Abstract
Species distributions and abundances are undergoing rapid changes worldwide. This highlights the significance of reliable, integrated information for guiding and assessing actions and policies aimed at managing and sustaining the many functions and benefits of species. Here we synthesize the types of data and approaches that are required to achieve such an integration and conceptualize ‘essential biodiversity variables’ (EBVs) for a unified global capture of species populations in space and time. The inherent heterogeneity and sparseness of raw biodiversity data are overcome by the use of models and remotely sensed covariates to inform predictions that are contiguous in space and time and global in extent. We define the species population EBVs as a space–time–species–gram (cube) that simultaneously addresses the distribution or abundance of multiple species, with its resolution adjusted to represent available evidence and acceptable levels of uncertainty. This essential information enables the monitoring of single or aggregate spatial or taxonomic units at scales relevant to research and decision-making. When combined with ancillary environmental or species data, this fundamental species population information directly underpins a range of biodiversity and ecosystem function indicators. The unified concept we present links disparate data to downstream uses and informs a vision for species population monitoring in which data collection is closely integrated with models and infrastructure to support effective biodiversity assessment.
Similar content being viewed by others
Main
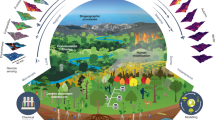
Biodiversity and the many ecosystem functions and services it underpins are undergoing significant and often rapid changes worldwide1. A range of global initiatives and policy frameworks, including the Convention on Biological Diversity (CBD) and Sustainable Development Goals (SDGs), have aimed to reduce this change and to halt the loss of biodiversity, with limited progress to date2. Appropriately gauging the impact of such policies or the progress toward international biodiversity goals has a key requirement: the availability of information on the status and trends of biodiversity in a form that is easily understood, timely, scientifically rigorous, standardized, relevant, global and representative of species populations across taxa and regions over time. Such information is particularly crucial in assessments, such as those carried out by the Intergovernmental Science–Policy Platform on Biodiversity and Ecosystem Services (IPBES)3, and is needed to construct ‘indicators’, which are aggregate measures that often address specific conservation targets4,5. Underpinning such metrics are core, essential measurements known as EBVs, which capture key constituent components of biodiversity change6,7, akin and complementary to the ‘essential climate variables’ supporting climate change assessment and policy8. Facilitated by the Group on Earth Observations Biodiversity Observation Network (GEO BON, http://geobon.org) and related efforts, the biodiversity science and observation community is now engaging in an effort to conceptualize and formulate these essential biodiversity components to enable more focused, integrated, and effective biodiversity monitoring in support of assessment and policy within a unified framework. This study represents the formal outcome of a process undertaken from 2015 through 2018 by the founding members of the GEO BON Species Populations Working Group9, which includes the authors of this Perspective, charged with providing the formal definitions, conceptualizations and recommendations addressing species distribution and abundance EBVs.
Changes in species distribution and abundance affect all biodiversity facets10, including the loss of potentially significant traits and functions1,11 and associated ecosystem consequences12,13. Patterns of spatial distribution and changes to these patterns inform us about the commonness, rarity and potential extinction risk for species14,15,16, determine the national and regional stewardship of species and are key to ensuring effective monitoring17, protection18,19 and population connectivity20 of species. Species-conservation goals often are particularly relevant to conservation legislation, and species population information used in tracking progress for CBD 2020 Targets 5, 11, 12 and 19 and SDG Goals 14 and 15, among others. When linked to data on surrounding conditions, occurrence information may provide insight into the realized environmental niche spaces of species21, which is key to capturing future consequences of global change22,23. Finally, species distribution and abundance have a range of other applications in science and society and facilitate app-based biodiversity discovery, learning and citizen science24,25,26,27.
Many countries already support a range of survey activities, such as national atlases, monitoring programs focused on threatened or flagship species and large-scale sharing of biodiversity data28,29,30. Conservation organizations and researchers add to this effort, but usually with a focus on a particular region or set of species. Critically, however, while often successful in addressing specific jurisdictional, organizational or scientific agendas, this data collection is naturally taxonomically, spatially, temporally and ecologically limited, biased and unrepresentative of overall biodiversity31,32,33,34, skewing the decisions that the data inform35. Sound measurement of progress toward policy targets and effective decision-making requires an information base geared toward overcoming these limitations through data and metadata capture and organization26,36,37,38 as well as harmonization and integration26,39,40. With species populations unconstrained by national borders and measurement of conservation progress requiring comparable and complete information, this integration should not only be explicit about its biodiversity representation but truly standardized and global in nature.
Here we characterize the elements of a capture of species population information that spans the entire Earth system and introduce the conceptual framework for the two ‘species population EBVs’ (SP EBVs) applicable to terrestrial, fresh water and marine environments. Specifically, we provide operational definitions for the species abundance EBV (SA EBV), which addresses counts of individuals for a given location in space and time, and the species distribution EBV (SD EBV), which is conceptually similar to the SA EBV but is simplified to a binary form and is usually more attainable. To address global policy and decision requirements, these EBVs need to fulfil four key criteria: (1) cover an explicit and, for a given taxonomic scope, maximally representative set of species; (2) have a near-global scope or, at minimum, address a given taxonomic scope to its full spatial extent so that national stewardship responsibilities are captured; (3) be geographically and temporally contiguous or maximally representative and (4) offer information at spatial and temporal resolutions that are useful for decision-makers and policy creation. As raw data alone is usually unable to fulfil these criteria, model-based and covariate-supported data integration is vital. We argue that the combination of global-scale remote sensing, new modelling methodology and novel computational and informatics solutions along with different species population data types now enable the required characterization.
Species occurrence data
The dynamics of species distribution and abundance are effectively assessed, and data to inform them fruitfully characterized, along the axes of space, time and taxonomic diversity41,42. Along these dimensions, an array of different data types from vastly varying sources contributes information about the occurrence of species. These usually vary by their spatial and temporal extent, resolution and frequency and are of different value in characterizing species distributions and their changes26. Currently, species distributions and the SD EBV are much more readily addressed than abundances (SA EBV); to introduce the concept, we first focus on species occurrence data. In its basic form, spatiotemporal species occurrence information requires at least a binary distinction of presence (≥1 individual) and/or non-detection (0 individuals). While all data types can provide evidence of species presence, only some are informative about absence (Fig. 1). Yet, reliable absence information is important for ascertaining change, including local emigrations and extirpations (absence given prior presence) or immigrations and introductions (presence given prior absence). Effective information integration for both SP EBVs therefore first requires a synthesis of the absolute and complementary value of core data types. As we illustrate (Fig. 2), these different data types and sources combine to offer very heterogeneous occurrence evidence.
The three major data types that we describe (incidental records, inventories over small or large areas and expert synthesis maps) differ in spatiotemporal scope, in taxonomic scope, in the quality of the presence (+) and absence information (–) that they provide and in spatiotemporal specificity, which in turn determine the key characteristics that they can inform. For further use and legend for the ‘Output’ field, see Fig. 2.
Left, Consider the occurrence of species in three countries (A–C) over a defined time period that has been informed by multiple data types (1–5). To facilitate the integration, data are summarized over a contiguous grid (right) with cells of equal size that includes all data types and species of a group and their global extent. Cells sizes could vary geographically to reflect data density, and, for marine or freshwater ecosystems, cells may represent units of volume or linear or area space. The grid allows geographic space to be collapsed into one dimension with, for example, neighbouring cells arranged in roughly sequential order along the vertical axis and over time for graphical presentation. This species–space–time–gram enables the visual characterization of available data for the two dimensions (space and time) and multiple species. Cells can be spatially aggregated at governance scales, such as regions, countries or counties and, potentially, themselves be politically defined. In our example (circled numbers on both sides of the figure), an expert synthesis map (1) characterizes the static species ranges for the 20-year period and is deemed by experts to reliably separate presence (green) and absence (orange) for the chosen space–time grid. This is complemented by two large-area inventories (2), developed before the year 2000, indicating — with little spatiotemporal specificity — species presence in parts of Country C, and likely absence in all of Country A. Two sets of small-area inventories exist, including single-year visits (3) and multi-year atlas efforts (4), all providing spatiotemporally explicit evidence that may hold reliable absence information. Finally, multiple incidental records (5) provide presence data for select grid cells and years. Assuming, for now, a high suitability of small-scale inventories for grid-level absence inference, the raw occurrence data does enable the detection of local extinctions (i) or non-native occurrence or invasions (ii).
