Abstract
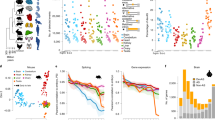
The mechanisms by which entire programmes of gene regulation emerged during evolution are poorly understood. Neuronal microexons represent the most conserved class of alternative splicing in vertebrates, and are critical for proper brain development and function. Here, we discover neural microexon programmes in non-vertebrate species and trace their origin to bilaterian ancestors through the emergence of a previously uncharacterized ‘enhancer of microexons’ (eMIC) protein domain. The eMIC domain originated as an alternative, neural-enriched splice isoform of the pan-eukaryotic Srrm2/SRm300 splicing factor gene, and subsequently became fixed in the vertebrate and neuronal-specific splicing regulator Srrm4/nSR100 and its paralogue Srrm3. Remarkably, the eMIC domain is necessary and sufficient for microexon splicing, and functions by interacting with the earliest components required for exon recognition. The emergence of a novel domain with restricted expression in the nervous system thus resulted in the evolution of splicing programmes that qualitatively expanded the neuronal molecular complexity in bilaterians.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Code availability
All of the software used to analyse the data is publicly available and listed in the Reporting Summary. VastDB files to run vast-tools are available to download for each species (https://github.com/vastgroup/vast-tools), as indicated in the Methods. Custom codes to generate orthologous gene clusters and figure plots are available upon request.
Data availability
Raw RNA-Seq data were submitted to the Sequence Read Archive (SRP149913). Mass spectrometry data were submitted to the ProteomeXchange Consortium: AP-MS through MassIVE (https://massive.ucsd.edu; accession codes: MSV000082361 and PXD009779) and RNP mass spectrometry via PRIDE59 (PXD010034). All other RNA-Seq datasets used in the study are listed in Supplementary Dataset 3.
References
Irimia, M. et al. A highly conserved program of neuronal microexons is misregulated in autistic brains. Cell 159, 1511–1523 (2014).
Li, Y. I., Sanchez-Pulido, L., Haerty, W. & Ponting, C. P. RBFOX and PTBP1 proteins regulate the alternative splicing of micro-exons in human brain transcripts. Genome Res. 25, 1–13 (2015).
Quesnel-Vallières, M., Irimia, M., Cordes, S. P. & Blencowe, B. J. Essential roles for the splicing regulator nSR100/SRRM4 during nervous system development. Genes Dev. 29, 746–759 (2015).
Nakano, Y. et al. A mutation in the Srrm4 gene causes alternative splicing defects and deafness in the Bronx waltzer mouse. PLoS Genet. 8, e1002966 (2012).
Quesnel-Vallieres, M. et al. Misregulation of an activity-dependent splicing network as a common mechanism underlying autism spectrum disorders. Mol. Cell 64, 1023–1034 (2016).
Calarco, J. A. et al. Regulation of vertebrate nervous system alternative splicing and development by an SR-related protein. Cell 138, 898–910 (2009).
Raj, B. et al. Global regulatory mechanism underlying the activation of an exon network required for neurogenesis. Mol. Cell 56, 90–103 (2014).
Parras, A. et al. Autism-like phenotype and risk gene mRNA deadenylation by CPEB4 mis-splicing. Nature 560, 441–446 (2018).
Gonatopoulos-Pournatzis, T. et al. Genome-wide CRISPR–Cas9 interrogation of splicing networks reveals a mechanism for recognition of autism-misregulated neuronal microexons. Mol. Cell 72, 510–524 (2018).
Putnam, N. et al. The amphioxus genome and the evolution of the chordate karyotype. Nature 453, 1064–1071 (2008).
Blencowe, B. J., Issner, R., Nickerson, J. A. & Sharp, P. A. A coactivator of pre-mRNA splicing. Genes Dev. 12, 996–1009 (1998).
Eldridge, A. G., Li, Y., Sharp, P. A. & Blencowe, B. J. The SRm160/300 splicing coactivator is required for exon-enhancer function. Proc. Natl Acad. Sci. USA 96, 6125–6130 (1999).
