Abstract
Around 150 million years ago, eusocial termites evolved from within the cockroaches, 50 million years before eusocial Hymenoptera, such as bees and ants, appeared. Here, we report the 2-Gb genome of the German cockroach, Blattella germanica, and the 1.3-Gb genome of the drywood termite Cryptotermes secundus. We show evolutionary signatures of termite eusociality by comparing the genomes and transcriptomes of three termites and the cockroach against the background of 16 other eusocial and non-eusocial insects. Dramatic adaptive changes in genes underlying the production and perception of pheromones confirm the importance of chemical communication in the termites. These are accompanied by major changes in gene regulation and the molecular evolution of caste determination. Many of these results parallel molecular mechanisms of eusocial evolution in Hymenoptera. However, the specific solutions are remarkably different, thus revealing a striking case of convergence in one of the major evolutionary transitions in biological complexity.
Similar content being viewed by others
Main
Eusociality, the reproductive division of labour with overlapping generations and cooperative brood care, is one of the major evolutionary transitions in biology1. Although rare, eusociality has been observed in a diverse range of organisms, including shrimps, mole rats and several insect lineages2,3,4. A particularly striking case of convergent evolution occurred within the holometabolous Hymenoptera and in the hemimetabolous termites (Isoptera), which are separated by over 350 Myr of evolution5. Termites evolved within the cockroaches around 150 Myr ago, towards the end of the Jurassic period6,7, about 50 Myr before the first bees and ants appeared5. Therefore, identifying the molecular mechanisms common to both origins of eusociality is crucial to understanding the fundamental signatures of these rare evolutionary transitions. While the availability of genomes from many eusocial and non-eusocial hymenopteran species8 has allowed extensive research into the origins of eusociality within ants and bees9,10,11, a paucity of genomic data from cockroaches and termites has precluded large-scale investigations into the evolution of eusociality in this hemimetabolous clade.
The conditions under which eusociality arose differ greatly between the two groups. Termites and cockroaches are hemimetabolous and so show a direct development, while holometabolous hymenopterans complete the adult body plan during metamorphosis. In termites, workers are immatures and only reproductive castes are adults12, while in Hymenoptera, adult workers and queens represent the primary division of labour. Moreover, termites are diploid and their colonies consist of both male and female workers, and usually a queen and king dominate reproduction. This is in contrast to the haplodiploid system found in Hymenoptera, in which all workers and dominant reproductives are female. It is therefore intriguing that strong similarities have evolved convergently within the termites and the hymenopterans, such as differentiated castes and a nest life with reproductive division of labour. The termites can be subdivided into wood-dwelling and foraging termites. The former belong to the lower termites and produce simple, small colonies with totipotent workers that can become reproductives. Foraging termites (some lower and all higher termites) form large, complex societies, in which worker castes can be irreversible12. For this reason, higher, but not lower, termites can be classed as superorganismal13. Similarly, within Hymenoptera, varying levels of eusociality exist.
Here we provide insights into the molecular signatures of eusociality within the termites. We analysed the genomes of two lower and one higher termite species and compared them to the genome of the German cockroach, Blattella germanica, as a closely related non-eusocial outgroup. Furthermore, differences in expression between nymphs and adults of the cockroach were compared to differences in expression between workers and reproductives of the three termites, to gain insights into how expression patterns changed along with the evolution of castes. Using 15 additional insect genomes to infer background gene family turnover rates, we analysed the evolution of gene families along the transition from non-social cockroaches to eusociality in the termites. In this study, we concentrated particularly on two hallmarks of insect eusociality, as previously described for Hymenoptera, with the expectation that similar patterns occurred along with the emergence of termites. These are the evolution of a sophisticated chemical communication, which is essential for the functioning of a eusocial insect colony3,14,15, and major changes in gene regulation along with the evolution of castes9,10. We also tested whether transposable elements spurred the evolution of gene families that were essential for the transition to eusociality.
