Abstract
Accelerating international trade and climate change make pathogen spread an increasing concern. Hymenoscyphus fraxineus, the causal agent of ash dieback, is a fungal pathogen that has been moving across continents and hosts from Asian to European ash. Most European common ash trees (Fraxinus excelsior) are highly susceptible to H. fraxineus, although a minority (~5%) have partial resistance to dieback. Here, we assemble and annotate a H. fraxineus draft genome, which approaches chromosome scale. Pathogen genetic diversity across Europe and in Japan, reveals a strong bottleneck in Europe, though a signal of adaptive diversity remains in key host interaction genes. We find that the European population was founded by two divergent haploid individuals. Divergence between these haplotypes represents the ancestral polymorphism within a large source population. Subsequent introduction from this source would greatly increase adaptive potential of the pathogen. Thus, further introgression of H. fraxineus into Europe represents a potential threat and Europe-wide biological security measures are needed to manage this disease.
Similar content being viewed by others
Main
Human movement and climate change may increase the incidence of plant pathogen introductions as new environments become favourable1. Dutch elm disease is one example of a pathogen epidemic that emerged and caused the loss of billions of elm trees globally2. Presently, the ascomycete fungus Hymenoscyphus fraxineus is causing disease and death in European common ash, Fraxinus excelsior, and narrow-leaved ash, F. angustifolia. H. fraxineus jumped host from Asian ash species where it is a leaf pathogen with little impact on its host. In Europe it is killing ash at an alarming rate and displacing the non-aggressive indigenous fungus, H. albidus3.
The disease was first observed in Europe in north-western Poland in 1992 and, moving west, was identified in the UK in 20123. Less than 5% of trees are partially resistant or tolerant4,5 to ash dieback disease, which is characterized by dark brown/orange lesions on leaves followed by wilting, necrotic lesions on shoots, then diamond-shaped lesions on the stems and finally dieback of the crown6 (Fig. 1). The loss of leaves in the crown of mature trees proceeds over years and leads, in severe cases, to tree death. H. fraxineus is a heterothallic fungus7 that has been shown to reproduce asexually in vitro8, but in the wild, sexual reproduction occurs annually on fallen leaf rachises in the leaf litter6 (Fig. 1).
a,b, Brown/orange lesions on leaves (a) followed by wilting (b). c, Disease progression: lesions on shoots followed by diamond-shaped lesions on the stems. d,e, Wild, sexual reproduction occurs annually on fallen leaf rachises in the leaf litter. f, Dieback of the crown6.
A pathogen’s evolutionary potential is rooted in its genetic diversity (or its effective population size)9. All else being equal, large populations can adapt more quickly than small ones for two reasons; first, a large population carries more polymorphism and so there is a greater chance that a favourable mutation is segregating. Second, the impact of random genetic drift is lower for larger populations and therefore, natural selection is better able to drive those favourable mutations to fixation10. However, pathogen introductions are often associated with genetic bottlenecks, or founder effects, which reduce the level of genetic diversity and the efficacy of selection. This disparity between reduced polymorphism and invasion success is known as the genetic paradox of biological invasions11. The potato late blight pathogen, Phytophthora infestans, is another example of a successful bottlenecked pathogen introduction12. However devastating a pathogen may be, multiple introductions can increase diversity above that of native populations11. Early P. infestans invasions were dominated by a single clonal lineage (US-1), which was later superseded as diversity and severity increased12. The Dutch elm disease pandemic(s) also had an initial rapid spread of Ophiostoma ulmi which was later replaced by O. novo-ulmi (also subspecies americana), and went on to devastate elm populations across two continents2. The amount of H. fraxineus genetic variation introduced to Europe, and the potential for further introduction, is therefore of critical concern.
Estimates of H. fraxineus microsatellite allelic richness suggest as few as two haploid individuals may have invaded Europe13. Interestingly, the viral pathogen of H. fraxineus (mitovirus 1) also has two genetic groups present in the European population14. Given the impact of the pathogen invasion so far, a founder effect of just two haploid individuals would represent a genetic paradox and so it is important to measure genetic diversity across the genome as well as meaningful, adaptive diversity in the host interaction genes (effectors). Determining the genetic diversity present in the native range is key to understanding the consequences of any further introductions. Here, we assembled and annotated a high-quality draft genome of H. fraxineus. We quantified the level of genetic variation in 43 H. fraxineus haploid isolates from across Europe as well as 15 haplotypes from part of its native range (Japan). To understand the adaptive potential of the ash dieback pathogen in Europe as well as the potential of future invasions, we determined the effective population size, the bottleneck into Europe and estimated the size of the source population. In addition, we mined putative effector genes from the genome of H. fraxineus. These genes encode secreted proteins considered to play a key role in establishing fungal infections and facilitating disease development15. We measured signals of adaptive diversity in the putative effectors and all other genes. The proportion of introduced adaptive variation and the potential for further introduction are a key focus of the present work.
Results and discussion
Population genetic diversity of the emerging invasive H. fraxineus pathogen measured among 43 isolates from across Europe is an eighth of that observed in 15 haplotypes from a single wood in Japan. This general reduction in polymorphism, caused by a bottleneck of two divergent individuals, could reflect a reduction in the pathogen’s adaptive potential in its introduced environment. This reduction in genetic diversity is present in effectors as well as all other genes. Effectors from European isolates retain a greater degree of adaptive variation but far less than in the native range and we discuss the implications for the predicted level of virulence in Europe over the long term.