Khanna, M. et al. A systematic characterization of Cwc21, the yeast ortholog of the human spliceosomal protein SRm300. RNA 15, 2174–2185 (2009).
Grainger, R. J., Barrass, J. D., Jacquier, A., Rain, J. C. & Beggs, J. D. Physical and genetic interactions of yeast Cwc21p, an ortholog of human SRm300/SRRM2, suggest a role at the catalytic center of the spliceosome. RNA 15, 2161–2173 (2009).
Blencowe, B. J. et al. The SRm160/300 splicing coactivator subunits. RNA 6, 111–120 (2000).
Nakanishi, N., Renfer, E., Technau, U. & Rentzsch, F. Nervous systems of the sea anemone Nematostella vectensis are generated by ectoderm and endoderm and shaped by distinct mechanisms. Development 139, 347–357 (2012).
Wongpalee, S. P. et al. Large-scale remodeling of a repressed exon ribonucleoprotein to an exon definition complex active for splicing. eLife 5, e19743 (2016).
McKeown, A. N. et al. Evolution of DNA specificity in a transcription factor family produced a new gene regulatory module. Cell 159, 58–68 (2014).
Logeman, B. L., Wood, L. K., Lee, J. & Thiele, D. J. Gene duplication and neo-functionalization in the evolutionary and functional divergence of the metazoan copper transporters Ctr1 and Ctr2. J. Biol. Chem. 292, 11531–11546 (2017).
Arnegard, M. E., Zwickl, D. J., Lu, Y. & Zakon, H. H. Old gene duplication facilitates origin and diversification of an innovative communication system—twice. Proc. Natl Acad. Sci. USA 107, 22172–22177 (2010).
Wang, J. et al. LSD1n is an H4K20 demethylase regulating memory formation via transcriptional elongation control. Nat. Neurosci. 18, 1256–1264 (2015).
Matsushita, M., Yamamoto, R., Mitsui, K. & Kanazawa, H. Altered motor activity of alternative splice variants of the mammalian kinesin-3 protein KIF1B. Traffic 10, 1647–1654 (2009).
Tapial, J. et al. An atlas of alternative splicing profiles and functional associations reveals new regulatory programs and genes that simultaneously express multiple major isoforms. Genome Res. 27, 1759–1768 (2017).
Braunschweig, U. et al. Widespread intron retention in mammals functionally tunes transcriptomes. Genome Res. 24, 1774–1786 (2014).
Giampietro, C. et al. The alternative splicing factor Nova2 regulates vascular development and lumen formation. Nat. Commun. 6, 8479 (2015).
Gueroussov, S. et al. An alternative splicing event amplifies evolutionary differences between vertebrates. Science 349, 868–873 (2015).
Han, H. et al. MBNL proteins repress ES-cell-specific alternative splicing and reprogramming. Nature 498, 241–245 (2013).
Solana, J. et al. Conserved functional antagonism of CELF and MBNL proteins controls stem cell-specific alternative splicing in planarians. eLife 5, e16797 (2016).
Burguera, D. et al. Evolutionary recruitment of flexible Esrp-dependent splicing programs into diverse embryonic morphogenetic processes. Nat. Commun. 8, 1799 (2017).
Altenhoff, A. M., Gil, M., Gonnet, G. H. & Dessimoz, C. Inferring hierarchical orthologous groups from orthologous gene pairs. PLoS ONE 8, e53786 (2013).
Alexeyenko, A., Tamas, I., Liu, G. & Sonnhammer, E. L. Automatic clustering of orthologs and inparalogs shared by multiple proteomes. Bioinformatics 22, e9–e15 (2006).
Singh, P. P., Arora, J. & Isambert, H. Identification of ohnolog genes originating from whole genome duplication in early vertebrates, based on synteny comparison across multiple genomes. PLoS Comput. Biol. 11, e1004394 (2015).
Katoh, K., Kuma, K., Toh, H. & Miyata, T. MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Res. 33, 511–518 (2005).