Incidental observations
These are single records that lack information about co-observed species, taxonomic scope and, usually, sampling protocol, such as most museum records and many citizen-science contributions. They are therefore unable to directly inform non-detections, yet can often offer species presences for detailed locations (hence the term ‘presence-only’ data). Thanks to increasing amateur data collection24,27, advance of animal tracking43,44, activities of aggregators like the Global Biodiversity Information Facility (GBIF) and the Ocean Biogeographic Information System (OBIS), existing protocols facilitating sharing and interoperability (DarwinCore37), and easy-to-use modelling tools45, this data type and its direct use have seen strong growth. However, major taxonomic, ecological and geographic biases in data availability still exist31,46, impeding straightforward interpretation.
Inventories
This data type differs in two key aspects — it has a defined taxonomic and spatiotemporal scope that is larger than that of a single observed individual or species (Fig. 1). Inventories may address all members of a taxon (for example, mammals or bryophytes) or a group defined in another way (for example, trees or phytoplankton above a certain size). This enables inference about non-detections — that is, members of the focal species group that were not recorded (hence the term ‘presence–absence’ data). Such non-detections can provide information about absences, but the reliability of such inference depends on the overall survey effort and effectiveness and sampling protocol. For small-area inventories addressing relatively immobile or readily detected organisms, such as vegetation plots or a short survey transect, a given effort may provide very spatiotemporally specific evidence and potentially reliable absences. For more mobile groups that are harder to detect, larger or longer-lasting survey campaigns, such as atlas efforts or sensor/trap-based surveys, may offer more reliable absence evidence, but at the cost of spatiotemporal specificity. Over much larger extents, such as countries, counties, islands or national parks, large-area inventories like multispecies ‘checklists’ or state-level databases for select species of national interest (for example, pests or non-natives) are often based on multiple data sources and protocols and are considered ‘summary inventories’38. Depending on effort and rigor, they may provide both reliable presence and absence information, but usually with limited spatial or temporal specificity. Despite the fundamental role of inventories for ascertaining potential absences, especially in the context of growing modelling methodology47, they have seen limited mobilization and integration with other data types. Key past causes for this include the lack of (1) data and metadata, (2) infrastructure to facilitate the capture and use of this information and (3) community-wide appreciation and incentives for data and metadata sharing. With a prototype inventory data standard (Humboldt Core38) and associated informatics tools at Map of Life and format extensions at GBIF and OBIS in place, future growth in the capture of effort and complete metadata looks promising.
Expert synthesis maps
These are binary or categorical distribution maps that are developed by species experts. They aim to separate coarsely occupied areas from those without species occurrence and typically cover a longer timeframe, often decades rather than years48,49. Similar to large-area inventories but addressing only a single species at a time, they usually summarize multiple sources and data types from multiple time points (with details and provenance usually not retained). These data are based on by-region expert determinations or are interpolated to inform occurrence boundaries that are often hand-drawn based on sources including taxonomic monographs, handbooks, large-scale field guides and conservation reports. Such expert predictions are now sometimes quantitatively supported by species distribution modelling50,51,52,53,54 with pixel suitability scores subsequently thresholded, masked or otherwise modified by the expert to exclude areas presumed to be unoccupied55. While both data and (human or machine-based) models underpinning such predictions are usually a ‘black box’, they can offer information vital to delineating the geographical scope for a species within which presence may be expected and outside of which absence is likely. This often implies broad temporal scope and thus limited opportunity for direct change inference and a lack of spatial detail: hand-drawn presence–absence boundaries may have substantial inaccuracies, often include substantial areas of false presence and sometimes also include false absences49,56,57. However, other data may allow validation of a reliable spatial grain for this data type to be used for presence–absence information, for example ca. 100 km for global bird expert maps49,58, and potentially finer in model-supported expert predictions55.
Model-based integration and prediction
The species population EBVs
As shown above and in our data ‘species–space–time–gram’ (Fig. 2), raw biodiversity data alone are usually able to characterize species distributions in space and time only in an idiosyncratic way and with limited sensitivity for detecting change. General reasons include sparse data and taxonomically, spatially and environmentally uneven coverage31; ecological, environmental or phenological variation in species detectability59; and highly heterogeneous spatial and temporal grains of available data. Thus, on their own, raw data fail some or all of our initially formulated four key criteria for SP EBVs. Status and trend metrics and indicators that are mere aggregates of raw data are likely to be biased and sub-optimal and potentially may mislead downstream inference. This can be addressed through the use of spatiotemporally contiguous environmental and other species-level covariates in a statistical framework (Fig. 3). Observations representing different data types collected over heterogeneous spatiotemporal scales are unified in a space–time analysis grid or, in aggregate form, a species–space–time cube in which cells represent a model-based measure of presence or abundance. Individual cells may refer to space of any dimensionality, including linear or three-dimensional in the case of aquatic habitats. Cell size would ideally represent the relevant scale of occurrence or change processes60 or, more operationally, be adaptively driven by available data, intended output and acceptable uncertainty (see below) and may thus vary by species group. Models integrating the respective data types and signals among species, locations and time then enable predictions of a cell’s suitability, presence probability, or abundance of a species at particular points in space and time, while including measurements of uncertainty. Applied contiguously over an extent encompassing the geographic ranges of species in the taxonomic scope, this enables assessment and monitoring of occupied areas and associated statistical signals of local, regional and global change (Fig. 3).
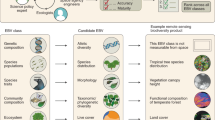
Bottom, Heterogeneous occurrence data, remotely sensed environmental conditions and ecological species attributes facilitate global predictions of species distributions in space and time. The SP EBVs are the predicted probability of occurrence (SD EBV) or the predicted number of individuals (SA EBV) over contiguous spatial and temporal units that cover the spatial extent of each species group member. For example, when data are aggregated for a single species, the SD EBV measures changes in distribution or population size. Centre, When data are aggregated for single cells, the SD EBV informs about community change in, for example, species richness or compositional similarity, or — via ancillary data — functional or phylogenetic turnover. Top, The fundamental contribution of SP EBVs for indicators of biodiversity and ecosystems status and trends. On their own, or combined with ancillary data on species or locations as well as EBVs from different classes, the SP EBVs underpin a large range of uses in policy, conservation, management, research and society. Inspired by earlier work42, the species–space–time–gram (cube) concept and graph were originally developed by the authors of the present work and then shared with the GEO BON community in 2016.
EBV definition
We thus define the essential biodiversity variable for species distributions (SD EBV) as the probability of occurrence over contiguous spatial and temporal units addressing the global extent of a species group consisting of one to many members. With support from models, this space–time–species cube is characterized for all members of a taxonomically or ecologically defined species group over their respective global extent, with a cell size that is potentially variable. The conceptually equivalent species abundance EBV (SA EBV) is the predicted count of individuals over contiguous spatial and temporal units. Together, they and their potential spatial aggregates or combinations with species attributes (see below) represent the species populations EBV class, as in ref. 6.
