Abstract
Essential Biodiversity Variables (EBVs) allow observation and reporting of global biodiversity change, but a detailed framework for the empirical derivation of specific EBVs has yet to be developed. Here, we re-examine and refine the previous candidate set of species traits EBVs and show how traits related to phenology, morphology, reproduction, physiology and movement can contribute to EBV operationalization. The selected EBVs express intra-specific trait variation and allow monitoring of how organisms respond to global change. We evaluate the societal relevance of species traits EBVs for policy targets and demonstrate how open, interoperable and machine-readable trait data enable the building of EBV data products. We outline collection methods, meta(data) standardization, reproducible workflows, semantic tools and licence requirements for producing species traits EBVs. An operationalization is critical for assessing progress towards biodiversity conservation and sustainable development goals and has wide implications for data-intensive science in ecology, biogeography, conservation and Earth observation.
Similar content being viewed by others
Main
In 2013, the Group on Earth Observations Biodiversity Observation Network (GEO BON) introduced the framework of Essential Biodiversity Variables (EBVs) to derive coordinated measurements critical for detecting and reporting biodiversity change1. Through this process, 22 candidate EBVs were proposed and organized within six classes (‘genetic composition’, ‘species populations’, ‘species traits’, ‘community composition’, ‘ecosystem structure’ and ‘ecosystem function’)1. These EBVs provide a foundation for assessing progress towards national and international policy goals, including the 20 Aichi Biodiversity Targets developed by the Parties to the United Nations (UN) Convention on Biological Diversity (CBD) and the 17 Sustainable Development Goals (SDGs) identified by the UN 2030 Agenda for Sustainable Development2. EBVs are conceptually located on a continuum between primary data observations (‘raw data’) and synthetic or derived metrics (‘indicators’), and can be represented as ‘data cubes’ with several basic dimensions (for example, time, space, taxonomy or Earth observation data types)3,4,5. Hence, EBVs allow derivation of biodiversity indicators (for example, trends of biodiversity change) such as those developed for the Aichi Biodiversity Targets, with several EBVs (for example, species population abundance) informing multiple targets1,6. Specific EBVs in the classes species populations, ecosystem structure and ecosystem function are now being developed by GEO BON working groups7. However, other EBV classes have received less attention, and the research community has yet to fully coalesce efforts to develop the conceptual and empirical frameworks for those variables and their associated data products.
Species traits are a key component of biodiversity because they determine how organisms respond to disturbances and changing environmental conditions, with impacts at a population level and beyond8,9,10. Within the EBV framework, the EBV class ‘species traits’ has yet to be formally conceptualized in detail and therefore cannot yet be made operational. In line with previous work8,11,12, we here define a species trait as any phenological, morphological, physiological, reproductive or behavioural characteristic of an individual that can be assigned to a species (Box 1). Because the building of EBV data products requires standardization and harmonization of raw measurements1,3,5, we further define species traits EBVs as standardized and harmonized measurements of species’ characteristics that allow monitoring of intra-specific trait changes within species populations across space and time (Box 1). Specific species traits selected for EBVs (for example, body mass, plant height and specific leaf area as examples of morphological traits) allow quantification of how species respond to global change including climate change, biological invasions, overexploitation and habitat fragmentation8,13,14,15,16 (Box 1). The time frame of species traits responses should be policy relevant, that is, intra-specific trait changes should be detectable within a decade rather than only seasonally, annually or over evolutionary time scales6. This is needed because EBVs will feed into biodiversity change indicators (Box 1) that allow the assessment of progress towards policy goals including the SDGs and Aichi Biodiversity Targets as well as National Biodiversity Strategies and Action Plans (NBSAPs). They can also help to inform global and regional assessments of the Intergovernmental Platform on Biodiversity and Ecosystem Services (IPBES)1,17. Other aspects of species traits that reflect traits expressions at the community or ecosystem level are not considered here as they belong to other EBV classes (Box 1). To our knowledge there are currently no global data products available that allow direct measurement and monitoring of trait changes within species populations across time17.
Here, we develop the conceptual and empirical basis for species traits EBVs to help to operationalize the development of global EBV data products. We start by critically re-examining the current set of candidate species traits EBVs (phenology, body mass, natal dispersal distance, migratory behaviour, demographic traits and physiological traits). We then explore how trait data are collected, how they can be standardized and harmonized and what bottlenecks currently prevent them from becoming findable, accessible, interoperable and reusable (FAIR guiding principles)18. We further outline workflow steps to produce EBV data products of species traits, using an example of plant phenology. Our perspective provides a conceptual framework with practical guidelines for building global, integrated and reusable EBV data products of species traits. This will promote the use of species trait information in national and international policy assessments and requires significant advancements and new tools in ecology, biogeography, conservation and environmental science. Beyond the direct relevance to species traits EBVs, our perspective further explores cross-cutting issues related to data-intensive science, interoperability, and legal and policy aspects of biodiversity monitoring and Earth observation that will help to advance the EBV framework.
A critical re-examination
GEO BON has proposed six candidate EBVs in the EBV class species traits (Supplementary Table 1): phenology, body mass, natal dispersal distance, migratory behaviour, demographic traits and physiological traits. These candidate EBVs were discussed in detail during a three-day experts’ workshop in Amsterdam (March 2017) organized by the GLOBIS-B project (http://www.globis-b.eu/)19. We suggest several key improvements of that initial list of candidate species traits EBVs.