Irimia, M. & Roy, S. W. Spliceosomal introns as tools for genomic and evolutionary analysis. Nucleic Acids Res. 36, 1703–1712 (2008).
Yeo, G. W. & Burge, C. B. Maximum entropy modeling of short sequence motifs with applications to RNA splicing signals. J. Comput. Biol. 11, 377–394 (2004).
Corvelo, A., Hallegger, M., Smith, C. W. & Eyras, E. Genome-wide association between branch point properties and alternative splicing. PLoS Comput. Biol. 6, e1001016 (2010).
Gohr, A. & Irimia, M. Matt: Unix tools for alternative splicing analysis. Bioinformatics 35, 130–132 (2019).
Gonzalez, M. et al. Generation of stable Drosophila cell lines using multicistronic vectors. Sci. Rep. 1, 75 (2011).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Huelsenbeck, J. P. & Ronquist, F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17, 754–755 (2001).
Müller, T. & Vingron, M. Modeling amino acid replacement. J. Comput. Biol. 7, 761–776 (2000).
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. ProtTest 3: fast selection of best-fit models of protein evolution. Bioinformatics 27, 1164–1165 (2011).
Birney, E., Clamp, M. & Durbin, R. Genewise and genomewise. Genome Res. 14, 988–995 (2004).
Yachdav, G. et al. MSAViewer: interactive JavaScript visualization of multiple sequence alignments. Bioinformatics 32, 3501–3503 (2016).
Wheeler, T. J., Clements, J. & Finn, R. D. Skylign: a tool for creating informative, interactive logos representing sequence alignments and profile hidden Markov models. BMC Bioinformatics 15, 7 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Sakamoto, H., Inoue, K., Higuchi, I., Ono, Y. & Shimura, Y. Control of Drosophila sex-lethal pre-mRNA splicing by its own female-specific product. Nucleic Acids Res. 20, 5533–5540 (1992).
Fritzenwanker, J. H. & Technau, U. Induction of gametogenesis in the basal cnidarian Nematostella vectensis (Anthozoa). Dev. Genes Evol. 212, 99–103 (2002).
Lambert, J. P., Tucholska, M., Go, C., Knight, J. D. & Gingras, A. C. Proximity biotinylation and affinity purification are complementary approaches for the interactome mapping of chromatin-associated protein complexes. J. Proteomics 118, 81–94 (2015).
Youn, J. Y. et al. High-density proximity mapping reveals the subcellular organization of mRNA-associated granules and bodies. Mol. Cell 69, 517–532 (2018).
Liu, G. et al. Data independent acquisition analysis in ProHits 4.0. J. Proteomics 149, 64–68 (2016).
Deutsch, E. W. et al. A guided tour of the trans-proteomic pipeline. Proteomics 10, 1150–1159 (2010).
Shteynberg, D. et al. iProphet: multi-level integrative analysis of shotgun proteomic data improves peptide and protein identification rates and error estimates. Mol. Cell. Proteomics 10, M111.007690 (2011).
Teo, G. et al. SAINTexpress: improvements and additional features in Significance Analysis of INTeractome software. J. Proteomics 100, 37–43 (2014).
Mackereth, C. D. et al. Multi-domain conformational selection underlies pre-mRNA splicing regulation by U2AF. Nature 475, 408–411 (2011).
Blencowe, B. J. & Lamond, A. I. Purification and depletion of RNP particles by antisense affinity chromatography. Methods Mol. Biol. 118, 275–287 (1999).
Barabino, S. M., Blencowe, B. J., Ryder, U., Sproat, B. S. & Lamond, A. I. Targeted snRNP depletion reveals an additional role for mammalian U1 snRNP in spliceosome assembly. Cell 63, 293–302 (1990).
Dignam, J. D., Lebovitz, R. M. & Roeder, R. G. Accurate transcription initiation by RNA polymerase II in a soluble extract from isolated mammalian nuclei. Nucleic Acids Res. 11, 1475–1489 (1983).
