Abstract
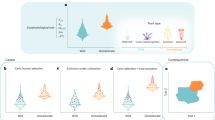
The origins of agriculture were key events in human history, during which people came to depend for their food on small numbers of animal and plant species. However, the biological traits determining which species were domesticated for food provision, and which were not, are unclear. Here, we investigate the phylogenetic distribution of livestock and crops, and compare their phenotypic traits with those of wild species. Our results indicate that phylogenetic clustering is modest for crop species but more intense for livestock. Domesticated species explore a reduced portion of the phenotypic space occupied by their wild counterparts and have particular traits in common. For example, herbaceous crops are globally characterized by traits including high leaf nitrogen concentration and tall canopies, which make them fast-growing species and proficient competitors. Livestock species are relatively large mammals with low basal metabolic rates, which indicate moderate to slow life histories. Our study therefore reveals ecological differences in domestication potential between plants and mammals. Domesticated plants belong to clades with traits that are advantageous in intensively managed high-resource habitats, whereas domesticated mammals are from clades adapted to moderately productive environments. Combining comparative phylogenetic methods with ecologically relevant traits has proven useful to unravel the causes and consequences of domestication.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All phenotypic traits of mammalian species included in this study are available from the literature (see Methods). For plants, most data are available from the database TRY59 (https://www.try-db.org), and all original sources of TRY data are listed in Supplementary References 1. All references for data not included in TRY are available in the Supplementary References 2. Unpublished data owned by R.M. and J.M.B. are available from Supplementary Data 3. Unpublished data from the University of Sheffield database of weed functional attributes can be requested from G.J. Lists of livestock and crop taxa are available in Supplementary Table 1 and Supplementary Data 1, respectively. Phylogenetic trees used in this study are available in Supplementary Data 4. Data on geography and climate at domestication sites are available as Supplementary Data 5.
References
RBG Kew. The State of the World’s Plants Report — 2016 (Royal Botanic Gardens, Kew, London, 2016).
Zeven, A. C. & De Wet, J. M. J. Dictionary of Cultivated Plants and Their Regions of Diversity: Excluding Most Ornamentals, Forest Trees and Lower Plants (Pudoc, Wageningen, 1982); http://agris.fao.org/agris-search/search.do?recordID=XE8282385
Dirzo, R. & Raven, P. H. Global state of biodiversity and loss. Annu. Rev. Environ. Resour. 28, 137–167 (2003).
Meyer, R. S. & Purugganan, M. D. Evolution of crop species: genetics of domestication and diversification. Nat. Rev. Genet. 14, 840–852 (2013).
Zeder, M. A. in Biodiversity in Agriculture: Domestication, Evolution, and Sustainability (eds Gepts, P. et al.) 227–259 (Cambridge Univ. Press, New York, 2012).
Clutton-Brock, J. A Natural History of Domesticated Mammals (Cambridge Univ. Press, New York, 1999).
Wilson, D. & Reeder, D. A. Mammal Species of the World (Johns Hopkins Univ. Press, Baltimore, 2005).
Gepts, P. in Plant Breeding Reviews Vol. 24 (ed. Janick, J.) Ch. 1, 1–44 (John Wiley & Sons, Hoboken, 2010).
Blomberg, S. P. & Garland, T. Jr. Tempo and mode in evolution: phylogenetic inertia, adaptation and comparative methods. J. Evol. Biol. 15, 899–910 (2002).
Diamond, J. Evolution, consequences and future of plant and animal domestication. Nature 418, 700–707 (2002).
Salman-Minkov, A., Sabath, N. & Mayrose, I. Whole-genome duplication as a key factor in crop domestication. Nat. Plants 2, 16115 (2016).
Whitehead, S. R., Turcotte, M. M. & Poveda, K. Domestication impacts on plant–herbivore interactions: a meta-analysis. Phil. Trans. R. Soc. B 372, 20160034 (2017).