Identified inconsistencies
We identified several inconsistencies in the proposed candidate list of species traits EBVs (summarized in Supplementary Table 2). First, some previously listed measurements — such as ocean and river flows, extent of wetlands and net primary productivity — do not occur at the species level (Box 1) and should therefore be placed within community or ecosystem-scale EBV classes such as community composition, ecosystem function or ecosystem structure. Second, several candidate EBVs (for example, body mass and natal dispersal distance) are narrowly defined compared to other candidate EBVs (for example, phenology, demographic traits, physiological traits), resulting in an inconsistent scope across EBVs. Third, a few candidate EBVs represent a similar category but are split into different EBVs (for example, both natal dispersal distance and migratory behaviour are aspects of movement behaviour), and should therefore be represented together. Fourth, the candidate EBV ‘demographic traits’ reflects population-level quantities that cannot be measured on individual organisms (for example, population growth rate, generation time, survival rate). These population-level metrics are derived from data that are captured by the EBV population structure by age/size/stage class belonging to another EBV class (species populations). It is therefore inconsistent to capture the same set of underlying measurements in two different EBV classes.
Suggestions for improvement
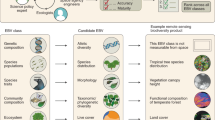
Based on our assessment, we suggest reducing the initial candidate list to five species traits EBVs (Fig. 1): phenology (timing of periodic biological events), morphology (dimensions, shape and other physical attributes of organisms), reproduction (sexual or asexual production of new individual organisms), physiology (chemical or physiological functions promoting organism fitness) and movement (spatial mobility of organisms) (see overview in Fig. 1 and detailed description in Supplementary Note 1). This improves the previous classification of species traits EBVs by standardizing the breadth and scope of EBVs, better recognizing the importance and relevance of reproductive traits and excluding ecosystem variables that cannot be measured at the scale of the individual and are thus not species-specific traits (Supplementary Note 1). These five species traits EBVs provide a conceptual framework for the EBV class species traits and are relevant to the Aichi Biodiversity Targets and SDGs (Fig. 1, Supplementary Table 3). Because GEO BON has the main responsibility for developing EBVs, we suggest that the new GEO BON working group on species traits (as recommended in the GEO BON implementation plan 2017–20207) should take our suggestions into consideration when updating the EBV class species traits.
We suggest five EBVs within the EBV class ‘species traits’, comprising (1) phenology, (2) morphology, (3) reproduction, (4) physiology and (5) movement. For each EBV, a definition, examples of species trait measurements, temporal sensitivity and societal relevance are given. Societal relevance refers to those Aichi Biodiversity Targets and SDGs to which the specific EBV is of highest relevance (for details on societal relevance see Supplementary Note 2 and Supplementary Table 2). Photo credits: Katja-Sabine Schulz.
Collecting trait data
Many trait databases have recently emerged that support assembling trait measurements from published literature, specimen collections, in situ collections and close-range, airborne or spaceborne remote sensing (for examples see Supplementary Table 4). Nevertheless, the total demand for species traits in the EBV context is still unmet for the following reasons.
Aggregated species-level trait values are not sufficient
Many ongoing trait data collections assemble species trait information from published literature (Fig. 2). When aggregated to the species-level without location and time information (for example, mean species body length for morphology, or typical month of flowering or fruiting for phenology), this information does not allow measurement of trait changes within species populations over space or time, and hence lacks the ability to yield species traits EBVs (Fig. 2, Box 1). However, if the variation in the aggregated trait (that is, variance) can be calculated from a sufficiently large sample, then changes in species populations over time (or space) can be statistically estimated15,20,21,22. Nevertheless, many projects aggregate trait data at the species level from multiple sources such as published and unpublished trait datasets, natural history collections, citizen science projects and text mining23,24,25,26,27,28. These trait data remain limited in their application for species traits EBVs if they do not keep the resolution of the original data in terms of space, time and individual measurement information. The lack of individual or population measures therefore makes it difficult to assess intra-specific trait changes and the drivers and scales at which they operate.
Several methods are used to assemble comprehensive trait databases, for example, from published literature, specimen collections, in situ monitoring and remote sensing (close-range, airborne and spaceborne). These methods can be ordered along a gradient of increasing temporal frequency of observations. Aggregation of trait data from multiple sources often does not provide measurements repeated in time and hence typically does not allow monitoring of trait changes within species populations. More information about trait databases (abbreviations) is provided in Supplementary Table 3.
Natural history collections offer historical data that remain underutilized
Museum and herbarium specimens allow study of individuals’ traits in species populations of the recent past29. Specimen collections can therefore be an important source for individual-level trait measurements through time (Fig. 2). For example, specimens have been used to document temporal changes in morphology (for example, bird and beetle body size30,31) and phenology (for example, timing of flowering32,33) during the past century. Billions of specimens are available for study, but efforts to digitize and store trait data associated with specimens are still in their infancy29. Hence, trait data from digitized specimen collections remain underutilized and are currently too often constrained and biased in space, time and number of individuals25. New ways to digitize biocollections and to automate trait data extraction from specimens are needed25, and analyses must take into account the constraints and biases inherent in these data34.
In situ monitoring of traits is promising but labour intensive
A promising approach for developing species traits EBVs is to collect in situ trait data through monitoring schemes (Fig. 2). These include repeated trait measurements (for example, of animal body size, plant size, lichen length, flower and fruit phenology, leaf morphology and chemistry) with standardized protocols using long-term ecological research sites35,36 or national and international monitoring programmes and citizen science networks20,37,38. Such sites and networks can monitor a comprehensive set of trait measurements for targeted species or sites through time and at continental extents38,39, but remain costly and labour intensive. The future collection of trait data time series through in situ monitoring therefore requires prioritization according to global and regional biodiversity and sustainability goals, and a robust temporal replication and spatial/environmental stratification of the sampling design40.
