss National Science Foundation grant No. 31003A_169978, Eawag and ETH Zürich.
Author information
Authors and Affiliations
Contributions
J.v.G. conceived the project. All authors were involved in designing the project, planning the research and interpreting the results. J.v.G. conducted the research, created the figures and wrote a first version of the manuscript. M.A. and A.W. contributed to subsequent versions of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Text 1–4, Supplementary Figs. 1–34, Supplementary Tables 4, 7–11.
Supplementary Table 1
Reconstructions of global transcription network of Bacillus subtilis
Supplementary Table 2
List of expression profiles included from study of Nicolas and colleagues
Supplementary Table 3
Gene regulatory network underlying lifestyle switches
Supplementary Table 5
List of genomes included in phylogenetic analysis
Supplementary Table 6
Bi-directional best BLAST hits for all genomes with respect to reference genome
Rights and permissions
About this article
Cite this article
van Gestel, J., Ackermann, M. & Wagner, A. Microbial life cycles link global modularity in regulation to mosaic evolution. Nat Ecol Evol 3, 1184–1196 (2019). https://doi.org/10.1038/s41559-019-0939-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-019-0939-6
This article is cited by
-
Adaptation and phenotypic diversification of Bacillus thuringiensis biofilm are accompanied by fuzzy spreader morphotypes
npj Biofilms and Microbiomes (2022)
-
Mutability of demographic noise in microbial range expansions
The ISME Journal (2021)
-
Modularity of the life cycle
Nature Ecology & Evolution (2019)