H. fraxineus version 2 genome approaches chromosome-level contiguity
The H. fraxineus version 2 genome assembly (Hf-v2.0) is 62.28 Mbp distributed across 145 scaffolds. Here, we analyse scaffolds greater than 10 kbp, a total of 23 scaffolds or 62.08 Mbp (Fig. 2; Supplementary Analysis 2). Both Hf-v1.116 and Hf-v2.0 have been released (open resources) online and are hosted on the Open Ash Dieback github repository (https://github.com/ash-dieback-crowdsource). The Hf-v2.0 gene annotation pipeline built on that of Hf-v1.116 and identified 10,945 genes (11,097 transcripts, 1,516.50 mean CDS length, 3.49 mean exons per gene; Supplementary Analysis 3). Telomeric repeats are identified at both ends of 14 scaffolds and at one end of all but one of the remaining scaffolds (Supplementary Table 2). Genetic linkage between scaffolds in the progeny of a H. fraxineus lab cross suggests linkage between three pairs of scaffolds missing a telomere (Supplementary analysis 4). By joining these linked scaffolds, we estimate 20 chromosomes (pseudomolecules) for H. fraxineus (Fig. 2b).
a, AT richness sliding window (AT scale = 0.5–0.85; window 63 kbp by 5 kbp slide) with outer ticks indicating gaps in the assembly >2.5 kbp. b, Scaffolds (ticks = 100 kbp) red, green and yellow pairs of scaffolds appear to be from the same linkage group. c, Repeat density (0.0–1.0; 100 kbp). d,e, Gene (blue) and effector (red) density, respectively (0.0–1.0; 100 kbp). f, Genetic differentiation (FST) sliding window between Japanese and European populations (0.01–0.90; 100 kbp by 5 kbp). g, Nucleotide diversity (π) sliding window within Japanese (red) and European (yellow) populations (0.0–0.032; 100 kbp by 5 kbp). h, SNP density sliding window within Japanese (red) and European (yellow) populations (0.0–0.17; 100 kbp by 5 kbp). Key shows track reference and orientation.
The European invasion population is bottlenecked and sexually recombining diversity from Asia
In 43 European and 15 Japanese haplotypes, we identified 6.26 million single nucleotide polymorphisms (SNPs) overall (SNPs segregating in: Japan = 4.5 × 106, Europe = 0.67 × 106; Supplementary Analysis 5). An SNP network of all genes shows us that the European population is genetically divergent and bottlenecked from the native population (Fig. 3). The disparity in nucleotide diversity (π) between Europe and Japan (Fig. 3) could have resulted from a source population much smaller than that of Japan. Tajima’s D17 is a statistic, centred around zero, that is sensitive to changes in effective population size (and/or the mode of selection). In Japan, we observe a Tajima’s D value close to zero (\({\bar{x}}\) = −0.22), which indicates neutrality with purifying selection operating on genes (\({\bar{x}}\) = −0.89; Fig. 3). In Europe, the genomic signal is broadly positive (\({\bar{x}}\) = 1.28; Fig. 3). This positive value is generated by a contraction in population size balancing allele frequencies by reducing the frequency of common alleles without equivalent reduction in the frequency of rarer alleles (see Supplementary Fig. 10). This balancing of allele frequencies without a real reduction in SNP density is consistent with a small founder European population from a larger source.
a,b, European (a) and Japanese (b) sampling locations (to scale). c, A neighbour-net network of the concatenated coding regions of all genes in the genome (16.6 Mbp) for all European (n = 43) and Japanese haplotypes (n = 15). d, Boxplot of nucleotide diversity (π per 100 kbp) for 642 windows in Japanese (Jp) and European (Eu) populations. Nucleotide diversity is significantly greater in Japan than in Europe (Wilcoxon (π), Jp genome > Eu genome: W(1257) = 393,210, n = 1,258, ***P < 0.001). e, Boxplot of Tajima’s D (D per 100 kbp) for 642 windows across the genome and for each polymorphic gene within Japanese (n = 10,869) and European (n = 6,559) populations. In Japan, a signal of neutrality is present across the genome and tends to be negative in genes (purifying selection). The European population has a much broader positive distribution, consistent with a recent population decline. Boxplots show the median, upper and lower quartiles with outliers plotted outside whiskers which extend 1.5 times beyond the interquartile range.
Sex is a fundamental determinant of plant pathogen adaptive potential not least because it allows recombination of host interaction genes, or effectors18. Here, as well as observing sexual reproduction between European isolates in the lab (Supplementary Analysis 4) we see a breakdown in linkage disequilibrium in the wild in Europe (Supplementary Analysis 6). At the genome level Japanese and European populations are divergent (FST = 0.55 (95% confidence interval = 0.54–0.56)) but recombination decouples relationships between genes. Across the genome, genetic differentiation correlates positively with diversity present in the Japanese population (R2 = 0.18, n = 627, P < 0.001) but negatively with diversity present in the European population (R2 = 0.51, n = 610, P < 0.001). That is to say that, the more diversity present at a locus in Japan, the less likely it is to be shared with Europe and, the more diversity present at a locus in Europe the more likely it is to be shared with Japan; a signal consistent with directionality from east to west (Fig. 2fg, Supplementary Fig. 11). Preventing continued gene flow from the native range into Europe is important because sexual reproduction between divergent demes may be important for plant pathogen adaptation19.
The European population was founded by two individuals from a large diverse population
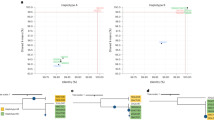
A previous study20 reported similar levels of genetic diversity (based on 11 microsatellite loci) between central European populations and those on the epidemic disease front. The authors suggest that for introduced diversity to reach the epidemic disease front, either the centre of genetic diversity quickly recombined and spread by range expansion, or only a small number of divergent H. fraxineus isolates arrived in Europe and present-day diversity is the product of recombination among those. Here, we use third-base positions of the coding regions of core eukaryotic genes (CEGs) to distinguish between these scenarios and describe the invasion process. CEGs are highly conserved essential proteins, encoded in all eukaryotic genomes21. Their importance to fundamental eukaryotic biology makes the 387 CEGs we identified in all H. fraxineus individuals an ideal set to explore the European invasion. The bottleneck into Europe reduced the number of CEG haplotypes to 2.3 with 42% of all CEGs being monomorphic. The average number of Japanese haplotypes was 12.6 with no less than two per locus. Strikingly, European CEGs grouped into two divergent haplotypes. Furthermore, the level of divergence between those haplotypes reflects that across Japanese CEG networks (Fig. 4, Supplementary Fig. 20). It is our interpretation that these divergent haplotype pairs have been introduced by two haploid founders, as suggested previously13. Moreover, the divergence between these European haplotype pairs, without intermediates, represents the ancestral divergence of a large population from which the European population was founded.