Remote sensing observations are promising but often not species specific
Airborne, spaceborne and close-range remote sensing techniques are promising tools (Fig. 2) because they can extend the geographic and temporal dimensions of trait measurements considerably9,41,42,43. Increasingly, ground-based light detection and ranging (that is, terrestrial LiDAR) is automating in situ data collection and allows retrieval of species trait information for individual plants (for example, height44 and leaf water content45). Moreover, sensor-derived trait data can provide individual- or population-level trait measurements from close-range instruments such as camera traps, phenology cameras46,47, field spectrometers48, wireless sensor networks, unmanned aerial vehicle (UAV) and aircraft mounted instruments such as airborne LiDAR and hyperspectral sensors49,50. Combining airborne LiDAR and imaging spectroscopy also allows mapping of individual-level variation in morphological and physiological traits (for example, canopy height, leaf chlorophyll and water content) at regional scales43. For species traits EBVs, the remotely sensed trait measurements require fine enough spatial resolution to attribute them to an individual or population of a particular species (Box 1). A synergy of hyperspectral and LiDAR remote sensing with airborne sensors has great potential for developing species traits EBVs, but is not available at a global extent. Spaceborne remote sensing systems can provide global coverage, but they still show a large deficit for providing an operational combination of data at high spatial and spectral resolution9,42,51. In other words, spaceborne instruments are in their infancy for monitoring species traits due to limitations with very high spatial resolution (pixel area) and spectral resolution (high number and small width of spectral bands), though new spaceborne imaging spectrometers and LiDAR are planned which will go some way towards closing this gap42,52,53. Further developments in instrumentation and data52, planned satellite sensor missions53, species-level spectral library databases (for example, EcoSIS; https://ecosis.org) and spectranomics54,55 — the coupling of spectroscopy with plant phylogeny and canopy chemistry — will further enhance the ability to retrieve species-specific trait data.
Standardizing trait data
A current bottleneck for integrating trait datasets from multiple sources is that measurements, data and metadata are not sufficiently standardized. We highlight three focal areas to improve this.
Standardizing protocols for measuring traits
The use of standardized measurement protocols during the phase of trait data collection is foundational for integrating data into EBV data products. Good examples of comprehensive protocols for standardized measurements of morphological, reproductive, physiological and behavioural traits exist for vascular plants56,57 and terrestrial invertebrates58. However, such comprehensive definitions of measurement protocols are still missing for most traits and taxa, and some remain little-known and difficult to access59. This is particularly true for remote sensing measurements of species traits (for example, leaf chlorophyll concentration and canopy chlorophyll content) where the instrumentation and required pre-processing of data to derive information on species-specific traits may vary considerably even within the same class of sensors (for example, within different types of spectrometers, phenology cameras or LiDAR instruments). A coordinated effort is therefore needed to develop and harmonize standardized measurement protocols for various taxa and across data types, sensors and regions, and to support consistent monitoring across political boundaries.
Standardizing trait terminology
Aggregating trait data from multiple sources requires standardized lists of trait terms or controlled vocabularies (that is, carefully selected lists of words and phrases)11,27,60,61. For instance, in the marine domain the formalization of a standardized list of trait terms and definitions has been achieved across a wide range of taxa26,60. Similar examples exist for other taxa and realms, for example, the thesaurus of plant characteristics11. Nevertheless, comprehensive trait vocabularies that provide standardized terms, definitions, units and synonyms for trait data and their metadata remain scarce. The further development and linking of such trait vocabularies is therefore needed to achieve semantic interoperability and facilitate integration of trait datasets11,23,27,62.
Ontologies
Integrating trait data from disparate sources requires mapping trait data to ontologies23,25,61,63,64,65,66, that is, to semantic models that allow formal descriptions of the relationships among trait concepts and vocabulary terms (Box 2). For trait data in particular, not only information about the occurrence of a species and the identification process needs to be reported, but also information about the entity (that is, whether specific parts of organisms, individual organisms, populations or species are measured), the measurement focus (for example, mass, length or area), the measurement units (for example, plant height in m, leaf nitrogen content in mg g–1, photosynthetic rate in μmol m2 s–1) and the protocols used. Because many traits exhibit phenotypic plasticity, information about the individuals’ living conditions before trait measurements (for example, if a plant was exposed to direct sunlight or shaded in the understory) is also essential to understand and interpret trait measurements67. Such reporting can be standardized by connecting two types of ontology: (1) observation and measurement ontologies for traits and environmental conditions and (2) ontologies for entities and qualities (Box 2). Various examples of both types of ontology already exist (Box 2), but their wider integration for developing comprehensive species traits data products has not yet been achieved.
Making trait data open and machine-readable
A workflow-oriented production of EBVs requires trait datasets and their metadata to be openly accessible and machine-readable3,18. Although openness and sharing of biodiversity data are improving68,69,70 and trait databases increasingly develop data management policies around open access principles (see Supplementary Note 3 for an assessment of openness of individual species traits datasets), the actual levels of open and FAIR18 access to trait data are still lagging behind the ideal, although remote-sensing data are increasingly freely available, especially through space agencies (for example, NASA and the European Space Agency). Here, we highlight two key steps for enhancing openness and machine-driven integration of trait datasets.