Environmental data
This conceptualization of the SP EBVs is uniquely facilitated by the environmental data revolution, specifically the availability of worldwide high-resolution remote sensing products. A number of sources, such as the National Aeronautics and Space Administration (NASA)’s Landsat, NASA’s moderate resolution imaging spectroradiometer (MODIS), NASA’s other upcoming missions61, the European Space Agency (ESA)’s Sentinel, and other (increasingly including commercial) ventures, provide data of growing spatiotemporal and spectral detail and extent and are increasingly representative of ecological drivers. The fine spatial and temporal resolution of data allows an environmental characterization of biodiversity data at spatiotemporal grains near that of in situ records. This enables an increasingly scale-conscious niche capture and provides critical spatiotemporal sensitivity and flexibility for inferring and predicting distributions and their change62,63. And these environmental measurements are increasingly relevant for species population processes, addressing biological drivers such as land cover, topographic and habitat heterogeneity, fine-scale weather variation, plant functional traits or productivity64,65,66,67,68 and, in select cases, even species directly63,69,70. Data across the depth-gradient for the oceans are more limited, but the first near-global distribution modeling71,72,73 and the first near-global characterizations of freshwater conditions are emerging74.
Models
The underlying modelling concepts and techniques supporting our approach are based broadly on species distribution models50,51,52,54, which identify the environmental conditions associated with species occurrence and allow their mapping in space. These conditions are usually identified inductively from occurrence data, but may also be informed deductively through species attributes, such as known ecological or physiological associations with land cover, elevation/depth and climatic or water conditions75,76,77. Dispersal and biotic constraints limit actual distributions, and for SP EBVs, predictions of the realized niche in the existing biological and spatial template is required, rather than the fundamental niche, which is more appropriate for projection into different time and space21. Expert-assessed, data-driven or phylogenetically inferred biotic species dependencies can be linked to hosts, prey, predators or other actors to inform distribution or abundance predictions78,79. These and landscape characteristics may also be gauged from the species co-occurrence data itself, for example, through a focus on assemblages and their compositional dissimilarity80 or a new generation of ‘joint’ distribution models for multiple species81,82. These methods are particularly promising for the strength of information gained across species in the face of limited occurrence data.
Almost all species distribution modelling applications to date use temporally static or collapsed input data only, even as they aim to project distributions changes in time (‘space for time substitution’52,83,84). This temporal stationarity assumption represents a key constraint for assessing temporal change85. With sufficient data, a more appropriate, yet somewhat inefficient, approach is to run separate models for different time periods. More desirable are models that are fit across the entire spatial and temporal scope of available data and that explicitly address spatiotemporal co-dependencies and signals of change86. This is addressed by dynamic distribution or dynamic occupancy models that set out to parameterize and predict variation in occurrence jointly in space and time82,87,88,89,90 as well as assemblage dissimilarity models applied to temporal turnover with no explicit consideration of species-level patterns or separation of spatial and temporal drivers91,92. The first large-scale demonstrations of dynamic occupancy approaches with spatiotemporal change assessment are now emerging93,94. We see great potential to extend and implement these methods as the backbone for addressing species occurrence in contiguous space and time.
Model-based data integration
The vast majority of species distribution prediction efforts to date are based on presence-only models using environmental covariates alone. This constrains delineation of non-environmental distribution limits, such as past or current physical or ecological barriers95,96. With sufficient data, this limitation can be addressed through hierarchical spatial models97 or alternatively by combining presences with an expert synthesis map data to restrict model predictions98. Presence-only models are limited in that they only provide relative cell suitability and use binary presence–absence thresholds contingent on ancillary information, expert judgment and assumptions99,100. We suggest that here the inclusion of inventory data will be critical and lead to a new generation of species distribution modelling approaches. Inventory data implicitly provide information on species absences and, through the use of an occupancy modelling framework40, enable the assessment of species- and environment-specific detection probabilities101,102 and a quantification of absolute occurrence probabilities103,104. While repeated sampling following a standardized protocol may be ideal, such data is obviously often limited to few or unrepresentative species and regions. We therefore argue that in many cases, alternative or even self-assessed information on the completeness of an inventory, and implicitly the level of detection or absence information afforded by it, may strengthen occurrence predictions38. We highlight the need for more statistical work to address this potential and acknowledge that species with extremely limited observational data will continue to pose a challenge, especially for trend and aggregate assemblage metrics. Combined with in situ abundance data or size estimates of home range, the same framework can address abundance105,106. Point process models in particular unify distribution and abundance predictions and seem especially suited to be a statistical framework39,107,108, in principle enabling smooth transition between the SD EBV and SA EBV.
A key challenge for model-based integration, exacerbating the known issue of gaps and biases in spatial biodiversity data, is the heterogeneity of taxa and sampling methods and of spatial and temporal grain of available data. Notably, presence and absence evidence have fundamentally different spatiotemporal grain properties. The presence of a single individual for a given place and time automatically implies the presence of the species for all larger spatial or temporal units that contain it. In contrast, an observed absence at the level of a small plot or short trawl does not imply absence at the level of an encompassing coarser-grain grid cell. Equally, for mobile organisms a reliable absence during a weeklong survey does not imply absence in that year. Hierarchical statistical models, often using Bayesian approaches, have been developed specifically to address the cross-type and cross-scale nature of occurrence data and used to combine inventory and incidental data109 or data from disparate spatial scales110. Such approaches enable predictions at a common spatial (and hypothetically temporal) resolution111, that is, the up- or downscaling of underlying heterogeneous data to a single spatiotemporal prediction grid for both species distribution and abundance112,113. The issue of scale is intimately connected to that of uncertainty, as occurrence at the continental or centennial scale is naturally fraught with less uncertainty than that at the 100-km or annual scale. For most uses, predictions at finer scales are preferred as long as uncertainty is captured, which is increasingly being facilitated by Bayesian and related modelling techniques114,115. We consider the capture, reporting, spatial visualization and cascading of uncertainty into aggregate products as key for supporting effective data collection and sound policy and management decisions. We note that the interconnections between the scale of process, evidence and predictions and the trade-offs between scale, uncertainty and sensitivity are key areas in need of further research.
Uses and applications
The envisioned essential species distribution information, or SD EBV, offers an exceptional breadth of applications in biodiversity and ecosystem monitoring and assessment (Fig. 3). Consider the idealized case of data and models providing annual occurrence probabilities and associated uncertainties for hundreds of species globally over a medium spatial grid sized at 100 km, 10 km or even 1 km. Such an empirically driven SD EBV enables the monitoring of species distribution dynamics (contractions, expansions and redistributions) and of the sizes and levels of fragmentation of geographic ranges. For any cell location, it provides information about community richness, composition and its change (thus addressing variables in the community EBV class sensu6), including immigration and loss of native species. When aggregated with data from regions or the globe, it offers compound characterizations of both species and community change in a larger-scale context and is thus able to directly inform global indicators of change, such as the suite of GEO-BON-endorsed biodiversity indicators116.
Ancillary data on species and places allow for enriched characterizations. The SD EBV joined with data on traits or functional roles of species may, for example, support inference about functional biodiversity losses10,117 or the potential ecological impacts of species invasions13. Combined with species-level estimates of life history and home range sizes, the SD EBV has the potential to support more temporally sensitive and accurate estimates of species extinction risk. Linking in spatial data on environmental change enables the identification of drivers of change. When combined with spatial protected area information, the SD EBV can support monitoring of progress for international biodiversity conservation targets or help identify new conservation opportunities, including in support of the Half-Earth Project118 or related aspirations. Extending the SD EBV to address abundance estimates for the same space–time–species cells, the SA EBV can offer even greater ecological and conservation relevance (for example, see refs. 94,119). Combined with ancillary data, such as species-typical body mass and function, it can inform biomass and biomass/abundance-scaled functional changes and more. Where attributes have high intraspecific variation, for example, due to local adaptation, these extended uses of SD EBV and SA EBV will benefit from in situ species-, community-, or even ecosystem-level measurements that can offer vital local detail.