Evolution of genomes, proteomes and transcriptomes
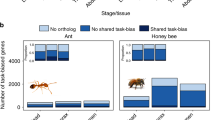
We sequenced and assembled the genome of the German cockroach, B. germanica (Ectobiidae), and of the lower, drywood termite Cryptotermes secundus (Kalotermitidae; for assembly statistics, see Supplementary Table 1). The cockroach genome (2.0 Gb) is considerably larger than all three termite genomes. The genome size of C. secundus (1.30 Gb) is comparable to the higher, fungus-growing termite Macrotermes natalensis (1.31 Gb, Termitidae)16, but more than twice as large as the lower, dampwood termite Zootermopsis nevadensis (562 Mb, Termopsidae)17. The smaller genomes of termites compared to the cockroach are in line with previous size estimations based on C-values18. The proteome of B. germanica (29,216 proteins) is also much larger than in the termites, where we find the proteome size in C. secundus (18,162) to be similar to those of the other two termites (M. natalensis: 16,140; Z. nevadensis: 15,459; Fig. 1). In fact, the B. germanica proteome was the largest among all 21 arthropod species analysed here (Fig. 1). Strong evidential support for over 80% of these proteins in B. germanica (see Supplementary Information) and large expansions in many manually annotated gene families offer high confidence in the accuracy of this proteome size. We also compared gene expression between the four species. When comparing worker expression with queen expression in the termites and nymph expression (fifth and sixth instars) with adult female expression in B. germanica, we found shifts in specificity of expression for termites compared to the cockroach in several gene families (Fig. 2). It has previously been reported for the primitively eusocial paper wasp Polistes canadensis that the majority of caste-biased genes, especially those upregulated in workers, are novel genes19. The authors suggested that this may be a feature of early eusociality. We did not find the same pattern for the termites. Species-specific genes (those without an orthologue) were not enriched for differentially expressed genes in any of the termites, with slight peaks among Blattodea- and Isoptera-specific genes (Supplementary Fig. 1).
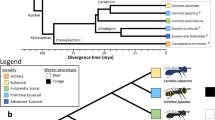
From left to right: a phylogenetic tree of 20 insect species with Strigamia maritima (centipede) as the outgroup (species of newly sequenced genomes presented in this study are underlined); level of eusociality (one red insect: simple eusociality; two red insects: advanced eusociality; black fly: non-eusocial); fractions of repetitive content (yellow) within genomes of selected species (for sources, see Supplementary Information); proportions of species-specific gene family expansions (green), contractions (red) and stable gene families (black), the size of the pies represents the relative size of the gene family change (based on total numbers); a bar chart showing protein orthology across taxonomic groups within each genome. Ma, million years ago. Smar, Strigamia maritima; Edan, Ephemera danica; Rpro, Rhodnius prolixus; Nvit, Nasonia vitripennis; Amel, Apis mellifera; Pcan, Polistes canadensis; Hsal, Harpegnathos saltator; Lhum, Linepithema humile; Cflo, Camponotus floridanus; Pbar, Pogonomyrmex barbatus; Sinv, Solenopsis invicta; Aech, Acromyrmex echinatior; Acep, Atta cephalotes; Tcas, Tribolium castaneum; Aaeg, Aedes aegypti; Dmel, Drosophila melanogaster; Lmig, Locusta migratoria; Bger, Blattella germanica; Znev, Zootermopsis nevadensis; Csec, Cryptotermes secundus; Mnat, Macrotermes natalensis.
Shown from left to right are: a simple phylogeny97 describing important novelties along the evolutionary trajectory to termites (numbers in brackets are genome sizes); life cycles; differential expression (log2(fold change) > 1 and P < 0.05; DESeq279; sample sizes are shown in the last column) between workers and queens (between nymphs and adult females in B. germanica) of the selected gene families (Desat, desaturases; Elong, elongases; H’ween, Halloween genes) and total numbers within all genes; the numbers denote total numbers of genes in each gene family.
Gene family expansions assisted by TEs in termites
The transitions to eusociality in ants10 and bees9 have been linked to major changes in gene family sizes. Similarly, we detected significant gene family changes on the branch leading to the termites (seven expansions and ten contractions; Supplementary Fig. 2 and Supplementary Table 2). The numbers of species-specific, significant expansions and contractions of gene families varied within termites (Z. nevadensis: 15/5; C. secundus: 27/3; M. natalensis: 24/20; Supplementary Fig. 2 and Supplementary Tables 3–5). Interestingly, in B. germanica we measured 93 significant gene family expansions but no contractions (Supplementary Table 6), which contributed to the large proteome.