Neighbour-net networks were generated individually for the third-base position of 387 CEGs. a,b, A selection of networks built using three individual core genes are shown separately from European (a) and Japanese (b) H. fraxineus populations (see Supplementary Fig. 20 for combined networks). Encircled numbers are used in a to indicate numbers of individuals that share each haplotype. The three gene networks from the European population in a are the same genes as those from the Japanese population in b and drawn to the same scale. c,d, Density plots show the relative pairwise distance amongst haplotypes of all 387 CEGs in Europe (c) and Japan (d). In Japan, pairwise distances between haplotypes range in their divergence (d). In Europe, the majority of genes are either identical or at the complete opposite ends of the divergence range (c). The high level of divergence that separates European haplotypes, which are shared by many individuals, is visible in the three gene networks (a) but also in the measure of pairwise haplotype divergence of all 387 CEGs (c). This represents the presence of two major divergent haplotype groups in Europe.
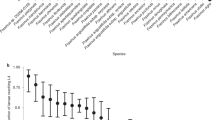
The observed estimate for the haploid effective population size (ϑπ = 2Neμ) in Japan is 2.46 million individuals (π = 0.0246) and the estimate from Europe is 0.67 million individuals (π = 0.00672; μ = 5 × 10−9 per base per generation). However, this European estimate of nucleotide diversity is inflated by the ancestral divergence of the two haploid founders. We used this divergent polymorphism to estimate the size of the European source population. Coalescence simulations of a hypothetical European source, bottlenecked to two individuals, show that as the source effective population size increases, so too does the average divergence between founder haplotypes. We find that a source effective population size between 2.2 and 2.8 million individuals (median 2.5 million) most accurately replicates the observed European haplotype divergence (Fig. 5). Our estimate of the size of the source of the European H. fraxineus population is therefore equivalent to that from a single Japanese wood. These estimates may reflect the equilibrium diversity in any given H. fraxineus population within its native range, but importantly also suggests that the European invaders could have come from a single site, perhaps even from a single ascocarp (fruiting body).
The coalescence of the third base pair of 387 core eukaryotic genes in a population was simulated (×1,000) at an effective population size of between 1 million and 4 million haploid individuals. This (equilibrium) population was bottlenecked to two individuals and the diversity at all polymorphic loci was recorded. 2.5 million individuals best accounted for the observed diversity in present day European divergent haplotypes (observed SNPs/bp = 0.0305). Plot shows median and 95% confidence intervals.
10% of the H. fraxineus proteome putatively interacts with the host
Effectors are a broad classification of proteins that are secreted by bacteria, oomycetes, fungi, nematodes and aphids in order to interact with the host, disable host defence components and facilitate colonization15. As such, effectors are in a co-evolutionary arms race with host resistance genes and are studied for insights into pathogen adaptive potential22. We used localization signals of N-terminal presequences to identify secreted proteins in the H. fraxineus genome (Fig. 2e). These 1,132 predicted secreted proteins are potential effectors, which clustered into 566 tribes (Supplementary Table 5). We did not identify any conserved fungal N-terminal signal motifs. A reduced set of 223 putative effectors were identified using a machine-learning approach (Supplementary Analysis 7). Nevertheless, we consider the broadest set of secreted proteins in subsequent analyses to avoid missing out on potentially important effectors.
Putative effector genes are spread across the genome, away from repeat regions, similar to non-effector genes (Fig. 2c–e; Supplementary Figs. 21,22) which contrasts with predictions of the two-speed genome model, in other filamentous plant pathogens23 (for example, Sclerotinia sclerotiorum24). Absence of the two-speed evolution model has been observed in the genomic landscapes of members of the Magnaporthaceae family25. Perhaps, in H. fraxineus, the requirement for a dynamic process that operates to shuffle and generate novel effectors is lower in this sexually reproducing pathogen with high haplotype diversity26. The presence of sex and high haplotype diversity in turn indicate a long-term balanced relationship between host and pathogen within the native range27
Structure and function of H. fraxineus putative effectors
Of the 1,132 predicted H. fraxineus secreted proteins, 62% carry Pfam domains associated with highly diverse biological functions (Supplementary Table 6). The largest subgroups include apoplastic catalytic enzymes, with 127 proteins predicted to have glycosyl hydrolase activity. Amongst these are putative cell-wall-degrading enzymes such as cellulases, pectinoesterases and cutinases. 77 predicted secreted proteins have oxidoreductase activity, 71 carry a cytochrome P450, 56 harbour conserved domains of unknown function and 396 did not carry a Pfam domain. H. fraxineus has a large number of predicted secreted Cytochrome P450 proteins. Cytochrome P450s’ modes of action are typically via the monooxygenase reaction, which is important in the generation and destruction of chemicals especially aromatics. In H. fraxineus, they may be important in pathogenesis, especially as they carry a signal of diversifying selection (Supplementary Analysis 8). Potential roles for P450s in pathogenesis include: (1) destruction of ash-tissue-derived aromatic compounds with antifungal properties. Plant secondary metabolism pathways are active during the infection process of Magnaporthe oryzae28. (2) Penetration of ash tree tissues through extracellular oxidation of wood and metabolism of hydrocarbon compounds, which are the main constituents of host plants cuticle29. (3) P450s are part of the biosynthetic pathways for mycotoxins and phytohormones secreted by H. fraxineus during invasion of ash tissue, similar modes of action have been shown for Fusarium multifunctional cytochrome P450 monooxygenase Tri430. The presence of such a high predicted number of P450s in the secretome of H. fraxineus suggests biochemistry unique to H. fraxineus.