Use standardized copyright waivers and licences
Waivers and licences support legal interoperability by clearly defining the conditions for both creation and use of combined or derivative data products, and allow users to legally access and use data without seeking additional authorization from the rights holders71. Many trait datasets do not yet use standardized copyright waiver or licence information such as those published through the Creative Commons (CC) framework72. In the context of EBVs, the formal designation with a CC0 copyright waiver or an open CC BY licence have been recommended because they minimize constraints on legal interoperability that emerge from restrictions on data use, modification and sharing3. Although a waiver of copyright through CC0 makes sharing and reuse much easier, the appropriate ‘attribution’ and maintenance of data provenance is important in a scientific context18, and the CC BY licence provides the opportunity for acknowledgement and citation.
Provide standardized and machine-readable metadata
Many trait datasets are already available through web portals and other developed infrastructures (Supplementary Table 4), but access to standardized and machine-readable trait data and metadata remains a key bottleneck for technical and legal interoperability. For instance, licence and citation information is often not available in standardized and machine-readable form (for example, by using hyperlinks or embedded code, Supplementary Note 3) and many research projects publish their trait data on file hosting services (for example, Figshare, Dryad, Zenodo and so on) where no data and metadata standards are forced upon the uploaded material27. Moreover, metadata on the level of individual trait records is usually missing and data provenance is rarely documented (Supplementary Note 3). Hence, sufficient, consistent and well-documented metadata in a standardized form should be provided to successfully integrate trait measurements into workflows for building EBV data products of species traits.
A workflow for integrating EBV-relevant trait data
The production of species traits EBVs can only be achieved if multiple trait datasets are harmonized and combined into open, accessible and reusable products3. However, most trait data are currently stored in siloed resources and not available in an interoperable and machine-readable format. We therefore outline a generalized workflow for integrating EBV-relevant trait data (Fig. 3) and show how this workflow is currently applied to produce a new integrated plant phenology dataset (Box 3).
Initial species trait measurements are collected through human observations and remote sensing and subsequently quality checked and bundled into datasets (1). Because such datasets often have different sampling protocols, reporting processes and metadata descriptions, they commonly end up as siloed datasets in file hosting services with little metadata documentation and data standardization. To achieve integration of different measurements and data collections, datasets must be harmonized through standardization of data and metadata and mapped to community-developed standards, including metadata standards, controlled vocabularies and ontologies (2). Standardization often includes a second QA and QC process to assure data quality across datasets (not shown). Such harmonized data products can then be made accessible through open licences, databases that employ semantic web standards and APIs, and web platforms or widely used software (3).
Collecting and provisioning trait data
The first part of the workflow represents the collection and initial processing of raw measurements of traits (for example, on flower and leaf phenology) following standardized sampling protocols, for example, by people (specimen collection and in situ observations) or close-range, airborne and spaceborne remote sensing (Fig. 3, top). After collection, raw data are validated through data quality assurance (QA, for example, by following standard protocols for trait data cleaning) and quality control (QC, for example, normalizing trait distributions, checking for outliers) (Fig. 3, top). Metadata about trait data collection and validation processes (for example, description of protocols) and about the dataset itself (for example, specimen IDs, ownership and licensing) need to be associated with the data when bundling the trait datasets (Box 3). Most currently existing trait datasets are only published in repositories with little metadata documentation and data standardization, but efforts to integrate them into more comprehensive data products are beginning to emerge.
Converting trait data into interoperable formats
To achieve integrated trait data products, data and metadata from different sources have to be standardized (Fig. 3, middle). This involves converting all data to comparable units and formats, the mapping of trait data to ontologies and automated reasoning over mapped data to discover new facts (Fig. 3, middle). The use of ontologies, for example, the Plant Phenology Ontology (PPO)73 for flower and leaf phenology traits (Box 3), provides a formal, generalized, logical structure that helps to automate integration across different datasets. Ontologies can also be used to further improve quality of trait data integration through inferring new facts through machine reasoning (see Box 3 for examples). This process converts trait datasets into fully interoperable formats and enables future researchers as well as machines to interpret the data.
Providing integrated and reusable trait data products via web services
To make an integrated trait data product FAIR18 (see above), a public domain designation (for example, CC0) or an open access licence (for example, CC BY) should be applied and provided together with other metadata in a machine-readable format (Fig. 3, bottom). In the best case, licence information should be available for each trait record and original source (Box 3). Further, it is important that data structures of trait data products align with semantic web standards (for example, multi-layered, relational databases rather than two-dimensional data tables). Hence, trait data products should be housed with a graph database that allows on-the-fly reasoning via semantic queries, or with relational database if on-the-fly reasoning is not needed (Fig. 3, bottom). In both cases, an application programming interface (API) should allow communication and access to the trait data product via a web platform (Box 3) or via widely used software such as R or Python (Fig. 3, bottom).
Towards operationalizing species traits EBVs
Species traits are a key component of biodiversity, but species trait information is currently not well represented in indicators of biodiversity change used for national and international policy assessments2,17,74. The increasing willingness to share trait data in an open and machine-readable way (see Supplementary Note 3), coupled with emerging semantic tools (for example, new plant trait vocabularies11, ontologies64,73 and preliminary suggestions for trait data standards27) and a massive collection of trait data through in situ monitoring schemes and close-range sensors (for example, for phenology20,39,47,75) as well as on-going and forthcoming airborne and spaceborne missions (including radar, optical sensors, radiometers and spectrometers42,43,50,53,76), suggest that comprehensive data products on species traits are within reach in the near future. However, a cultural shift towards more openness, interoperability and reproducibility is needed within the broader science community18,19,77 — including ecologists, biogeographers, global change biologists, biodiversity informaticians and Earth scientists — and with support from global coordinating institutions such as GEO BON, IPBES and the CBD.