The termite and cockroach genomes contain a higher level of repetitive DNA compared to the hymenopterans we analysed (Fig. 1). C. secundus and B. germanica genomes both contain 55% repetitive content (Supplementary Table 7), which is higher than in both Z. nevadensis (28%) and the higher termite M. natalensis (46%; Fig. 1)20. As also found in Z. nevadensis and M. natalensis20, LINEs and especially the subfamily BovB were the most abundant transposable elements (TEs) in the B. germanica and C. secundus genomes, indicating that a proliferation of LINEs may have occurred in the ancestors of Blattodea (cockroaches and termites).
We hypothesized that these high levels of TEs may be driving the high turnover in gene family sizes within the termites and B. germanica21. Expanded gene families indeed had more repetitive content within 10-kb flanking regions in all three termites (P < 1.3 × 10−8; Wald t-test; Supplementary Tables 8 and 9), in particular in the higher termite M. natalensis. In contrast, gene family expansions were not correlated with TE content in flanking regions for B. germanica. These results suggest that a major expansion of LINEs at the root of the Blattodea clade contributed to the evolution of gene families within termites, probably via unequal crossing-over21; however, the expansions in B. germanica were not facilitated by TEs. It can therefore be speculated that the large expansion of LINEs within Blattodea allowed the evolution of gene families that ultimately facilitated the transition to eusociality.
Expansion and positive selection of ionotropic receptors
Insects perceive chemical cues from toxins, pathogens, food and pheromones with three major families of chemoreceptors, the odorant (ORs), gustatory and ionotropic (IRs) receptors22. ORs, in particular, have been linked to colony communication in eusocial Hymenoptera, where they abound14,15,23. Interestingly, as previously detected for Z. nevadensis17, the OR repertoire is substantially smaller in B. germanica and all three termites compared to hymenopterans. IRs, on the other hand, which are less frequent in hymenopterans, are strongly expanded in the cockroach and termite genomes (Fig. 3 and Supplementary Fig. 3). Intronless IRs, which are known to be particularly divergent24, show the greatest cockroach- and Blattodea-specific expansions (Fig. 3a, Blattodea-, Cockroach- and Group D-IRs). By far the most IRs among all investigated species were found in B. germanica (455 complete gene models), underlining that the capacity for detecting many different kinds of chemosensory cues is crucial for this generalist that thrives in challenging, human environments. In line with a specialization in diet and habitat, the total number of IRs is lower within the termites (Z. nevadensis: 141; C. secundus: 135; M. natalensis: 75). Nevertheless, IRs are more numerous in termites than in all other analysed species (except Nasonia vitripennis: 111). This is strikingly similar to the pattern for ORs in Hymenoptera, which are also highly numerous in non-eusocial outgroups as well as in eusocial sister species14,23,25.
a,b, IR (a) and OR (b) gene trees of 13 insect species. In each tree, only well-supported clades (support values > 85) that include B. germanica or termite genes are highlighted within the gene trees. The lengths of the coloured bars represent the number of genes per species within each of these clades. The red asterisk in a denotes the putative root of intronless IRs. c, The upper schematic diagram depicts the 2D structure of an IR, containing ligand-binding lobes (S1 and S2), transmembrane regions (TM1–3) and the pore domain (P). Below, the sequence of the domains along the peptide is represented, showing that the sites, which are under significant positive selection (red bars; codeml site models 7 and 8) within Blattodea IRs for M. natalensis (P < 1.7 × 10−10; likelihood-ratio test, 5 sequences, 413 aligned codons), are all situated within the ligand-binding lobes and on or around the putative ligand-binding sites (asterisks)86. d, The same representation for ORs, which include eight transmembrane regions. Positive selection was found for M. natalensis (P = 1.1 × 10−10; 5 sequences, 1,001 aligned codons) and C. secundus (P = 5.6 × 10−16; likelihood ratio test, 26 sequences, 1,913 aligned codons) of the orange clade, each at two codon positions within the second transmembrane region and at a third position within the carboxy-terminal extracellular region for M. natalensis.