For nations, the presented framework and associated infrastructure (see below) enable a substantially improved capacity to track biodiversity change and to assess progress toward national and international commitments35 by: (1) informing local predictions with globally pooled and integrated data and by involving a concerted expert effort for data input and validation and potentially offering more complete and rigorous status and trend estimates than an isolated national analysis or system, especially in undersampled regions; (2) capturing, through its global scope, various nations’ stewardship of species (for example, the proportion of a species’ global population that a given nation holds) and changes to it; (3) enabling indicators that, because of their global and standardized nature, inform progress toward internationally agreed biodiversity targets in a comparable way; and (4) guiding and providing infrastructure, tools and dashboards that support in-country and international assessment and reporting on species populations (facilitated by GEO BON and its partners), such as Map of Life, the BON-in-a-Box web toolkit, or the GEO BON EBV portal.
Infrastructure
The SP EBVs require rethinking traditional approaches to producing knowledge products. At the heart of the SP EBVs is the recognition of harmonization and model-based integration of multiple types of biodiversity and environmental data from heterogeneous sources that address different scales and are stored in multiple formats. This requires workflows that connect data to models in order to produce and disseminate credible and transparent modelled products. Such workflows necessitate a network of tools and infrastructure to address four main steps (Fig. 4): (1) data generation, contribution and aggregation, (2) data integration, (3) modelling and production of SP EBVs and (4) delivery and use of SP EBVs. In step 1, biodiversity data producers, such as taxon-, region- or data-type-specific networks, sampling campaigns (for example, through citizen science) or national/institutional interests (for example, country atlases and national marine surveys) are improving the sophistication of their database tools and data size. Unfortunately, much data still remain unavailable owing to lack of sharing or restrictive licensing, but data contributions are facilitated by a range of platforms (for a review of examples and associated standard and workflow issues, see ref. 36). For incidental records, data and metadata standards are enabling such data to effectively support EBV development. Key examples of such networks include GBIF and OBIS, which operate globally via connection to national nodes and thematic networks. Such global networks are important infrastructures to ensure availability, repeatability, standardization and archiving in support of downstream data integration and use120. However, still nascent and of immediate need is infrastructure that can play a similar role for more complex data types, such as select inventory data that require detailed metadata to most effectively feed into models, data-type or model-focused effort. Data integration and modelling (steps 2 and 3) are the backbone for producing SP EBVs. An informatics framework is needed that is strongly informed by research and is based on incorporation of environmental sensor data, flexible modelling methodology that integrates data types and scalable computational statistical approaches and associated cloud-based data management. The infrastructure should form a community-platform for the best-possible development of standardized, scientifically rigorous and transparent SP EBVs. This should include dashboards that enable taxon-region experts to provide community feedback and product improvement as well as data evaluation and product delivery back to their respective platforms and networks. Species distributions and data traverse national boundaries. We thus consider a harmonized global infrastructure that addresses core model-based integration steps as key to achieving standardized, geographically comparable information and downstream indicator products. Data integration and modelling steps and downstream products are otherwise likely to be near impossible to standardize.
a, Data contribution and validation. Networks composed of individuals, organization and institutions with an array of data types engage with platforms that support the standardized capture of data and metadata and provide tools for quality control and taxonomic harmonization. Infrastructure, such as GBIF and OBIS, are established as global examples of infrastructure in this space with the capacity to effectively store, improve and mobilize data onward for further use. b, Data integration. In this step, workflows and dashboards unify disparate data types and sources in a common spatiotemporal framework and supply networks of data providers, experts and other users with initial reporting on raw data coverage and trends in space and time. c, Modelling. After raw data is annotated with, for example, remotely-sensed environmental data, a default set of dynamic models, adapted to data type and quantity, is applied for predictions in space and (past to current) time, producing the EBV. Dashboards allow both taxon- and region-specific experts and modelling specialists to engage with, and iteratively improve, models and products. b and c, and parts of a concerning particularly model-relevant data and metadata, are enabled by infrastructure (such as Map of Life) that addresses global data integration, annotation, modelling and feedback to networks of experts. d, Use. A range of users, from policy, management, research, advocacy and the broader public, engages with summary maps or with products derived from the SP EBVs, such as indicators, which all can be connected with ancillary biodiversity or spatial data (Fig. 4). Further development of products and their communication is performed by scientific, international, national and non-governmental groups and associated platforms, including GEO BON, CBD and conservation organizations.
This approach places a premium on community involvement, and such a platform is meant to strongly support the needs of different communities to develop and curate their own SP EBVs products while enabling the best-possible transnational information products. In the absence of such a central resource, we foresee the potential for scattered and hard-to-replicate outputs of SP EBVs that are unable to be further integrated into the most usable synthesis products. We expect that this approach will not operate via a single actor or access point, but through coordinated efforts to develop shared methods and protocols, most likely using shared, cloud-based workflows. With support from a range of funding, science and technology partners, the core elements of this infrastructure are built or prototyped in Map of Life (http://mol.org/), which, with further development, we consider well placed to serve a key role in coordinating many components of data assembly and especially modelled outputs for the SD EBV. Finally (step 4), the visualization and programmatic delivery of basic SP EBV information and directly connected indicators should be well served through the infrastructure driving its production. But we envision a range of other national or global infrastructure, such as that associated with GEO BON, the CBD, conservation organizations, national agencies and others would similarly host programmatically accessed EBV information and/or combine with ancillary data to address community-specific needs or to build additional products.
Evidence base
Dataset information value
With billions of organisms on this planet that are changing in distribution and abundance at any moment, it is clearly impossible to fill the space–time–species cube in fine spatial, temporal and taxonomic resolution for all life and at the planetary scale. Even with the aid of ongoing environmental data collection and models, the operational grain for a given species group will be finer (and typical uncertainty levels will be lower) for those with many and well-stratified records in space and time (and environment) and high detection levels, such as birds, and coarser for those with more limited and/or spatially clustered data, such as ferns (Fig. 5a,b). This operational grain in turn is intimately connected to our ability to infer change — the coarser it is, the more constrained is our ability to infer species population trends, and particularly so for species groups with fine-scale spatiotemporal dynamics (Fig. 5c). Critically, even in the best-known species groups, viable spatiotemporal prediction grains may only arise in a proportion of species, usually an ecologically and spatially highly non-random subset103,121,122. Accordingly, taxonomic or functional representativeness of the predicted space–time–species cube may be high for some well- or easily-studied groups, but not others. Such limited taxonomic or functional coverage (representativeness) has strong repercussions for the relevance and generality of detected change (Fig. 5d). Consequently, the value of a dataset on a given species group is not determined by a single attribute, but is instead driven by its minimum ‘performance’ across a range of attributes (Fig. 6). We advocate for an expanded assessment of existing and planned data collections that accounts for this multitude of components and that provides a valuation of their ability to inform model-based predictions relative to other data.
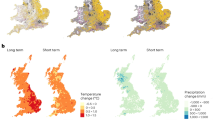
a, Prediction uncertainty is affected not just by record count but a suite of other issues and varies strongly among species groups and regions. b, Species groups with lower prediction uncertainty can support a finer spatiotemporal grain for EBV production. c, The magnitude of uncertainty or prediction grain in turn has direct effects on the potential for change inference, with exact effect depending on how readily such change is detectable for a species group at a given space–time grid, which will vary with the spatiotemporal scale of their distribution (for example, over the same spatiotemporal grain small-bodied versus large-bodied species groups will show different sensitivity). d, The generality and relevance of change inferences decreases quickly with any taxonomic or ecological biases in the data — for example, if some functional groups of a taxon are missing. US, United States; UK, United Kingdom; PNG, Papua New Guinea; the countries here are simply used to symbolize current perceived differences in available data, qualitatively informed by ref. 31.