Our refined list of species traits EBVs (Fig. 1) provides an improved conceptual framework for how phenological, morphological, reproductive, physiological and movement-related trait measurements can represent biodiversity in the EBV context and hence support international policies for biodiversity conservation and sustainable development. The specific species traits EBVs contain essential information with ecological, societal and policy relevance for biodiversity that cannot be substituted by other species traits EBVs (Supplementary Note 2). For instance, morphological and physiological measurements of leaves (for example, leaf area, nitrogen and chlorophyll content), stems (for example, height and stem density) and diaspores (for example, seed mass) allow quantification of fundamental dimensions of plant ecological strategies and how these organisms respond to competition, stress, environmental change and disturbances8,12,43,50. Phenological trait information of amphibians (spawning), birds (egg laying), plankton (population peaks), fish (spawning), insects (flight periods), mammals (birth dates) and plants (flowering, fruiting, leafing) is highly relevant for tracking changes in species’ ecology in response to climate change21 and other global changes (for example, nitrogen deposition inducing delayed foliar senescence). Morphological measurements (body sizes) of commercially relevant fish species78,79,80 can allow assessments of sustainable food production and harvesting (Box 1). Similarly, morphological, reproductive and physiological traits of microbial species (for example, cell size, lifetime pattern of growth and microbial resistance to viruses) are essential for predicting their responses to environmental change81. A key aspect for the future operationalization of species traits EBVs is that they should be measurable with available technologies and have a proven track record of feasibility6. We suggest that a focus on trait measurements representing plant phenology, morphology and physiology (for example, from both in situ monitoring20,39,47,75 and remote sensing9,12,42,43,49,50,82) as well as animal morphology15,79 and movement83 could provide a realistic prioritization for operationalizing species traits EBVs.
Compiling the necessary data for EBVs globally remains a major challenge, especially for species traits7,17. A key bottleneck is that the repeated and systematic collection of in situ trait data is not only costly and difficult but also spatially discontinuous. The global, spatially contiguous and periodic nature of spaceborne remote sensing observations therefore offers potential for building EBVs82. To date, spaceborne remote sensing products (for example, related to land surface phenology, canopy biochemistry and vegetation height) allow the mapping of ecosystem structure and processes as well as functional diversity9,43,51,84, but not the quantification of species-level traits1,82 because the spatial resolution is not fine enough to allow attribution of trait measurements to an individual or a population of a single species (Box 1). With airborne remote sensing it is possible to continuously map individual-level trait variation in morphological and physiological traits at fine (metre) resolution across regional scales (for example, forest trees43), often allowing assignment of trait measurements to the species level85,86. Since species-level resolution is required for many policy targets76, assigning trait measurements to taxonomic information is key for monitoring intra-specific trait changes. A deeper integration of in situ and various close-range remote sensing trait measurements as well as a synergy of hyperspectral and LiDAR airborne remote sensing might help to achieve this. An avenue for building contiguous species traits EBVs could be to use information from Earth observation data for interpolating in situ trait point samples for building continuous landscape maps of trait distributions76. This would require the development of statistical and mechanistic models that allow mapping and prediction of trait distributions across space and time87. In this context, specimens from natural history collections could become useful for obtaining baseline trait data for regions that have been poorly studied88.
Moving forward
Many dimensions of biodiversity still remain invisible when measuring and monitoring global biodiversity change2,17,76. Species traits EBVs will provide a deeper understanding of the species-level responses to global change and the benefits and services that individual species provide to humanity. For operationalizing species traits EBVs, we recommend the biodiversity research community to support trait data harmonization, reproducible workflows, interoperability and ‘big data’ biodiversity informatics for species traits19,23,27,89,90. Specifically, we suggest the following concrete steps to facilitate the building of EBV data products of species traits:
-
Support the recording of species traits across time through repeated and periodic collection of in situ measurements of traits, through digitization of trait information from literature and biocollections and through developing species traits data products from close-range, airborne and spaceborne remote sensing observations.
-
Develop and apply standardized protocols, controlled trait vocabularies and trait data standards when measuring, harmonizing and combining trait data and metadata.
-
Support the semantic integration of trait data by mapping trait datasets to ontologies, facilitate training courses about semantic standards of the World Wide Web Consortium (W3C) and promote training tools for trait data integration within research institutions and educational programmes of universities.
-
Publish trait databases with standardized licence information in machine-readable form and designate data as open access (for example, through CC BY) or in the public domain (for example, CC0). Encourage others to share trait data.
-
Develop and apply reproducible statistical and mechanistic models for integrating in situ trait data with remote sensing observations to allow mapping and prediction of trait distributions across space and time.
-
Establish consortia and interest groups on species traits. Contribute to the GEO BON working group on species traits and raise awareness of the need for semantic, technical and legal interoperability of trait data.
-
Foster the integration of species traits EBVs into biodiversity indicators and biodiversity and sustainability goals.
These activities — which require substantial financial and in kind investments from universities, research infrastructures, governments, space agencies and other bodies — will facilitate the building of global EBV data products of species traits and allow significant steps towards incorporating intra-specific trait variability into global, regional and national biodiversity and policy assessments.
References
Pereira, H. M. et al. Essential Biodiversity Variables. Science 339, 277–278 (2013).
Geijzendorffer, I. R. et al. Bridging the gap between biodiversity data and policy reporting needs: An Essential Biodiversity Variables perspective. J. Appl. Ecol. 53, 1341–1350 (2016).
Kissling, W. D. et al. Building essential biodiversity variables (EBVs) of species distribution and abundance at a global scale. Biol. Rev. 93, 600–625 (2018).
Giuliani, G. et al. Building an Earth Observations Data Cube: lessons learned from the Swiss Data Cube (SDC) on generating Analysis Ready Data (ARD). Big Earth Data 1, 100–117 (2017).