Vizcaino, J. A. et al. ProteomeXchange provides globally coordinated proteomics data submission and dissemination. Nat. Biotechnol. 32, 223–226 (2014).
Acknowledgements
The authors thank B. Lehner and S. W. Roy for critical reading of the manuscript, M. Akam and K. Siggens for providing access to Strigamia specimens, M. Sattler and P. Zou for pETM11 clones, S. Taylor for providing the HeLa Flp-In cell line, B. Bergum (Flow Cytometry Core Facility at the University of Bergen) for assistance with Nematostella fluorescence-activated cell sorting, and the CRG Genomics Unit. Animal silhouettes were obtained from PhyloPic. This work has been funded by the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (ERC-StG-LS2-637591 to M.Ir. and ERC-AdvG-670146 to J.V.), the Spanish Ministry of Economy and Competitiveness (BFU2014-55076-P and BFU2017-89201-P to M.Ir., BFU2014-005153 to J.V., and the ‘Centro de Excelencia Severo Ochoa 2013–2017’ (SEV-2012-0208)), AGAUR, Fundación Botín (to J.V.) and the Canadian Institutes of Health Research (to B.J.B. and A.-C.G.). RNP mass spectrometric analyses were performed at the CRG/UPF Proteomics Unit (part of ProteoRed-PRB3, supported by PE I+D+i 2013–2016 (PT17/0019) of the ISCIII and ERDF) by ‘Programa CERCA Generalitat de Catalunya’ and ‘Secretaria d’Universitats i Recerca del Departament d’Economia i Coneixement’ (2017SGR595). A.T.-M. held an FPI-SO fellowship, and Y.M. a Marie Skłodowska-Curie individual fellowship.
Author information
Authors and Affiliations
Contributions
A.T-M. performed the bioinformatic analyses and molecular biology and cell culture experiments. S.B. performed the cell culture, spliceosome assembly and RNP mass spectrometry experiments under the supervision of J.V. Y.M. performed the bioinformatics analyses. J.R. performed and analysed the AP-MS experiments under the supervision of A-C.G. and B.J.B. M.Ig. performed the Nematostella experiments under the supervision of F.R. J.P. performed the Strigamia dissections and molecular biology work. D.O. performed the co-immunoprecipitation experiments under the supervision of B.J.B. T.G., I.A., F.G. and M.S. performed the Drosophila and molecular biology experiments. M.Ir. conceived and supervised the study, provided resources and performed the bioinformatic analyses. A.T-M., S.B., M.Ir. and B.J.B. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–7
Supplementary Data 1
Conserved microexons across phyla
Supplementary Data 2
Mass spectrometry results
Supplementary Data 3
RNA-seq data used in this study
Supplementary Data 4
Orthologous genes in the six bilaterian species studied
Supplementary Data 5
Srrm protein sequences used for comparative analyses
Supplementary Data 6
List of primer sequences used in this study
Rights and permissions
About this article
Cite this article
Torres-Méndez, A., Bonnal, S., Marquez, Y. et al. A novel protein domain in an ancestral splicing factor drove the evolution of neural microexons. Nat Ecol Evol 3, 691–701 (2019). https://doi.org/10.1038/s41559-019-0813-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-019-0813-6
This article is cited by
-
Unique transcriptomes of sensory and non-sensory neurons: insights from Splicing Regulatory States
Molecular Systems Biology (2024)
-
Pancreatic microexons regulate islet function and glucose homeostasis
Nature Metabolism (2023)
-
NvPrdm14d-expressing neural progenitor cells contribute to non-ectodermal neurogenesis in Nematostella vectensis
Nature Communications (2023)
-
High frequency of alternative splicing variants of the oncogene Focal Adhesion Kinase in neuroendocrine tumors of the pancreas and breast
Frontiers of Medicine (2023)
-
Histone demethylase Lsd1 is required for the differentiation of neural cells in Nematostella vectensis
Nature Communications (2022)