Milla, R., Osborne, C. P., Turcotte, M. M. & Violle, C. Plant domestication through an ecological lens. Trends Ecol. Evol. 30, 463–469 (2015).
Preece, C. et al. Were Fertile Crescent crop progenitors higher yielding than other wild species that were never domesticated? New Phytol. 207, 905–913 (2015).
Darwin, C. The Variation of Animals and Plants under Domestication Vol. 2 (O. Judd, London, 1868).
Abbo, S. et al. Plant domestication versus crop evolution: a conceptual framework for cereals and grain legumes. Trends Plant Sci. 19, 351–360 (2014).
Harlan, J. R. Crops and Man (American Society of Agronomy, Madison, 1992).
Meyer, R. S., DuVal, A. E. & Jensen, H. R. Patterns and processes in crop domestication: an historical review and quantitative analysis of 203 global food crops. New Phytol. 196, 29–48 (2012).
Sánchez-Villagra, M. R., Geiger, M. & Schneider, R. A. The taming of the neural crest: a developmental perspective on the origins of morphological covariation in domesticated mammals. R. Soc. Open Sci. 3, 160107 (2016).
Larson, G. & Fuller, D. Q. The evolution of animal domestication. Annu. Rev. Ecol. Evol. Syst. 45, 115–136 (2014).
Reich, P. B. The world‐wide ‘fast–slow’plant economics spectrum: a traits manifesto. J. Ecol. 102, 275–301 (2014).
Milla, R., Morente-López, J., Alonso-Rodrigo, J. M., Martín-Robles, N. & Chapin, F. S. III. Shifts and disruptions in resource-use trait syndromes during the evolution of herbaceous crops. Proc. R. Soc. B 281, 20141429 (2014).
Tribouillois, H. et al. A functional characterisation of a wide range of cover crop species: growth and nitrogen acquisition rates, leaf traits and ecological strategies. PLoS ONE 10, e0122156 (2015).
Milla, R. & Matesanz, S. Growing larger with domestication: a matter of physiology, morphology or allocation? Plant Biol. 19, 475–483 (2017).
Díaz, S. et al. The global spectrum of plant form and function. Nature 529, 167–171 (2015).
Ricklefs, R. E. & Wikelski, M. The physiology/life-history nexus. Trends Ecol. Evol. 17, 462–468 (2002).
Lovegrove, B. G. The zoogeography of mammalian basal metabolic rate. Am. Nat. 156, 201–219 (2000).
Lovegrove, B. G. The influence of climate on the basal metabolic rate of small mammals: a slow-fast metabolic continuum. J. Comp. Physiol. B 173, 87–112 (2003).
Zohary, D., Tchernov, E. & Horwitz, L. K. The role of unconscious selection in the domestication of sheep and goats. J. Zool. 245, 129–135 (1998).
Cunniff, J. et al. Functional traits differ between cereal crop progenitors and other wild grasses gathered in the Neolithic Fertile Crescent. PLoS ONE 9, e87586 (2014).
Biro, P. & Stamps, J. Are animal personality traits linked to life-history productivity? Trends Ecol. Evol. 23, 361–368 (2008).
Dittmann, M. T. et al. Characterising an artiodactyl family inhabiting arid habitats by its metabolism: low metabolism and maintenance requirements in camelids. J. Arid Environ. 107, 41–48 (2014).
Careau, V., Bininda‐Emonds, O. R. P., Thomas, D. W., Réale, D. & Humphries, M. M. Exploration strategies map along fast–slow metabolic and life‐history continua in muroid rodents. Funct. Ecol. 23, 150–156 (2009).
Réale, D. et al. Personality and the emergence of the pace-of-life syndrome concept at the population level. Phil. Trans. R. Soc. B 365, 4051–4063 (2010).
Found, R. & St. Clair, C. C. Ambidextrous ungulates have more flexible behaviour, bolder personalities and migrate less. Open Sci. 4, 160958 (2017).