Poor performance in a single criterion can result in limited overall value of a dataset for the purposes of global mapping and monitoring in this framework, and useful datasets would have at least medium performance in all criteria and would score high in as many as possible. This is highlighted with an initial characterization of some example datasets that could feed into EBV production (reflecting their status in the year 2018).
Effective data contributions
What then are the most valuable data contributions from individuals, organizations and government agencies in the context of these grain and coverage limitations? It follows from above that the value of additional data sampling and mobilization activities must be seen in the context of remote-sensing-aided models as well as other available data types and sources26,123. For charismatic, readily identified or common species in particular, citizen-science activities may now represent a relatively inexpensive, ongoing form of data collection, although highly geographically biased. Amateur-deployed visual or acoustic sensors or even environmental DNA data collection also increasingly contribute data at a relatively low cost124,125. In a more organized and standardized form, such as in many volunteer ‘atlas’ efforts or bird counts126, these sorts of citizen-scientist-supported activities hold the potential to reduce spatial and taxonomic data biases and, through capture of inventory information, absence inference. For select species and regions, conservation organizations may have a particular role in organizing and funding such sampling activities. Despite the vast potential this form of data collection holds for some taxa, for many species groups, taxon-region expert networks hosted by scientific societies and museums are key in providing guidance and quality control for amateur contributions or driving forward primary data collection as well as taxonomic assignment and harmonization.
National activities
Countries have a strong self-interest in effective biodiversity monitoring to maintain and sustainably use their own biological resources and to gauge their progress toward international obligations. To this end, many have set up national monitoring activities and are looking to the global observation and science community for facilitation and guidance17,30,33. While less so than bottom-up amateur efforts, most nationally organized data collection is still marked by heterogeneous methodology that usually fails to address detection probabilities, spatiotemporal biases and limited taxonomic coverage28,29, and sometimes scientific data sharing principles are not followed127,128. These activities also usually do not yet consider ancillary data and model-based inference in sampling design127,129. Some countries, such as Switzerland (http://atlas.vogelwarte.ch/), are leading the way in country-wide designs aimed at delivering highly effective data while not yet fully considering other citizen-science data collections and data from beyond their border, which both can improve prediction and inference123. For larger and more biodiverse countries that are less populated and have fewer resources, these sorts of campaigns are out of reach, highlighting the importance of national activities that are cost-effective and impactful.
For effective national contributions to essential species population information, we put forth a few key insights. (1) National monitoring efforts should be guided by existing biodiversity and remote sensing data collection and their model-based integration. The most (cost-)effective information gains may be achieved from strategically complementing past or ongoing data collection and taking advantage of statistical inference frameworks. (2) Country-level monitoring should be minimally isolated. Effective data collection relies on international coordination and collaboration and potentially cross-border support — species distributions span borders, and changes missed in one part of the range compromise inference for all countries that hold stewardship for a species. (3) Data should be published and open access. Only a full sharing of both data and metadata and their integration with other data in one place unlocks their full value and the potential for identifying data gaps and most strategic sampling opportunities. Such sharing can be incentivized by infrastructure that supports data discovery and measures data use and value in a global EBV context. (4) Support and guidance for bottom-up efforts may maximize cost-effectiveness. With strong and growing engagement of citizen scientists, some of the most effective gains for national monitoring may arise from supporting existing or nascent taxon-region specific networks and efforts and incentivizing them for more stratified sampling, full data and metadata sharing and engagement with experts for quality control and potentially greater taxonomic coverage. These may be rewarding areas of investment by governments and conservation organizations.
The necessary internationally integrated and science-driven approach to effective biodiversity monitoring systems at national and global scales holds an important role for GEO BON, which has this guidance and operationalization as its core mission. The SP EBVs framework enables the harmonization of data from an array of survey designs and technologies to deliver on the broad array of needs for biodiversity information, while avoiding potentially restrictive standards. GEO BON’s usage of the SP EBV framework and the associated research network and infrastructure and its engagement of National Biodiversity Observation Networks have the potential to make national monitoring activities more effective and to increase the value of their contributions to global biodiversity monitoring and knowledge advance.
Conclusions and recommendations
This conceptual and infrastructure framework for the production and use of essential information on species populations is intended to foster more effective and rigorous collection, communication and use of biodiversity data to support research, conservation, management and policy. By focusing attention on how, via standardized integration and modelling, different types of data from heterogeneous sources are improved in information value, the framework enables effective delivery of species population information for multiple management and policy objectives. Undoubtedly, the path toward a globally implemented and operational SD EBV (and even more so an SA EBV) will be long, and uncertainties or spatiotemporal grains may often impede inference. But the aspirational goal of a best-possible completion of the SP EBVs (Fig. 3) and the formulation of required elements and steps would herald a new and global phase of species population information collection, synthesis, and use. To attain this vision, we make the following key recommendations:
-
Enhanced data and metadata publication and sharing by countries, agencies, conservation organizations, research networks and individuals. Too much collected species occurrence-relevant data remains locked up, or its use is restricted by licensing, a key cause behind existing spatial biodiversity data gaps. We urge all to contribute to publishing and sharing mechanisms that inform SP EBVs globally, for example, via central aggregators GBIF and OBIS, via taxon-region specific efforts with unrestricted data and metadata sharing or via a direct link or publication to global SP EBV producing infrastructure such as Map of Life.
-
Collection, recognition and sharing of inventory data and detection-relevant metadata. We highlight inventories and associated non-detection information as key for effective species population inference. All too often, these data are simplified to presence records, or other detection-relevant information, for example, on taxonomic scope or observer qualification, is not being shared. We advocate for general infrastructure to support this capture.
-
Recognition of the relative, complementary value any primary biodiversity datasets have in the context of other biodiversity data, environmental data and models. This directly emerges from the SP EBV concept, and we suggest its use for designing monitoring efforts or incentivizing and supporting data collection.
-
Recognition of the role of agency-based and private remote sensing. Near-global remote sensing, in particular as conducted by United States’ NASA, the European Space Agency ESA and other space agencies that make their data freely available, is a key enabler of species population inference, empowering beneficiaries of biodiversity change information worldwide. This arena of impact and societal value deserves stronger recognition and support by agencies, governments and business.
-
Recognition of the role of science and research networks. The SP EBVs and their downstream uses only become unlocked through the engagement of scientists at the cutting edge of statistical methods development, big data integration and biodiversity informatics, strongly linked to relevant outcomes. By their nature, these activities are outside the scope of single agencies or conservation organizations that traditionally dominate biodiversity indicator and target discussions. Scientific projects based at academic institutions, research networks and organizations providing robust, policy-relevant science’ such as GEO BON and Future Earth, play a key role here by connecting monitoring and research with policy. This includes model development and infrastructure for the production of SP EBVs and filling of the space–time–species cube (Fig. 3). Given the complex and rapidly evolving informatics and modelling methodology, this task is best suited for development and hosting by academia and associated research networks.
-
Funding support for taxon-region expert and amateur networks, methods development and integration infrastructure. All three components are vital for an effective compilation and use of SP EBVs. Yet, with few exceptions, science funding agencies, conservation organization and foundations tend to be unwilling or unable to support such basic research and development activities. We encourage recognition of the societal benefits for all that a pooling of resources or dedicated engagement by some would enable.
References
Cardinale, B. J. et al. Biodiversity loss and its impact on humanity. Nature 486, 59–67 (2012).
Tittensor, D. P. et al. A mid-term analysis of progress toward international biodiversity targets. Science 346, 241–244 (2014).
Larigauderie, A. & Mooney, H. A. The Intergovernmental science-policy Platform on Biodiversity and Ecosystem Services: moving a step closer to an IPCC-like mechanism for biodiversity. Curr. Opin. Environ. Sustain. 2, 9–14 (2010).
Mace, G. M. & Baillie, J. E. M. The 2010 biodiversity indicators: challenges for science and policy. Conserv. Biol. 21, 1406–1413 (2007).