Transcribed low-complexity domains may provide effectors with flexibility and diversity driven by adaptive evolution31. Of 1,132 effectors, 22% contain short repeats (for example, tetratricopeptide repeats and leucine-rich repeats; Supplementary Table 7). Some are predicted to have a nuclear localization signal often indicative of intracellular function in host cells whilst others have possible apoplastic function. Many putative effectors (37%), are cysteine rich with between one and five predicted disulphide bonds (Supplementary Table 8), which are predicted to confer stability, biological activity and resistance to proteases in the apoplastic space32 and inside infected host cells33.
We identify three effectors (one of them previously identified16) with an NPP1 (necrosis-inducing Phytophthora protein) domain, which is present in fungal, oomycete and bacterial proteins that induce hypersensitive-reaction-like cell death upon infiltration into plant leaves34. Two other proteins have predicted cell death activity and two more carry S1/P1 nuclease activity domains, which are involved in non-specific cleavage of RNA and single-stranded DNA. Host cell death induction is suggested to be common among facultative parasites that engage in a necrotrophic lifestyle35.
Adaptive effector diversity is present in Europe, but is far greater in the native range
We analysed genetic diversity in all the genes of the European and Japanese populations (Supplementary Table 9). First, there remains a significant positive correlation in the level of polymorphism present in genes between the invasive European and native Japanese populations, despite an 81% reduction in SNP density in Europe (Fig. 6). This reduction in the level of polymorphism has impacted putative effector and other genes alike, whereas, in Japan putative effector genes maintain significantly greater levels of polymorphism than that of other genes (Fig. 6).
a, The Japanese population contains an increased level of diversity to that in Europe but despite the founder effect, genes that are more polymorphic in Japan tend to be more polymorphic in Europe (Eu–Jp SNP density: gene, R2 = 0.664, n = 3214, p < 0.001; effector, R2 = 0.605, n = 369, p < 0.001). Linear model calculated using log–log data and plotted here on the original scale (see Supplementary Fig. 23 for log–log scale). b, The level of genetic diversity in Europe is not significantly different between potential effectors (Wilcoxon (π), Eu genes ~ Eu effectors: W(10,942) = 5,420,600, n = 10,943, p = 0.78 (not significant)) whereas, in Japan putative effectors carry significantly more genetic diversity (Wilcoxon (π), Jp genes < Jp effectors: W(10,942) = 5,671,200 n = 10943, *p = 0.041). Nucleotide diversity over all genes combined is significantly greater in Japan than in Europe (Wilcoxon (π), Jp all genes > Eu all genes: W(21,885) = 102,060,000 n = 21,886, ***p < 0.001). c, The signal of adaptive diversity (PN:PS) is significantly different between putative effectors and all other genes (permutation test of equality: n = 9,258, p < 0.001; joint kernel density shaded). The increase in effector PN:PS to other genes, over this range, is evidence for reduced efficacy of purifying selection in this classification of genes. For genes with a PN:PS greater than one and less than two (c, inset) effector genes again peak outside the joint kernel density, consistent with the operation of positive selection driving adaptive change in these effectors (permutation test of equality: n = 183, p = 0.01). d, The overall level of adaptive polymorphism is greater in effectors than other genes in both Japan and Europe (Wilcoxon (PN:PS), Jp genes < Jp effectors: W(9,510) = 4,891,800, n = 9,511, ***p < 0.001; Eu genes < Eu effectors: W(6,080) = 1,750,600, n = 6081, *p = 0.045). However, the strength of this signal has been reduced in the European population. Boxplots show the median, upper and lower quartiles with outliers plotted outside whiskers which extend 1.5 times beyond the interquartile range (see Supplementary Figs. 24,25 for outliers).
Putative effectors in Europe have an increase in SNPs that affect splice regions as well as 5′ UTRs. Importantly however, there are also increases in SNP densities in synonymous, intron, up- and downstream positions in putative effectors relative to other genes (Supplementary Analysis 8). Therefore, increases within these effector features could be the product of increased SNP density through linkage and balancing selection36 operating on effectors.
Pathogen effectors must adapt to avoid host recognition and so they are expected to undergo positive, diversifying and balancing selection15. To investigate the role of selection in the maintenance of polymorphism we measured adaptive diversity using the mean pairwise ratio of non-synonymous (PN) to synonymous (PS) polymorphism present in all genes. The PN:PS ratio can detect the presence of adaptive evolution when applied to short regions of a gene, for example, a binding domain, because a value greater than one indicates positive or diversifying selection (functional divergence)37 and PN:PS close to zero indicates negative or purifying selection. Here, we apply the PN:PS ratio across whole genes, which we presume have different modes of selection operating across them. However, we expect effectors will retain an overall higher PN:PS value despite dilution by purifying selection operating over the rest of the gene.
Approximately 97% of all PN:PS values in the European and Japanese effectors are lower than one. Within this range, we observe a significant increase in the effector PN:PS over that of other genes between 0.2–0.5 (Fig. 6). This represents a reduction in the strength of purifying selection in effectors relative to other genes. At PN:PS values greater than one, we observe a significant peak in the density for effectors which, indicates a number of effectors evolving under contemporary positive selection (Fig. 6). Effectors in Japan have higher PN:PS values than other genes (Fig. 6). In Europe, effector adaptive diversity has been maintained despite the bottleneck, albeit at a lower level than in the native range (Fig. 6). Finally, consistent with linkage and balancing selection36, the level of synonymous polymorphism is also higher in effectors in both Japan and Europe (Supplementary Fig. 25). Pseudogenes too, may have a PN:PS ratio approaching one. Pseudogenization could be more readily observed for effectors increasingly recognized by the immune system38. Here, we do not observe a significant increase in start loss, stop gain or stop loss variants in effectors over other genes (Supplementary Analysis 8).