Tchernov, E. & Horwitz, L. K. Body size diminution under domestication: unconscious selection in primeval domesticates. J. Anthropol. Archaeol. 10, 54–75 (1991).
Rensch, B. Evolution Above the Species Level (Columbia Univ. Press, New York, 1960).
Vigne, J.-D. The origins of animal domestication and husbandry: a major change in the history of humanity and the biosphere. C. R. Biol. 334, 171–181 (2011).
Li, Y. et al. Habitat filtering determines the functional niche occupancy of plant communities worldwide. J. Ecol. 106, 1001–1009 (2018).
Donovan, L. A., Mason, C. M., Bowsher, A. W., Goolsby, E. W. & Ishibashi, C. D. A. Ecological and evolutionary lability of plant traits affecting carbon and nutrient cycling. J. Ecol. 102, 302–314 (2014).
Rotundo, J. L. & Cipriotti, P. A. Biological limits on nitrogen use for plant photosynthesis: a quantitative revision comparing cultivated and wild species. New Phytol. 214, 120–131 (2017).
Denison, R. F. Darwinian Agriculture: How Understanding Evolution Can Improve Agriculture (Princeton Univ. Press, Princeton, 2012).
Miller, A. J. & Gross, B. L. From forest to field: perennial fruit crop domestication. Am. J. Bot. 98, 1389–1414 (2011).
Brown, J. & West, G. Scaling in Biology (Oxford Univ. Press, Oxford, 2000).
West, G. B., Brown, J. H. & Enquist, B. J. General model for the origin of allometric scaling laws in biology. Science 276, 122–126 (1997).
Reich, P., Tjoelker, M., Machado, J.-L. & Oleksyn, J. Universal scaling of respiratory metabolism, size and nitrogen in plants. Nature 439, 457–461 (2006).
Preece, C. et al. How did the domestication of Fertile Crescent grain crops increase their yields? Funct. Ecol. 31, 387–397 (2017).
Vasseur, F., Violle, C., Enquist, B. J., Granier, C. & Vile, D. A common genetic basis to the origin of the leaf economics spectrum and metabolic scaling allometry. Ecol. Lett. 15, 1149–1157 (2012).
Martin, A. R. et al. Inter- and intraspecific variation in leaf economic traits in wheat and maize. AoB Plants 10, ply006 (2018).
Martin, A. et al. Intraspecific trait variation across multiple scales: the leaf economics spectrum in coffee. Funct. Ecol. 31, 604–612 (2017).
Diamond, J. Guns, Germs, and Steel: The Fates of Human Societies (W. W. Norton, London, 1997).
Smythe, N. & Brown de Guanti, O. Domestication and Husbandry of the Paca (Agouti paca) FAO conservation guide 26 (Food and Agriculture Organization of the United Nations, Rome, 1995).
Moreira, J. R. & Pinheiro, M. S. in Capybara: Biology, Use and Conservation of an Exceptional Neotropical Species (eds Moreira, J. R. et al.) 333–344 (Springer, New York, 2013).
Chamberlain, S. A. & Szocs, E. taxize: taxonomic search and retrieval in R. F1000Research 2, 191 (2013).
Vincent, H. et al. A prioritized crop wild relative inventory to help underpin global food security. Biol. Conserv. 167, 265–275 (2013).
Mansfeld, R. & Hanelt, P. Mansfeld’s Encyclopedia of Agricultural and Horticultural Crops (Springer, Berlin, 2001).
Cayuela, L., Granzow‐de la Cerda, Í., Albuquerque, F. S. & Golicher, D. J. Taxonstand: an R package for species names standardisation in vegetation databases. Methods Ecol. Evol. 3, 1078–1083 (2012).
Kissling, W. D. et al. Establishing macroecological trait datasets: digitalization, extrapolation, and validation of diet preferences in terrestrial mammals worldwide. Ecol. Evol. 4, 2913–2930 (2014).