Noss, R. F. Indicators for monitoring biodiversity: a hierarchical approach. Conserv. Biol. 4, 355–364 (1990).
Pereira, H. M. et al. Ecology. Essential biodiversity variables. Science 339, 277–278 (2013).
Navarro, L. M. et al. Monitoring biodiversity change through effective global coordination. Curr. Opin. Environ. Sustain. 29, 158–169 (2017).
Bojinski, S. et al. The concept of essential climate variables in support of climate research, applications, and policy. Bull. Am. Meteorol. Soc. 95, 1431–1443 (2014).
Species populations. GEO BON Group on Earth Observations (GEO BON, accessed 12 July 2018); https://geobon.org/ebvs/working-groups/species-populations
Jarzyna, M. A. & Jetz, W. A near half-century of temporal change in different facets of avian diversity. Glob. Change Biol. 23, 2999–3011 (2017).
Keil, P., Storch, D. & Jetz, W. On the decline of biodiversity due to area loss. Nat. Commun. 6, 8837 (2015).
Simberloff, D. et al. Impacts of biological invasions: what’s what and the way forward. Trends Ecol. Evol. 28, 58–66 (2013).
Latombe, G. et al. A vision for global monitoring of biological invasions. Biol. Conserv. https://doi.org/10.1016/j.biocon.2016.06.013 (2016).
Mace, G. M. et al. Quantification of extinction risk: IUCN’s system for classifying threatened species. Conserv. Biol. 22, 1424–1442 (2008).
Cardillo, M. et al. The predictability of extinction: biological and external correlates of decline in mammals. Proc. Biol. Sci. 275, 1441–1448 (2008).
McGeoch, M. A. & Latombe, G. Characterizing common and range expanding species. J. Biogeogr. 43, 217–228 (2016).
Schmeller, D. S., Evans, D., Lin, Y.-P. & Henle, K. The national responsibility approach to setting conservation priorities—recommendations for its use. J. Nat. Conserv. 22, 349–357 (2014).
Moilanen, A., Wilson, K.A. & Possingham, H. Spatial conservation prioritization: quantitative methods and computational tools. (Oxford University Press, 2009).
Pollock, L. J., Thuiller, W. & Jetz, W. Large conservation gains possible for global biodiversity facets. Nature 546, 141–144 (2017).
Mazaris, A. D. et al. Evaluating the connectivity of a protected areas’ network under the prism of global change: the efficiency of the European Natura 2000 network for four birds of prey. PLoS One 8, e59640 (2013).
Soberón, J. & Nakamura, M. Niches and distributional areas: concepts, methods, and assumptions. Proc. Natl Acad. Sci. USA 106(Suppl 2), 19644–19650 (2009).
La Sorte, F. A. & Jetz, W. Tracking of climatic niche boundaries under recent climate change. J. Anim. Ecol. 81, 914–925 (2012).
Tingley, M. W., Monahan, W. B., Beissinger, S. R. & Moritz, C. Birds track their Grinnellian niche through a century of climate change. Proc. Natl Acad. Sci. USA 106(Suppl 2), 19637–19643 (2009).
Sullivan, B. L. et al. eBird: A citizen-based bird observation network in the biological sciences. Biol. Conserv. 142, 2282–2292 (2009).
Goldsmith, G. R. The field guide, rebooted. Science 349, 594 (2015).
Jetz, W., McPherson, J. M. & Guralnick, R. P. Integrating biodiversity distribution knowledge: toward a global map of life. Trends Ecol. Evol. 27, 151–159 (2012).
Dickinson, J. L. et al. The current state of citizen science as a tool for ecological research and public engagement. Front. Ecol. Environ. 10, 291–297 (2012).
Marsh, D. M. & Trenham, P. C. Current trends in plant and animal population monitoring. Conserv. Biol. 22, 647–655 (2008).
Schmeller, D. S., Henle, K., Loyau, A., Besnard, A. & Henry, P.-Y. Bird-monitoring in Europe - a first overview of practices, motivations and aims. Nat. Conserv. 2, 41–57 (2012).
Pereira, H.M. et al. in The GEO Handbook on Biodiversity Observation Networks 79–105 (Springer, 2017).
Meyer, C., Kreft, H., Guralnick, R. & Jetz, W. Global priorities for an effective information basis of biodiversity distributions. Nat. Commun. 6, 8221 (2015).
Proença, V. et al. Global biodiversity monitoring: from data sources to Essential Biodiversity Variables. Biol. Conserv. 213, 253–263 (2016).
Turak, E. et al. Essential Biodiversity Variables for measuring change in global freshwater biodiversity. Biol. Conserv. 213, 272–279 (2017).
Hortal, J. et al. Seven shortfalls that beset large-scale knowledge of biodiversity. Annu. Rev. Ecol. Evol. Syst. 46, 523–549 (2015).
Geijzendorffer, I. R. et al. Bridging the gap between biodiversity data and policy reporting needs: an essential biodiversity variables perspective. J. Appl. Ecol. 53, 1341–1350 (2015).
Kissling, W. D. et al. Building essential biodiversity variables (EBVs) of species distribution and abundance at a global scale. Biol. Rev. Camb. Philos. Soc. 93, 600–625 (2018).
Wieczorek, J. et al. Darwin Core: an evolving community-developed biodiversity data standard. PLoS One 7, e29715 (2012).
Guralnick, R., Walls, R. & Jetz, W. Humboldt Core – toward a standardized capture of biological inventories for biodiversity monitoring, modeling and assessment. Ecography 41, 713–725 (2017).
Williams, P. J. et al. An integrated data model to estimate spatiotemporal occupancy, abundance, and colonization dynamics. Ecology 98, 328–336 (2017).
MacKenzie, D. I. Occupancy Estimation and Modeling: Inferring Patterns and Dynamics of Species Occurrence, 1st edn. (Academic Press, 2005).
Andrewartha, H.G. & Birch, L.C. The Distribution and Abundance of Animals (Chicago University Press, 1954).
Bell, G. The interpretation of biological surveys. Proc. Biol. Sci. 270, 2531–2542 (2003).
Kays, R., Crofoot, M. C., Jetz, W. & Wikelski, M. ECOLOGY. Terrestrial animal tracking as an eye on life and planet. Science 348, aaa2478 (2015).
Hussey, N. E. et al. ECOLOGY. Aquatic animal telemetry: a panoramic window into the underwater world. Science 348, 1255642 (2015).
Phillips, S. J., Anderson, R. P. & Schapire, R. E. Maximum entropy modeling of species geographic distributions. Ecol. Modell. 190, 231–259 (2006).
Boakes, E. H. et al. Distorted views of biodiversity: spatial and temporal bias in species occurrence data. PLoS Biol. 8, e1000385 (2010).
Fithian, W., Elith, J., Hastie, T. & Keith, D. A. Bias correction in species distribution models: pooling survey and collection data for multiple species. Methods Ecol. Evol. 6, 424–438 (2015).
Gaston, K. J. & Fuller, R. A. The sizes of species’ geographic ranges. J. Appl. Ecol. 46, 1–9 (2009).
Hurlbert, A. H. & Jetz, W. Species richness, hotspots, and the scale dependence of range maps in ecology and conservation. Proc. Natl Acad. Sci. USA 104, 13384–13389 (2007).
Guisan, A. & Thuiller, W. Predicting species distribution: offering more than simple habitat models. Ecol. Lett. 8, 993–1009 (2005).
Franklin, J. & Miller, J.A. Mapping Species Distributions: Spatial Inference and Prediction Vol. 338 (Cambridge University Press Cambridge, 2009).
Elith, J. & Leathwick, J. R. Species distribution models: ecological explanation and prediction across space and time. Annu. Rev. Ecol. Evol. Syst. 40, 677–697 (2009).