Schmeller, D. S. et al. An operational definition of essential biodiversity variables. Biodivers. Conserv. 26, 2967–2972 (2017).
Walters, M. et al. Essential Biodiversity Variables (Convention on Biological Diversity, 2013).
GEO BON Implementation Plan 2017–2020 Version 1.3 (GEO BON Secretariat, 2017).
Lavorel, S. & Garnier, E. Predicting changes in community composition and ecosystem functioning from plant traits: revisiting the Holy Grail. Funct. Ecol. 16, 545–556 (2002).
Lausch, A. et al. Linking Earth observation and taxonomic, structural and functional biodiversity: local to ecosystem perspectives. Ecol. Indic. 70, 317–339 (2016).
Gibert, J. P., Dell, A. I., DeLong, J. P. & Pawar, S. in Advances in Ecological Research Vol. 52 (eds Woodward, G., Pawar, S. & Dell, A. I.) 1–17 (Academic, Waltham, 2015).
Garnier, E. et al. Towards a thesaurus of plant characteristics: an ecological contribution. J. Ecol. 105, 298–309 (2017).
Díaz, S. et al. The global spectrum of plant form and function. Nature 529, 167–171 (2016).
Eskildsen, A. et al. Ecological specialization matters: long-term trends in butterfly species richness and assemblage composition depend on multiple functional traits. Divers. Distrib. 21, 792–802 (2015).
Hagen, M. et al. Biodiversity, species interactions and ecological networks in a fragmented world. Adv. Ecol. Res. 46, 89–210 (2012).
Genner, M. J. et al. Body size-dependent responses of a marine fish assemblage to climate change and fishing over a century-long scale. Glob. Change Biol. 16, 517–527 (2010).
Wilson, J. R. U., Dormontt, E. E., Prentis, P. J., Lowe, A. J. & Richardson, D. M. Something in the way you move: dispersal pathways affect invasion success. Trends Ecol. Evol. 24, 136–144 (2009).
Navarro, L. M. et al. Monitoring biodiversity change through effective global coordination. Curr. Opin. Environ. Sustain. 29, 158–169 (2017).
Wilkinson, M. D. et al. The FAIR Guiding Principles for scientific data management and stewardship. Sci. Data 3, 160018 (2016).
Kissling, W. D. et al. Towards global interoperability for supporting biodiversity research on essential biodiversity variables (EBVs). Biodiversity 16, 99–107 (2015).
Schwartz, M. D., Betancourt, J. L. & Weltzin, J. F. From Caprio’s lilacs to the USA National Phenology Network. Front. Ecol. Environ. 10, 324–327 (2012).
Thackeray, S. J. et al. Phenological sensitivity to climate across taxa and trophic levels. Nature 535, 241–245 (2016).
Kattge, J. et al. TRY – a global database of plant traits. Glob. Change Biol. 17, 2905–2935 (2011).
Parr, C. S. et al. TraitBank: practical semantics for organism attribute data. Semant. Web 7, 577–588 (2016).
Calewaert, J.-B., Weaver, P., Gunn, V., Gorringe, P. & Novellino, A. in Quantitative Monitoring of the Underwater Environment (eds Zerr, B. et al.) 31–46 (Springer, Cham, 2016).
Guralnick, R. P. et al. The importance of digitized biocollections as a source of trait data and a new VertNet resource. Database 2016, baw158 (2016).
MarLIN BIOTIC: Biological Traits Information Catalogue (Marine Biological Association of the United Kingdom, 2006).
Schneider, F. D. et al. Towards an ecological trait-data standard. Preprint at https://www.biorxiv.org/content/early/2018/05/31/328302 (2018).
Kissling, W. D. et al. Establishing macroecological trait datasets: digitalization, extrapolation, and validation of diet preferences in terrestrial mammals worldwide. Ecol. Evol. 4, 2913–2930 (2014).
Holmes, M. W. et al. Natural history collections as windows on evolutionary processes. Mol. Ecol. 25, 864–881 (2016).
Gardner, J. L. et al. Temporal patterns of avian body size reflect linear size responses to broadscale environmental change over the last 50 years. J. Avian Biol. 45, 529–535 (2014).
Tseng, M. et al. Decreases in beetle body size linked to climate change and warming temperatures. J. Anim. Ecol. 87, 647–659 (2018).
Miller-Rushing, A. J., Primack, R. B., Primack, D. & Mukunda, S. Photographs and herbarium specimens as tools to document phenological changes in response to global warming. Am. J. Bot. 93, 1667–1674 (2006).
Robbirt, K. M., Davy, A. J., Hutchings, M. J. & Roberts, D. L. Validation of biological collections as a source of phenological data for use in climate change studies: a case study with the orchid Ophrys sphegodes. J. Ecol. 99, 235–241 (2011).
Willis, C. G. et al. Old plants, new tricks: phenological research using herbarium specimens. Trends Ecol. Evol. 32, 531–546 (2017).
Kim, E.-S. Development, potentials, and challenges of the International Long-Term Ecological Research (ILTER) Network. Ecol. Res. 21, 788–793 (2006).
Gosz, J. R., Waide, R. B. & Magnuson, J. J. in Long-Term Ecological Research: Between Theory and Application (eds Müller, F., Baessler, C., Schubert, H. & Klotz, S.) 59–74 (Springer, Dordrecht, 2010).
Silvertown, J. A new dawn for citizen science. Trends Ecol. Evol. 24, 467–471 (2009).
Jackson, M. C. et al. in Advances in Ecological Research Vol. 55 (eds Dumbrell, A. J., Kordas, R. L. & Woodward, G.) 615–636 (Academic, Waltham, 2016).