Kattge, J. et al. TRY—a global database of plant traits. Global Change Biol. 17, 2905–2935 (2011).
Zanne, A. E. et al. Three keys to the radiation of angiosperms into freezing environments. Nature 506, 89–92 (2014).
Moles, A. T. et al. Global patterns in plant height. J. Ecol. 97, 923–932 (2009).
Glazier, D. S. & Eckert, S. E. Competitive ability, body size and geographical range size in small mammals. J. Biogeogr. 29, 81–92 (2002).
Wright, I. J. et al. The worldwide leaf economics spectrum. Nature 428, 821–827 (2004).
Moles, A. T. et al. A brief history of seed size. Science 307, 576–580 (2005).
Riesch, R., Martin, R. A., Lerp, H., Plath, M. & Wronski, T. Size and sex matter: reproductive biology and determinants of offspring survival in Gazella marica. Biol. J. Linn. Soc. 110, 116–127 (2013).
Jones, K. E. et al. PanTHERIA: a species‐level database of life history, ecology, and geography of extant and recently extinct mammals. Ecology 90, 2648 (2009).
Genoud, M., Isler, K. & Martin, R. Comparative analyses of basal rate of metabolism in mammals: data selection does matter. Biol. Rev. 93, 404–438 (2018).
Martin, A. R. & Isaac, M. E. Plant functional traits in agroecosystems: a blueprint for research. J. Appl. Ecol. 52, 1425–1435 (2015).
Nakagawa, S. & Freckleton, R. P. Missing inaction: the dangers of ignoring missing data. Trends Ecol. Evol. 23, 592–596 (2008).
Penone, C. et al. Imputation of missing data in life‐history trait datasets: which approach performs the best? Methods Ecol. Evol. 5, 961–970 (2014).
van Buuren, S. & Groothuis-Oudshoorn, K. mice: multivariate imputation by chained equations in R. J. Stat. Softw. 45, 1–67 (2011).
Santos, T. PVR: phylogenetic eigenvectors regression and phylogentic signal-representation curve. R package version 0.3 (2018); https://cran.r-project.org/web/packages/PVR/index.html
Bininda-Emonds, O. R. P. et al. The delayed rise of present-day mammals. Nature 446, 507–512 (2007).
Qian, H. & Jin, Y. An updated megaphylogeny of plants, a tool for generating plant phylogenies and an analysis of phylogenetic community structure. J. Plant Ecol. 9, 233–239 (2016).
Keck, F., Rimet, F., Bouchez, A. & Franc, A. phylosignal: an R package to measure, test, and explore the phylogenetic signal. Ecol. Evol. 6, 2774–2780 (2016).
de Villemereuil, P. & Nakagawa, S. in Modern Phylogenetic Comparative Methods and Their Application in Evolutionary Biology (ed. Garamszegi, L.) 287–304 (Springer-Verlag, Berlin, 2014).
Hadfield, J. & Nakagawa, S. General quantitative genetic methods for comparative biology: phylogenies, taxonomies and multi-trait models for continuous and categorical characters. J. Evol. Biol. 23, 494–508 (2010).
Felsenstein, J. Phylogenies and quantitative characters. Annu. Rev. Ecol. Syst. 19, 445–471 (1988).
Burnham, K. P. & Anderson, D. R. Model Selection and Multimodel Inference: A Practical Information-theoretic Approach (Springer-Verlag, New York, 2002).
Hadfield, J. MCMC methods for multi-response generalised linear mixed models: the MCMCglmm R package. J. Stat. Softw. 33, 1–22 (2010).
Harmon, L. J., Weir, J. T., Brock, C. D., Glor, R. E. & Challenger, W. GEIGER: investigating evolutionary radiations. Bioinformatics 24, 129–131 (2007).
Webb, C. O. & Donoghue, M. J. Phylomatic: tree assembly for applied phylogenetics. Mol. Ecol. Resour. 5, 181–183 (2005).