Ready, J. et al. Predicting the distributions of marine organisms at the global scale. Ecol. Modell. 221, 467–478 (2010).
Peterson, A.T. et al. Ecological Niches and Geographic Distributions (Princeton University Press, 2011).
Ballesteros-Mejia, L., Kitching, I. J., Jetz, W. & Beck, J. Putting insects on the map: near-global variation in sphingid moth richness along spatial and environmental gradients. Ecography 40, 698–708 (2017).
Hurlbert, A. H. & White, E. P. Disparity between range map- and survey-based analyses of species richness: patterns, processes and implications. Ecol. Lett. 8, 319–327 (2005).
Ferrier, S. Mapping spatial pattern in biodiversity for regional conservation planning: where to from here? Syst. Biol. 51, 331–363 (2002).
Jetz, W., Sekercioglu, C. H. & Watson, J. E. M. Ecological correlates and conservation implications of overestimating species geographic ranges. Conserv. Biol. 22, 110–119 (2008).
MacKenzie, D. I. et al. Estimating site occupancy rates when detection probabilities are less than one. Ecology 83, 2248–2255 (2002).
Mertes, K. & Jetz, W. Disentangling scale dependencies in species environmental niches and distributions. Ecography 40, 1604–1615 (2017).
Stavros, E. N. et al. ISS observations offer insights into plant function. Nat. Ecol. Evo. 1, 0194 (2017).
Anderson, C. B. Biodiversity monitoring, earth observations and the ecology of scale. Ecol. Lett. 21, 1572–1585 (2018).
He, K. S. et al. Will remote sensing shape the next generation of species distribution models? Remote Sens. Ecol. Conserv. 1, 4–18 (2015).
Hansen, M. C. et al. High-resolution global maps of 21st-century forest cover change. Science 342, 850–853 (2013).
Wilson, A. M. & Jetz, W. Remotely sensed high-resolution global cloud dynamics for predicting ecosystem and biodiversity distributions. PLoS Biol. 14, e1002415 (2016).
Tuanmu, M.-N. & Jetz, W. A global, remote sensing-based characterization of terrestrial habitat heterogeneity for biodiversity and ecosystem modelling. Glob. Ecol. Biogeogr. 24, 1329–1339 (2015).
Muller-Karger, F. et al. Satellite remote sensing in support of an integrated ocean observing system. IEEE Geosci. Remote Sens. Mag. 1, 8–18 (2013).
Jetz, W. et al. Monitoring plant functional diversity from space. Nat. Plants 2, 16024 (2016).
Fretwell, P. T. & Trathan, P. N. Penguins from space: faecal stains reveal the location of emperor penguin colonies. Glob. Ecol. Biogeogr. 18, 543–552 (2009).
Asner, G. P. & Martin, R. E. Spectranomics: emerging science and conservation opportunities at the interface of biodiversity and remote sensing. Glob. Ecol. Conserv. 8, 212–219 (2016).
Basher, Z. & Costello, M. J. The past, present and future distribution of a deep-sea shrimp in the Southern Ocean. PeerJ 4, e1713 (2016).
Duffy, J. E. et al. Envisioning a marine biodiversity observation network. Bioscience 63, 350–361 (2013).
Muller-Karger, F. E. et al. Advancing marine biological observations and data requirements of the complementary essential ocean variables (EOVs) and essential biodiversity variables (EBVs) frameworks. Front. Mar. Sci. 5, 211 (2018).
Domisch, S., Amatulli, G. & Jetz, W. Near-global freshwater-specific environmental variables for biodiversity analyses in 1 km resolution. Sci. Data 2, 150073 (2015).
Kaschner, K., Watson, R., Trites, A. & Pauly, D. Mapping world-wide distributions of marine mammal species using a relative environmental suitability (RES) model. Mar. Ecol. Prog. Ser. 316, 285–310 (2006).
Tuanmu, M.-N. & Jetz, W. A global 1-km consensus land-cover product for biodiversity and ecosystem modelling. Glob. Ecol. Biogeogr. 23, 1031–1045 (2014).
Ficetola, G. F., Rondinini, C., Bonardi, A., Baisero, D. & Padoa-Schioppa, E. Habitat availability for amphibians and extinction threat: a global analysis. Divers. Distrib. 21, 302–311 (2014).
Urban, M. C., Zarnetske, P. L. & Skelly, D. K. Moving forward: dispersal and species interactions determine biotic responses to climate change. Ann. NY Acad. Sci. 1297, 44–60 (2013).
Wisz, M. S. et al. The role of biotic interactions in shaping distributions and realised assemblages of species: implications for species distribution modelling. Biol. Rev. Camb. Philos. Soc. 88, 15–30 (2013).
Ferrier, S. & Guisan, A. Spatial modelling of biodiversity at the community level. J. Appl. Ecol. 43, 393–404 (2006).
Pollock, L. J. et al. Understanding co-occurrence by modelling species simultaneously with a Joint Species Distribution Model (JSDM). Methods Ecol. Evol. 5, 397–406 (2014).
Thorson, J. T. et al. Joint dynamic species distribution models: a tool for community ordination and spatio‐temporal monitoring. Glob. Ecol. Biogeogr. 25, 1144–1158 (2016).
Kappes, H., Sundermann, A. & Haase, P. High spatial variability biases the space-for-time approach in environmental monitoring. Ecol. Indic. 10, 1202–1205 (2010).
La Sorte, F. A., Lee, T. M., Wilman, H. & Jetz, W. Disparities between observed and predicted impacts of climate change on winter bird assemblages. Proc. Biol. Sci. 276, 3167–3174 (2009).
Elith, J., Kearney, M. & Phillips, S. The art of modelling range-shifting species. Methods Ecol. Evol. 1, 330–342 (2010).
Ferrier, S., Jetz, W. & Scharlemann, J. in The GEO Handbook on Biodiversity Observation Networks 239–257 (Springer, 2017).
Franklin, J. Moving beyond static species distribution models in support of conservation biogeography. Divers. Distrib. 16, 321–330 (2010).
Royle, J. A. & Kéry, M. A Bayesian state-space formulation of dynamic occupancy models. Ecology 88, 1813–1823 (2007).
Merow, C., Lafleur, N., Silander, J. A. Jr., Wilson, A. M. & Rubega, M. Developing dynamic mechanistic species distribution models: predicting bird-mediated spread of invasive plants across northeastern North America. Am. Nat. 178, 30–43 (2011).
MacKenzie, D. I., Nichols, J. D., Seamans, M. E. & Gutiérrez, R. J. Modeling species occurrence dynamics with multiple states and imperfect detection. Ecology 90, 823–835 (2009).
Ferrier, S., Manion, G., Elith, J. & Richardson, K. Using generalized dissimilarity modelling to analyse and predict patterns of beta diversity in regional biodiversity assessment. Divers. Distrib. 13, 252–264 (2007).
Hoskins, A. J. et al. Supporting global biodiversity assessment through high-resolution macroecological modelling: Methodological underpinnings of the BILBI framework. Preprint at https://doi.org/10.1101/309377 (2018).
Rodhouse, T. J. et al. Establishing conservation baselines with dynamic distribution models for bat populations facing imminent decline. Divers. Distrib. 21, 1401–1413 (2015).
Thorson, J. T. & Barnett, L. A. Comparing estimates of abundance trends and distribution shifts using single-and multispecies models of fishes and biogenic habitat. ICES J. Mar. Sci. 74, 1311–1321 (2017).
Barve, N. et al. The crucial role of the accessible area in ecological niche modeling and species distribution modeling. Ecol. Modell. 222, 1810–1819 (2011).
Svenning, J.-C. & Skov, F. Ice age legacies in the geographical distribution of tree species richness in Europe. Glob. Ecol. Biogeogr. 16, 234–245 (2007).