Conclusions
The bottleneck of H. fraxineus into Europe removed the majority of neutral and much of the adaptive genetic variation. Despite this, the pathogen has devastated ash from east to west and host defence is characterized in terms of levels of susceptibility5. Further introduction of pathogen genetic diversity runs the risk of increasing disease prevalence. Successive invasions can increase the level of genetic diversity above that of native populations39 and drive temporal fluctuations in adaptive diversity40. Increased virulence at invasion onset, due to uninfected host density, may be attenuated by natural selection on the pathogen as uninfected density reduces41 but this is much less probable where immigrations continue.
Tree pathogens can be introduced through trade in live trees and untreated wood1. If multiple native genotypes can invade, we face a situation where further immigration is likely to be accompanied by extreme increases in the level of adaptive polymorphism and disease severity. It is most important to prevent further introduction of pathogen diversity to Europe, particularly by prohibiting imports of Fraxinus species from East Asia, including material that may contain leaf debris bearing ascocarps.
The H. fraxineus native range, Asia, is large and the fungus lives there on the leaf litter of several ash species. Broader population genetic analyses over this range will allow us to address important questions on H. fraxineus adaptation to specific hosts. Moreover, the two-haploid founder scenario provides a unique system to unravel the population genetics of invasion with genome evolution. Finally, efforts to gain an understanding of disease progression will combine effector nucleotide statistics with targeted RNA-seq experiments.
Notwithstanding the potential impact of an introduction of the emerald ash borer42, current levels of European pathogen genetic diversity may infect and kill 95% of all European ash. We must consider the implications of any further introduction of diversity from the ash dieback pathogen to an already dire situation.
Methods
H. fraxineus collection and sequencing
44 H. fraxineus isolates were collected from Europe (21 UK, 13 Norway, 5 France, 4 Poland, 1 Austria) and 9 from Japan (Supplementary Table 1). Japanese isolates were made up of one haploid and eight fruiting bodies (see isolate phasing below). Fruiting bodies were stored in ethanol and haploid isolates were cultured on malt extract agar43. Species confirmation was performed by PCR and sequencing of ITS sequences. Isolates were genome sequenced at either Edinburgh Genomics or the Earlham Institute on MiSeq and HiSeq Illumina platforms at 25–160 sequence depth (Supplementary Analysis 1).
Genome assembly and annotation of H. fraxineus
As part of a commitment to open science and rapid community analyses, the genome of isolate KW1, Hf-v1.1, was released online in March 2013 on the Open Ash Dieback repository (oadb.tsl.ac.uk)16. Hf-v2.0, published here and released online in February 2015, was assembled using a 200 bp insert paired end (PE) and a long (5 kbp) mate pair (LMP) library. SOAPec error correction was performed on PE reads (overlapping merged) and SOAPdenovo v2.0444 was used to assemble and scaffold (Supplementary Analysis 2).
The annotation pipeline used for genome Hf-v1.116 was replicated on Hf-v2 adding newer RNA-seq data from the Open Ash Dieback repository (Supplementary Analysis 3). The Hf-v2 genome was repeat masked using de novo models, known fungal and low-complexity repeats. Annotations (9,737) from Hf-v1.1 were transferred to Hf-v2. Augustus v2.745 was used to predict additional protein-coding genes using protein and RNA-seq alignments (16 libraries, Supplementary Table 16) with the previously trained H. fraxineus model. We ran a transcript extension protocol on all genes to identify whether those genes within 100 bp of an upstream gene had evidence for extension. We also checked this extended gene dataset for potential secreted proteins. This protocol allowed the identification of 36 genes that had not previously been recognized as having a methionine start codon which were included into our effector-mining pipeline (Supplementary analysis 3).
We used a pipeline described previously32 to identify putative effectors. Briefly, transcripts with a signal peptide were identified and from these we remove transcripts with transmembrane or mitochondrial localization signals and then transcripts with repeats, the disulphide bonds and nuclear localization signals are identified. Finally, secreted proteins were clustered into tribes (see Supplementary analysis 8).
Mapping and SNP calling
Reads were trimmed for Illumina adapters and Phred quality (-q20) and read alignment was performed using BWA-MEM v0.7.746 after which duplicates were removed. VCFtools v0.1.1347 was used to filter variants to a minimum depth of ten, maximum depth of 1.8 × mean individual depth and a SNP genotype quality of at least 30. SNP sites with more than two alleles were excluded as probable errors, as were sites reported as heterozygous in haploids. Finally, sites that were missing in ≥20% individuals were removed. Three samples were removed from further analyses because of insufficient depth or evidence of contamination, leaving 20 UK, 13 Norway, 5 France, 4 Poland, 1 Austria, and 8 Japan (1 haploid 7 fruiting bodies; see Supplementary Analysis 1).
Phasing fruiting body samples
We used Shapeit248, to phase fruiting body data, this first applies a phase informative read step (extractPIRs) to group SNPs present on the same read pair and then phases remaining SNPs using population level polymorphism (assemble --states 1,000 --burn 60 --prune 60 --main 300 --effective-size 564,000 --window 0.5). All Japanese samples including the haploid strain (as control) were phased. As required indels were removed from the dataset before phasing. Effective population size (ϑ = 2Neμ) was calculated assuming a mutation rate of 5 × 10−9 per generation per site49 using the mean nucleotide diversity present in 100-kbp windows in the Japanese population.
SNP diversity and divergence analysis
Polymorphism statistics and sliding windows, π and FST were conducted using VCFtools v0.1.1347. DNAsp v5.10.0150 was used to calculate population genetic statistics per gene and PAML v4.951 was used (YN00) in pairwise mode to calculate the average PN:PS ratio nonsynonymous (PN) and synonymous (PS) values for polymorphism in each gene with at least one synonymous mutation. The sm package52 in R v3.2.1 compared the density distributions of PN:PS. Output data for all genes are combined in Supplementary Table 9 (see also Supplementary Analysis 6). Data are represented using SplitsTree v4.13.153 using the neighbour-net algorithm and CIRCOS v0.67-554.