Paradis, E., Claude, J. & Strimmer, K. APE: analyses of phylogenetics and evolution in R language. Bioinformatics 20, 289–290 (2004).
Cornwell, W. K., Schwilk, D. W. & Ackerly, D. D. A trait‐based test for habitat filtering: convex hull volume. Ecology 87, 1465–1471 (2006).
Barber, C. B. et al. geometry: mesh generation and surface tessellation. R package version 0.3-4 (2014); https://cran.r-project.org/web/packages/geometry/index.html
R Core Team. R: a language and environment for statistical computing (R Foundation for Statistical Computing, 2016); https://www.r-project.org/
Acknowledgements
R.M., J.C.-L. and J.M.B. were funded by grants CGL2014-56567-R, CGL2017-83855-R and PCIN-2014-053 (Ministerio de Economia y Competitividad, MINECO, Spain), and Eco-serve project (Biodiversa–FACCE, Horizon 2020, European Union). The study has been supported by the TRY initiative on plant traits (http://www.try-db.org). The TRY initiative and database is hosted, developed and maintained by J. Kattge and G. Bönisch (Max Planck Institute for Biogeochemistry, Jena, Germany). TRY is currently supported by DIVERSITAS/Future Earth and the German Centre for Integrative Biodiversity Research (iDiv) Halle-Jena -Leipzig. R.M. thanks C. F. Ingala from Universidad Rey Juan Carlos. E.E.S. thanks the FAPESP/BIOTA programme for financial support. J.K. thanks BACI (EU grant ID 640176). T.H. thanks the support of Australian Research Council (DP130013029). V.D.P. was supported by CNPq, Brazil, grant no. 307689/2014-0, K.K. was funded by the project Resilient Forests (KB-29-009-003).
Author information
Authors and Affiliations
Contributions
R.M. and J.M.B. designed the study and compiled the data. R.M., J.M.B., J.C.-L. and M.M.T. performed statistical analyses. R.M. and J.M.B. wrote a first draft of the paper. M.M.T., G.J., C.P.O. and C.V. extensively revised drafts. All authors contributed to the writing of, and approved, the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Methods, Supplementary Figures, Supplementary Tables, Supplementary Note, and Supplementary References of the Article.
Supplementary Data 1
List of 944 domesticated angiosperms included in this study, with information about family adscription and growth forms (herbaceous, graminoid and woody). Bold typing indicates the 181 species included in phenotypic space analyses.
Supplementary Data 2
Results of the analyses on Local Indicators of Phylogenetic Association analysis (LIPA). LIPA values for relative abundance of domesticates and domestication frequencies at family level are shown for mammals and angiosperms families.
Supplementary Data 3
Unpublished crop trait dataset of R.M. and J.M.B.
Supplementary Data 4
Phylogenetic trees used in this study.
Supplementary Data 5
Data, and sources for, wild progenitor assignment, and antiquity of domestication, for the domesticates included in phenotypic space analyses.
Rights and permissions
About this article
Cite this article
Milla, R., Bastida, J.M., Turcotte, M.M. et al. Phylogenetic patterns and phenotypic profiles of the species of plants and mammals farmed for food. Nat Ecol Evol 2, 1808–1817 (2018). https://doi.org/10.1038/s41559-018-0690-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-018-0690-4
This article is cited by
-
Variations and trade-offs in leaf and culm functional traits among 77 woody bamboo species
BMC Plant Biology (2024)
-
Early human selection of crops’ wild progenitors explains the acquisitive physiology of modern cultivars
Nature Plants (2024)
-
Evolutionary imbalance, climate and human history jointly shape the global biogeography of alien plants
Nature Ecology & Evolution (2023)
-
Accelerating crop domestication through genome editing for sustainable agriculture
Journal of Plant Biochemistry and Biotechnology (2023)
-
Signatures of selection in recently domesticated macadamia
Nature Communications (2022)