Latimer, A. M., Banerjee, S., Sang, H. Jr., Mosher, E. S. & Silander, J. A. Jr. Hierarchical models facilitate spatial analysis of large data sets: a case study on invasive plant species in the northeastern United States. Ecol. Lett. 12, 144–154 (2009).
Merow, C., Wilson, A. M. & Jetz, W. Integrating occurrence data and expert maps for improved species range predictions. Glob. Ecol. Biogeogr. 26, 243–258 (2017).
Calabrese, J. M., Certain, G., Kraan, C. & Dormann, C. F. Stacking species distribution models and adjusting bias by linking them to macroecological models. Glob. Ecol. Biogeogr. 23, 99–112 (2014).
Jiménez-Valverde, A. & Lobo, J. M. Threshold criteria for conversion of probability of species presence to either–or presence–absence. Acta Oecol. 31, 361–369 (2007).
Dorazio, R. M., Royle, J. A., Söderström, B. & Glimskär, A. Estimating species richness and accumulation by modeling species occurrence and detectability. Ecology 87, 842–854 (2006).
Iknayan, K. J., Tingley, M. W., Furnas, B. J. & Beissinger, S. R. Detecting diversity: emerging methods to estimate species diversity. Trends Ecol. Evol. 29, 97–106 (2013).
Lahoz-Monfort, J. J., Guillera-Arroita, G. & Wintle, B. A. Imperfect detection impacts the performance of species distribution models. Glob. Ecol. Biogeogr. 23, 504–515 (2014).
Lobo, J. M., Jiménez-Valverde, A. & Hortal, J. The uncertain nature of absences and their importance in species distribution modelling. Ecography 33, 103–114 (2010).
Jones, J. P. G. Monitoring species abundance and distribution at the landscape scale. J. Appl. Ecol. 48, 9–13 (2011).
Royle, J. A. & Nichols, J. D. Estimating abundance from repeated presence-absence data or point counts. Ecology 84, 777–790 (2003).
Renner, I. W. & Warton, D. I. Equivalence of MAXENT and Poisson point process models for species distribution modeling in ecology. Biometrics 69, 274–281 (2013).
Koshkina, V. et al. Integrated species distribution models: combining presence-background data and site-occupany data with imperfect detection. Methods Ecol. Evol. 8, 420–430 (2017).
Dorazio, R. M. Accounting for imperfect detection and survey bias in statistical analysis of presence‐only data. Glob. Ecol. Biogeogr. 23, 1472–1484 (2014).
Keil, P., Belmaker, J., Wilson, A. M., Unitt, P. & Jetz, W. Downscaling of species distribution models: a hierarchical approach. Methods Ecol. Evol. 4, 82–94 (2013).
Keil, P., Wilson, A. M. & Jetz, W. Uncertainty, priors, autocorrelation and disparate data in downscaling of species distributions. Divers. Distrib. 20, 797–812 (2014).
Hui, C. et al. Extrapolating population size from the occupancy-abundance relationship and the scaling pattern of occupancy. Ecol. Appl. 19, 2038–2048 (2009).
Barwell, L. J., Azaele, S., Kunin, W. E. & Isaac, N. J. Can coarse‐grain patterns in insect atlas data predict local occupancy? Divers. Distrib. 20, 895–907 (2014).
Golding, N. & Purse, B. V. Fast and flexible Bayesian species distribution modelling using Gaussian processes. Methods Ecol. Evol. 7, 598–608 (2016).
Beale, C. M. & Lennon, J. J. Incorporating uncertainty in predictive species distribution modelling. Phil. Trans. R. Soc. Lond. B 367, 247–258 (2012).
Pereira, H. M., Freyhof, J., Ferrier, S. & Jetz, W. Global Biodiversity Change Indicators 1–18 (GEO Biodiversity Observation Network, Leipzig, Germany, 2015).
Schipper, A. M. et al. Contrasting changes in the abundance and diversity of North American bird assemblages from 1971 to 2010. Global Change Biol 22, 3948–3959 (2016).
Wilson, E.O. Half-Earth: Our Planet’s Fight for Life (WW Norton & Company, 2016).
Amano, T. et al. Successful conservation of global waterbird populations depends on effective governance. Nature 553, 199–202 (2017).
Edwards, J. L. Research and Societal Benefits of the Global Biodiversity Information Facility. Bioscience 54, 486–487 (2004).
Isaac, N. J. B. & Pocock, M. J. O. Bias and information in biological records. Biol. J. Linn. Soc. 115, 522–531 (2015).
Meyer, C., Jetz, W., Guralnick, R. P., Fritz, S. A. & Kreft, H. Range geometry and socio-economics dominate species-level biases in occurrence information. Glob. Ecol. Biogeogr. 25, 1181–1193 (2016).
Honrado, J. P., Pereira, H. M. & Guisan, A. Fostering integration between biodiversity monitoring and modelling. J. Appl. Ecol. 53, 1299–1304 (2016).
Bush, A. et al. Connecting Earth observation to high-throughput biodiversity data. Nat. Ecol. Evol. 1, 0176 (2017).
Steenweg, R. et al. Scaling‐up camera traps: monitoring the planet’s biodiversity with networks of remote sensors. Front. Ecol. Environ. 15, 26–34 (2016).
Robertson, M., Cumming, G. & Erasmus, B. Getting the most out of atlas data. Divers. Distrib. 16, 363–375 (2010).
Wilkinson, M. D. et al. The FAIR Guiding Principles for scientific data management and stewardship. Sci. Data 3, 160018 (2016).
Costello, M. J. & Wieczorek, J. Best practice for biodiversity data management and publication. Biol. Conserv. 173, 68–73 (2014).
Carvalho, S. B., Gonçalves, J., Guisan, A. & Honrado, J. P. Systematic site selection for multispecies monitoring networks. J. Appl. Ecol. 53, 1305–1316 (2016).
Acknowledgements
This study and its definitions and recommendations are the official, consensus outputs of the Species Populations Working Group of GEO BON (https://geobon.org/ebvs/working-groups/species-populations) and are the outcome of multiple workshops and group meetings in 2015 to 2018. We are grateful to the GEO BON Secretariat and the German Centre for Integrative Biodiversity Research (iDiv) for facilitation. We acknowledge NSF grants DBI-1262600 and DEB-1441737 and NASA Grants 80NSSC17K0282 and 80NSSC18K0435 to W.J. The research described in this paper was in part carried out at the Jet Propulsion Laboratory, California Institute of Technology, under contract with the NASA; government sponsorship is acknowledged.
Author information
Authors and Affiliations
Contributions
All authors contributed to the working group discussions leading to this manuscript and provided text contributions and/or figure feedback. W.J. wrote the first draft of the text, drafted Figs. 1–3 and 5 and 6 and coordinated the manuscript work. R.G. drafted Fig. 4 and led the informatics text section. M.A.M. provided key text and figure contributions and co-led the group discussions.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jetz, W., McGeoch, M.A., Guralnick, R. et al. Essential biodiversity variables for mapping and monitoring species populations. Nat Ecol Evol 3, 539–551 (2019). https://doi.org/10.1038/s41559-019-0826-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-019-0826-1
This article is cited by
-
Leveraging virtual datasets to investigate the interplay of pollinators, protected areas, and SDG 15
Sustainable Earth Reviews (2024)
-
Quantifying forest degradation requires a long-term, landscape-scale approach
Nature Ecology & Evolution (2024)
-
Gaps and opportunities in modelling human influence on species distributions in the Anthropocene
Nature Ecology & Evolution (2024)
-
Predicting species abundance using machine learning approach: a comparative assessment of random forest spatial variants and performance metrics
Modeling Earth Systems and Environment (2024)
-
Monitoring of Organochlorine Pesticides (OCP) in Hair Samples of Wild Herbivorous Mammals Living in Remote and Protected Areas of the Far East and Siberia of Russia
Bulletin of Environmental Contamination and Toxicology (2024)