Parental cross and linkage analysis
Two H. fraxineus isolates of opposite mating types, LWD054 and LWD067, were collected from stem lesions on trees in Norfolk, UK in 2014. Crosses were made according to a previously reported method55 except that isolates were kept at 25 °C in 16 hours light. KASP56 markers were designed to SNPs between two H. fraxineus parents mapped to the Hf-v2 assembly (Supplementary Analysis 4). Linkage between scaffolds was ascertained using chi-squared test of sites close to ends of the 23 scaffolds, using 28 F1 offspring. Scaffolds at linkage group ends were examined for the presence of telomeric motifs.
Statistics
R v3.2.157 was used to fit simple regression models to test the level of association between population signals in order to show directionality in the invasion and portray the population contraction (founder effect) behind Tajima’s D. To evaluate the correlation in gene diversity (the proportion of SNPs per gene) in Japanese and the European populations we log-transformed SNP density excluding genes with less than two SNPs (leaving 6,216 genes). SNP density regression models were calculated on log–log data and plotted on the original scale. We also correlated feature coverage (that is, repeat content) of regions with gene and effector coverage to understand genome evolution. Two-sample Wilcoxon tests (one tailed indicated by ‘>’ or ‘<’; two tailed indicated by ‘~‘), were used to test for differences in signals of nucleotide diversity, Tajima’s D and PN:PS between non-effector and effector genes in European and Japanese populations. Bootstrap analyses compare variants effect by feature or gene (i.e. SnpEff output) and involved sampling feature with replacement (×1,000). Bootstrapped comparisons were tested using a randomization test and 95% confidence intervals are presented.
Ancestral effective population size
To estimate the size of the population from which the European population was founded we used the SNPs at the third base pair in codons within 387 core eukaryotic genes. CEGMA v2.421 initially identified 440 CEGs which was reduced to 387 genes that were confirmed by reciprocal blastp (BLAST v2.2.3158 (-evalue 1 × 10–5) with a minimum of 90% coverage (removing technical assembly errors and biological duplicates), removing CEGs with nonsense variants (expected pseudogenes) and a minimum mean depth of 10× per gene across all individuals (poorly covered CEGs). Remaining genes were then run through a pipeline to quantify the pairwise distance between alleles (the level of nucleotide polymorphism, or distance between haplotypes, is defined as the number of SNPs (S) divided by the total length of the sequence (N) using SplitsTree) and the diversity per gene (using DNAsp v5). Relative pairwise distances between (third base pair) haplotypes is plotted as a density including all 387 CEGs combined. CEG networks are shown as a visual demonstration of the shapes of three polymorphic networks in Japan and Europe HYMFR746836.2.0_000003240.1, -000004200.1, -000061770.1)
The average number of base pairs (third codon base pair) per CEG was 448 and this number was used as an input to fastsimcoal2 v2.5.2.859, a fast sequential Markov coalescent simulator used to estimate the size of the population from which the founders into Europe came. A simple model (×1,000) of a single (haploid) population of fixed effective size (Ne = 100,000–4,000,000) containing 387 unlinked CEGs that were freely mutating at a rate of 5 × 10−9 per base per generation49 was bottlenecked to two individuals. From these two individuals, we recorded the number of haplotypes and the haplotype divergence at each gene for comparison to observed data.
Reporting Summary
Further information on experimental design is available in the Nature Research Reporting Summary linked to this article.
Data availability
All MP and LMP sequencing data generated for the Hf-v2.0 genome (PRJEB21027), population genetics and parental cross reads have been submitted to the European Nucleotide Archive (under projects PRJEB21059, PRJEB21060, PRJEB21061, PRJEB21062, PRJEB21063, PRJEB21064 and accessions ERS480843–ERS480865; see Supplementary Table 1). The genome annotation is available at the Earlham Institute Open Data site (http://opendata.earlham.ac.uk/Hymenoscyphus_fraxineus/EI/v2/).
References
Ghelardini, L., Pepori, A. L., Luchi, N., Capretti, P. & Santini, A. Drivers of emerging fungal diseases of forest trees. For. Ecol. Manag. 381, 235–246 (2016).
Brasier, C. M. & Buck, K. W. Rapid evolutionary changes in a globally invading fungal pathogen (Dutch elm disease). Biol. Invasions 3, 223–233 (2001).
Drenkhan, R. et al. Hymenoscyphus fraxineus is a leaf pathogen of local Fraxinus species in the Russian Far East. Plant Pathol. 66, 490–500 (2017).
McKinney, L. V., Nielsen, L. R., Hansen, J. K. & Kjær, E. D. Presence of natural genetic resistance in Fraxinus excelsior (Oleraceae) to Chalara fraxinea (Ascomycota): an emerging infectious disease. Heredity 106, 788–797 (2011).
Sollars, E. S. et al. Genome sequence and diversity of European ash trees threatened by ash dieback. Nature 541, 212–216 (2017).
Downie, J. A. Ash dieback epidemic in Europe: How can molecular technologies help? PLoS Pathog. 13, 1–6 (2017).
Gross, A., Zaffarano, P. L., Duo, A. & Grünig, C. R. Reproductive mode and life cycle of the ash dieback pathogen Hymenoscyphus pseudoalbidus. Fungal Genet. Biol. 49, 977–86 (2012).
Fones, H. N., Mardon, C. & Gurr, S. J. A role for the asexual spores in infection of Fraxinus excelsior by the ash-dieback fungus Hymenoscyphus fraxineus. Sci. Rep. 6, 34638 (2016).
McDonald, B. A. & Linde, C. Pathogen population genetics, evolutionary potential, and durable resistance. Annu. Rev. Phytopathol. 40, 349–379 (2002).
Charlesworth, B. Effective population size and patterns of molecular evolution and variation. Nat. Rev. Genet. 10, 195–205 (2009).
Gladieux, P. et al. The population biology of fungal invasions. Mol. Ecol. 24, 1969–1986 (2015).