Elmendorf, S. C. et al. The plant phenology monitoring design for the National Ecological Observatory Network. Ecosphere 7, e01303 (2016).
Metzger, M. J. et al. Environmental stratifications as the basis for national, European and global ecological monitoring. Ecol. Indic. 33, 26–35 (2013).
Pettorelli, N. et al. Framing the concept of satellite remote sensing essential biodiversity variables: challenges and future directions. Remote Sens. Ecol. Conserv. 2, 122–131 (2016).
Skidmore, A. K. et al. Agree on biodiversity metrics to track from space. Nature 523, 403–405 (2015).
Schneider, F. D. et al. Mapping functional diversity from remotely sensed morphological and physiological forest traits. Nat. Commun. 8, 1441 (2017).
Ashburner, M. et al. Gene Ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Zhu, X. et al. Canopy leaf water content estimated using terrestrial LiDAR. Agr. Forest Meteorol. 232, 152–162 (2017).
Richardson, A. D., Klosterman, S. & Toomey, M. in Phenology: An Integrative Environmental Science (ed Schwartz, M. D.) 413–430 (Springer, Dordrecht, 2013).
Nasahara, K. N. & Nagai, S. Development of an in situ observation network for terrestrial ecological remote sensing: the Phenological Eyes Network (PEN). Ecol. Res. 30, 211–223 (2015).
Schweiger, A. K. et al. Plant spectral diversity integrates functional and phylogenetic components of biodiversity and predicts ecosystem function. Nat. Ecol. Evol. 2, 976–982 (2018).
Kampe, T. U., Johnson, B. R., Kuester, M. & Keller, M. NEON: the first continental-scale ecological observatory with airborne remote sensing of vegetation canopy biochemistry and structure. J. Appl. Remote Sens. 4, 043510 (2010).
Asner, G. P. et al. Airborne laser-guided imaging spectroscopy to map forest trait diversity and guide conservation. Science 355, 385–389 (2017).
Jetz, W. et al. Monitoring plant functional diversity from space. Nat. Plants 2, 16024 (2016).
Schimel, D. S., Asner, G. P. & Moorcroft, P. Observing changing ecological diversity in the Anthropocene. Front. Ecol. Environ. 11, 129–137 (2013).
Stavros, E. N. et al. ISS observations offer insights into plant function. Nat. Ecol. Evol. 1, 0194 (2017).
Asner, G. P. et al. Amazonian functional diversity from forest canopy chemical assembly. Proc. Natl Acad. Sci. USA 111, 5604–5609 (2014).
Asner, G. P. & Martin, R. E. Airborne spectranomics: mapping canopy chemical and taxonomic diversity in tropical forests. Front. Ecol. Environ. 7, 269–276 (2009).
Cornelissen, J. H. C. et al. A handbook of protocols for standardised and easy measurement of plant functional traits worldwide. Aust. J. Bot. 51, 335–380 (2003).
Pérez-Harguindeguy, N. et al. New handbook for standardised measurement of plant functional traits worldwide. Aust. J. Bot. 61, 167–234 (2013).
Moretti, M. et al. Handbook of protocols for standardized measurement of terrestrial invertebrate functional traits. Funct. Ecol. 31, 558–567 (2017).
Salgado-Negret, B. La Ecología Funcional Como Aproximación al Estudio, Manejo y Conservación de la Biodiversidad: Protocolos y Aplicaciones (Instituto de Investigación de Recursos Biológicos Alexander von Humboldt, 2015).
Costello, M. J. et al. Biological and ecological traits of marine species. PeerJ 3, e1201 (2015).
Park, C. A. et al. The Vertebrate Trait Ontology: a controlled vocabulary for the annotation of trait data across species. J. Biomed. Semant. 4, 13 (2013).
Baird, D. J. et al. Toward a knowledge infrastructure for traits‐based ecological risk assessment. Integr. Environ. Assess. Manag. 7, 209–215 (2011).
Mungall, C. J. et al. Integrating phenotype ontologies across multiple species. Genome Biol. 11, R2 (2010).
Hoehndorf, R. et al. The flora phenotype ontology (FLOPO): tool for integrating morphological traits and phenotypes of vascular plants. J. Biomed. Semant. 7, 65 (2016).
Walls, R. L. et al. Ontologies as integrative tools for plant science. Am. J. Bot. 99, 1263–1275 (2012).
Hughes, L. M., Bao, J., Hu, Z.-L., Honavar, V. & Reecy, J. M. Animal trait ontology: the importance and usefulness of a unified trait vocabulary for animal species. J. Anim. Sci. 86, 1485–1491 (2008).
Kattge, J. et al. A generic structure for plant trait databases. Methods Ecol. Evol. 2, 202–213 (2011).
Costello, M. J., Michener, W. K., Gahegan, M., Zhang, Z.-Q. & Bourne, P. E. Biodiversity data should be published, cited, and peer reviewed. Trends Ecol. Evol. 28, 454–461 (2013).
Penev, L. et al. Strategies and guidelines for scholarly publishing of biodiversity data. Res. Ideas Outcomes 3, e12431 (2017).
Michener, W. K. Ecological data sharing. Ecol. Inform. 29, 33–44 (2015).
RDA-CODATA Legal Interoperability Interest Group Legal Interoperability of Research Data: Principles and Implementation Guidelines (Research Data Alliance, 2016).
Carroll, M. W. Creative commons and the new intermediaries. Michigan State Law Rev. 2006, 45–65 (2006).
Stucky, B. J. et al. The Plant Phenology Ontology: a new informatics resource for large-scale integration of plant phenology data. Front. Plant Sci. 9, 517 (2018).