Fry, W. E. Phytophthora infestans: New tools (and old ones) lead to new understanding and precision management. Annu. Rev. Phytopathol. 54, 529–547 (2016).
Gross, A., Hosoya, T. & Queloz, V. Population structure of the invasive forest pathogen Hymenoscyphus pseudoalbidus. Mol. Ecol. 23, 2943–2960 (2014).
Schoebel, C. N., Botella, L., Lygis, V. & Rigling, D. Population genetic analysis of a parasitic mycovirus to infer the invasion history of its fungal host. Int. J. Lab. Hematol. 38, 42–49 (2016).
Rafiqi, M., Ellis, J. G., Ludowici, V. A., Hardham, A. R. & Dodds, P. N. Challenges and progress towards understanding the role of effectors in plant-fungal interactions. Curr. Opin. Plant Biol. 15, 477–482 (2012).
Saunders, D. G. et al. Crowdsourced analysis of ash and ash dieback through the Open Ash Dieback project: A year 1 report on datasets and analyses contributed by a self-organising community. Preprint at bioRxiv https://doi.org/10.1101/004564 (2014).
Tajima, F. Statistical-method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123, 585–595 (1989).
Croll, D., Lendenmann, M. H., Stewart, E. & McDonald, B. A. The impact of recombination hotspots on genome evolution of a fungal plant pathogen. Genetics 201, 1213–1228 (2015).
McMullan, M. et al. Evidence for suppression of immunity as a driver for genomic introgressions and host range expansion in races of Albugo candida, a generalist parasite. eLife 4, e04550 (2015).
Burokiene, D. et al. Genetic population structure of the invasive ash dieback pathogen Hymenoscyphus fraxineus in its expanding range. Biol. Invasions 17, 2743–2756 (2015).
Parra, G., Bradnam, K. & Korf, I. CEGMA: a pipeline to accurately annotate core genes in eukaryotic genomes. Bioinformatics 23, 1061–7 (2007).
Stukenbrock, E. H. & McDonald, B. A. Population genetics of fungal and oomycete effectors involved in gene-for-gene interactions. Mol. Plant. Microbe Interact. 22, 371–80 (2009).
Raffaele, S. & Kamoun, S. Genome evolution in filamentous plant pathogens: why bigger can be better. Nat. Rev. Microbiol. 10, 417–430 (2012).
Derbyshire, M. et al. The complete genome sequence of the phytopathogenic fungus Sclerotinia sclerotiorum reveals insights into the genome architecture of broad host range pathogens. Genome Biol. Evol. 9, 593–618 (2017).
Okagaki, L. H., Sailsbery, J. K., Eyre, A. W. & Dean, R. A. Comparative genome analysis and genome evolution of members of the Magnaporthaceae family of fungi. BMC Genom. 17, 135 (2016).
Stukenbrock, E. H. & Croll, D. The evolving fungal genome. Fungal Biol. Rev. 28, 1–12 (2014).
Tellier, A. & Brown, J. K. M. Stability of genetic polymorphism in host–parasite interactions. Proc. R. Soc. B 274, 809–817 (2007).
Collemare, J. et al. Magnaporthe grisea avirulence gene ACE1 belongs to an infection-specific gene cluster involved in secondary metabolism. New Phytol. 179, 196–208 (2008).
Syed, K. & Yadav, J. P450 monooxygenases (P450ome) of the model white rot fungus Phanerochaete chrysosporium. Crit. Rev. Microbiol. 38, 339–363 (2012).
Tokai, T. et al. Fusarium Tri4 encodes a key multifunctional cytochrome P450 monooxygenase for four consecutive oxygenation steps in trichothecene biosynthesis. Biochem. Biophys. Res. Commun. 353, 412–417 (2007).
Mesarich, C. H., Bowen, J. K., Hamiaux, C. & Templeton, M. D. Repeat-containing protein effectors of plant-associated organisms. Front. Plant Sci. 6, 872 (2015).
Saunders, D. G. et al. Using hierarchical clustering of secreted protein families to classify and rank candidate effectors of rust fungi. PLoS ONE 7, e29847 (2012).
Blondeau, K. et al. Crystal structure of the effector AvrLm4-7 of Leptosphaeria maculans reveals insights into its translocation into plant cells and recognition by resistance proteins. Plant J. 83, 610–624 (2015).
Fellbrich, G. et al. NPP1, a Phytophthora-associated trigger of plant defense in parsley and Arabidopsis. Plant J. 32, 375–390 (2002).
Gijzen, M. & Nürnberger, T. Nep1-like proteins from plant pathogens: Recruitment and diversification of the NPP1 domain across taxa. Phytochemistry 67, 1800–1807 (2006).
Hughes, A. L. & Yeager, M. Natural selection at major histocompatibility complex loci of vertebrates. Annu. Rev. Genet. 32, 415–435 (1998).
Stukenbrock, E. H. & Bataillon, T. A population genomics perspective on the emergence and adaptation of new plant pathogens in agro-ecosystems. PLoS Pathog. 8, e1002893 (2012).
Burstein, D. et al. Genomic analysis of 38 Legionella species identifies large and diverse effector repertoires. Nat. Genet. 48, 167–175 (2016).
Uller, T. & Leimu, R. Founder events predict changes in genetic diversity during human-mediated range expansions. Glob. Change Biol. 17, 3478–3485 (2011).
McMullan, M. & van Oosterhout, C. Inference of selection based on temporal genetic differentiation in the study of highly polymorphic multigene families. PLoS ONE 7, e42119 (2012).
Berngruber, T. W., Froissart, R., Choisy, M. & Gandon, S. Evolution of virulence in emerging epidemics. PLoS Pathog. 9, e1003209 (2013).
Villari, C., Herms, D. A., Whitehill, J. G. A., Cipollini, D. & Bonello, P. Progress and gaps in understanding mechanisms of ash tree resistance to emerald ash borer, a model for wood-boring insects that kill angiosperms. New Phytol. 209, 63–79 (2016).