Global Biodiversity Change Indicators: Model-Based Integration of Remote-Sensing and In Situ Observations that Enables Dynamic Updates and Transparency at Low Cost (GEO BON Secretariat, 2015).
van Vliet, A. J. H. et al. The European Phenology Network. Int. J. Biometeorol. 47, 202–212 (2003).
Bush, A. et al. Connecting Earth observation to high-throughput biodiversity data. Nat. Ecol. Evol. 1, 0176 (2017).
Lowndes, J. S. S. et al. Our path to better science in less time using open data science tools. Nat. Ecol. Evol. 1, 0160 (2017).
Worm, B. et al. Rebuilding global fisheries. Science 325, 578–585 (2009).
Coll, M. et al. Ecological indicators to capture the effects of fishing on biodiversity and conservation status of marine ecosystems. Ecol. Indic. 60, 947–962 (2016).
Welcomme, R. L. et al. Inland capture fisheries. Phil. Trans. R. Soc. B 365, 2881–2896 (2010).
Krause, S. et al. Trait-based approaches for understanding microbial biodiversity and ecosystem functioning. Front. Microbiol. 5, 251 (2014).
Skidmore, A. K. et al. Remote Sensing Enabled Essential Biodiversity Variables (Convention on Biological Diversity, 2018).
Kays, R., Crofoot, M. C., Jetz, W. & Wikelski, M. Terrestrial animal tracking as an eye on life and planet. Science 348, aaa2478 (2015).
Houborg, R., Fisher, J. B. & Skidmore, A. K. Advances in remote sensing of vegetation function and traits. Int. J. Appl. Earth Obs. Geoinform. 43, 1–6 (2015).
Asner, G. P. et al. Invasive species detection in Hawaiian rainforests using airborne imaging spectroscopy and LiDAR. Remote Sens. Environ. 112, 1942–1955 (2008).
Skidmore, A. K. et al. Forage quality of savannas — Simultaneously mapping foliar protein and polyphenols for trees and grass using hyperspectral imagery. Remote Sens. Environ. 114, 64–72 (2010).
Butler, E. E. et al. Mapping local and global variability in plant trait distributions. Proc. Natl Acad. Sci. USA 114, E10937–E10946 (2017).
Greve, M. et al. Realising the potential of herbarium records for conservation biology. S. Afr. J. Bot. 105, 317–323 (2016).
La Salle, J., Williams, K. J. & Moritz, C. Biodiversity analysis in the digital era. Phil. Trans. R. Soc. B 371, 20150337 (2016).
Merchant, N. et al. The iPlant collaborative: cyberinfrastructure for enabling data to discovery for the life sciences. PLoS Biol. 14, e1002342 (2016).
Violle, C. et al. Let the concept of trait be functional! Oikos 116, 882–892 (2007).
Lausch, A., Erasmi, S., King, D., Magdon, P. & Heurich, M. Understanding forest health with remote sensing -part I—a review of spectral traits, processes and remote-sensing characteristics. Remote Sens. 8, 1029 (2016).
Galetti, M. et al. Functional extinction of birds drives rapid evolutionary changes in seed size. Science 340, 1086–1090 (2013).
Madin, J. S., Bowers, S., Schildhauer, M. P. & Jones, M. B. Advancing ecological research with ontologies. Trends Ecol. Evol. 23, 159–168 (2008).
Walls, R. L. et al. Semantics in support of biodiversity knowledge discovery: an introduction to the Biological Collections Ontology and related ontologies. PLoS ONE 9, e89606 (2014).
Bruskiewich, R. M. et al. Linking genotype to phenotype: the International Rice Information System (IRIS). Bioinformatics 19, i63–i65 (2003).
Smith, C. L. & Eppig, J. T. The mammalian phenotype ontology: enabling robust annotation and comparative analysis. Wiley Interdisc. Rev. Syst. Biol. Med. 1, 390–399 (2009).
Buttigieg, P. L. et al. The environment ontology in 2016: bridging domains with increased scope, semantic density, and interoperation. J. Biomed. Semant. 7, 1–12 (2016).
Schimel, D., Hargrove, W., Hoffman, F. & MacMahon, J. NEON: a hierarchically designed national ecological network. Front. Ecol. Environ. 5, 59 (2007).
Cooper, L. & Jaiswal, P. in Plant Bioinformatics: Methods and Protocols (ed. Edwards, D.) 89–114 (Springer, New York, 2016).
Acknowledgements
This paper emerged from a workshop of the Horizon 2020 project GLOBIS-B (Global Infrastructures for Supporting Biodiversity research; http://www.globisb.eu/). Financial support came from the European Commission (grant 654003). We thank J. Konijn for administrative support and the members of the BIOMAC lab (https://www.biomac.org/) for discussion. L.M.N. is supported by the German Centre for Integrative Biodiversity Research (iDiv) Halle-Jena-Leipzig funded by the German Research Foundation (FZT 118). N.R. was funded by a research grant from Deutsche Forschungsgemeinschaft DFG (RU 1536/3-1).
Author information
Authors and Affiliations
Contributions
W.D.K. coordinated the study and wrote the draft manuscripts. R.W., A.B., M.O.J., J.K., R.P.G. and W.D.K. led the writing of sections. All authors provided substantial input into ideas and text, and commented on draft manuscripts.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kissling, W.D., Walls, R., Bowser, A. et al. Towards global data products of Essential Biodiversity Variables on species traits. Nat Ecol Evol 2, 1531–1540 (2018). https://doi.org/10.1038/s41559-018-0667-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-018-0667-3