Kowalski, T. Chalara fraxinea sp. nov. associated with dieback of ash (Fraxinus excelsior) in Poland. For. Pathol. 36, 264–270 (2006).
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1, 18 (2012).
Stanke, M. & Morgenstern, B. AUGUSTUS: A web server for gene prediction in eukaryotes that allows user-defined constraints. Nucleic Acids Res. 33, 465–467 (2005).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Delaneau, O., Zagury, J.-F. & Marchini, J. Improved whole-chromosome phasing for disease and population genetic studies. Nat. Methods 10, 5–6 (2013).
Kasuga, T., White, T. J. & Taylor, J. W. Estimation of nucleotide substitution rates in Eurotiomycete fungi. Mol. Biol. Evol. 19, 2318–2324 (2002).
Librado, P. & Rozas, J. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452 (2009).
Yang, Z. H. PAML 4: Phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Bowman, A. W. & Azzalini, A. Computational aspects of nonparametric smoothing with illustrations from the sm library. Comput. Stat. Data Anal. 42, 545–560 (2003).
Huson, D. H. & Bryant, D. Application of phylogenetic networks in evolutionary studies. Mol. Biol. Evol. 23, 254–267 (2006).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Wey, T., Schlegel, M., Stroheker, S. & Gross, A. MAT- gene structure and mating behavior of Hymenoscyphus fraxineus and Hymenoscyphus albidus. Fungal Genet. Biol. 87, 54–63 (2016).
Semagn, K., Babu, R., Hearne, S. & Olsen, M. Single nucleotide polymorphism genotyping using Kompetitive Allele Specific PCR (KASP): Overview of the technology and its application in crop improvement. Mol. Breed. 33, 1–14 (2014).
R: A Language and Environment for Statistical Computing (R Core Team, 2017).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–10 (1990).
Excoffier, L. & Foll, M. fastsimcoal: A continuous-time coalescent simulator of genomic diversity under arbitrarily complex evolutionary scenarios. Bioinformatics 27, 1332–1334 (2011).
Acknowledgements
We thank The Sainsbury Laboratory for hosting the Open Ash Dieback Git repository (oadb.tsl.ac.uk), and M. Caccamo and S. Janacek for assistance with project management. The submission of sequencing data was brokered by the COPO platform (https://copo-project.org), funded by the Biotechnology and Biological Sciences Research Council (BBSRC) (BB/L024055/1) and supported by CyVerse UK, part of the Earlham Institute National Capability in e-Infrastructure. The Ash Dieback research project was funded jointly by the BBSRC and the Department of Environment Food and Rural Affairs (Defra) via the Nornex project (BBS/E/J/000CA523), the Economic and Social Research Council, the Forestry Commission, the Natural Environment Research Council (NERC) and the Scottish Government, under the Tree Health and Plant Biosecurity Initiative (BB/L01291X/1). The UMR IAM is supported by a grant overseen by the French National Research Agency (ANR) as part of the Investissements d’Avenir programme (ANR-11-LABX-0002-01, Laboratory of Excellence ARBRE). Genome sequencing was done by Edinburgh Genomics, and the Earlham Institute (formerly The Genome Analysis Centre, Norwich), by members of the Platforms and Pipelines Group funded by BBSRC National Capability in Genomics grant (BB/J010375/1). Additional research funding was via a BBSRC Institute Strategic Programme Grant for Bioinformatics (BB/J004669/1) to the Earlham Institute.
Author information
Authors and Affiliations
Contributions
M.M., C.F., J.B., M.B., A.D. and M.C. contributed to the design of the project. E.O., A.E., K.Y., T.H., L.W., P.J., R.I., C.H., A.H., A.V., H.S. and A.D. contributed to collection of samples. Acquisition and processing of sequence data was done by M.M., W.V., A.E., J.W., Geo.K. and D.M.; M.M., M.R., Gem.K., B.C., G.G., L.B., E.O., D.G.S., L.P.A., B.W., D.S., J.B. and M.C. contributed to the analysis and interpretation of data. M.M., M.R., Gem.K., B.C., L.P.A., L.B., E.O., A.H., N.H., D.S., J.B., A.D. and M.C. contributed to drafting and revising of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Analysis
Supplementary Table 1
Sample collection, extraction and sequencing details along with accession numbers
Supplementary Table 2
KASP marker position and telomeric repeat length per scaffold
Supplementary Table 3
Linkage between markers designed across H. fraxineus scaffolds v2 genome assembly
Supplementary Table 4
Extended gene models with a signalP4 hit and methionine start codon
Supplementary Table 5
Putative effector tribe clustering using TribMCL
Supplementary Table 6
Effector candidates of H. fraxineus with corresponding Pfam signatures
Supplementary Table 7
Repeat-rich effector candidates of H. fraxineus
Supplementary Table 8
Cysteine-rich effector candidates of H. fraxineus
Supplementary Table 9
Population genetic statistics per gene in both the Japanese and European populations
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
McMullan, M., Rafiqi, M., Kaithakottil, G. et al. The ash dieback invasion of Europe was founded by two genetically divergent individuals. Nat Ecol Evol 2, 1000–1008 (2018). https://doi.org/10.1038/s41559-018-0548-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-018-0548-9
This article is cited by
-
Load of the ash dieback pathogen hymenoscyphus fraxineus differs in soil
Journal of Plant Diseases and Protection (2024)
-
Endophytic fungi related to the ash dieback causal agent encode signatures of pathogenicity on European ash
IMA Fungus (2023)
-
Recent co-evolution of two pandemic plant diseases in a multi-hybrid swarm
Nature Ecology & Evolution (2023)
-
Applying molecular and genetic methods to trees and their fungal communities
Applied Microbiology and Biotechnology (2023)
-
Evidence of a component Allee effect for an invasive pathogen: Hymenoscyphus fraxineus, the ash dieback agent
Biological Invasions (2023)