We scanned each IR group for signs of species-specific positive selection. Within the Blattodea-specific intronless IRs, we found two codon positions under significant selection for the higher termite M. natalensis (codeml site models 7 and 8; P = 5.4 × 10−5). Within a subgroup of five sequences, this was more strongly pronounced with seven codon positions under significant positive selection for M. natalensis (P < 1.7 × 10−10). The positively evolving codons are situated within the two ligand-binding lobes of the receptors (Fig. 3c), showing that a diversification of ligand specificity has occurred along with the transition to higher eusociality and a change from wood-feeding to fungus-farming in M. natalensis. Only two IRs were differentially expressed between nymphs and adult females in B. germanica. Underlining a change in expression along with the evolution of castes, we found 35 IRs to be differentially expressed between workers and queens in Z. nevadensis, 11 in C. secundus and 10 in M. natalensis (Fig. 2 and Supplementary Table 10). The possible role of IRs in pheromonal communication has been highlighted both in the cockroach Periplaneta americana26 and in Drosophila melanogaster27, where several IRs show sex-biased expression.
One group of ORs (orange clade in Fig. 3b) is evolving under significant positive selection at codon positions within the second transmembrane domain in M. natalensis (codeml site model; P = 1.1 × 10−11) and C. secundus (P = 5.6 × 10−16; Fig. 3d). Such a variation in the transmembrane domain can be related to ligand-binding specificity, as has been shown for a polymorphism in the third transmembrane domain for an OR in D. melanogaster28,29, adding further support for an adaptive evolution of chemoreceptors, in line with the greater need for a sophisticated colony communication in the termites. Similar to IRs, a higher proportion of ORs were differentially expressed between workers and queens in the three termites than between nymphs and adults in the cockroach (Fig. 2 and Supplementary Table 11), highlighting a change in expression and function along with the transition to eusociality. The evolution of chemoreceptors along with the emergence of the termites can also be related to adaptation processes and changes in diet compared to the cockroach. Experimental verification will help pinpoint which receptors are particularly important for communication.
CHC-producing enzymes have evolved caste-specificity
Despite their different ancestry, both termites and eusocial hymenopterans are characterized by the production of caste-specific cuticular hydrocarbons (CHCs)30,31,32, which are often crucial for regulating reproductive division of labour and chemical communication. Accordingly, we find changes in the termites in three groups of proteins involved in the synthesis of CHCs: desaturases (introduction of double bonds33), elongases (extension of C-chain length34) and CYP4G1 (last step of CHC biosynthesis35).
Desaturases are thought to be important for division of labour and social communication in ants36. As previously described for ants36, Desat B genes are the most abundant desaturase family in the termites and the cockroach (Supplementary Table 12), especially in M. natalensis where we found ten gene copies (significant expansion; P = 0.0003; Supplementary Table 5 and Supplementary Figure 4). As in ants, especially the first desaturases (Desat A–Desat E) vary greatly in their expression between castes and species in the three termites (Fig. 2 and Supplementary Table 13)36. In contrast to ants, where these genes are under strong purifying selection36, for the highly eusocial termite M. natalensis, we found significant positive selection within the Desat B genes (codeml site models 7 and 8; P = 1.1 × 10−16), indicating a diversification in function, possibly related to their greater diversification of worker castes (major and minor workers, major and minor soldiers). Although desaturases are often discussed in the context of CHC production and chemical communication, their biochemical roles are quite diverse36, and the positive selection we observe for M. natalensis may, at least in part, be related to their rather different ecology of foraging and fungus-farming rather than nest-mate recognition. Future experimental verification of the function of these genes will help better understand these observed genomic and transcriptomic patterns.
Underlining an increased importance of CHC communication in termites, the expression patterns of elongases (extension of C-chain length) differ considerably in the termites compared to the cockroach (Fig. 2 and Supplementary Table 14). In contrast to B. germanica, in which elongases are both nymph- (five genes) and adult-biased (four genes), only one or two elongase genes in each termite are queen-biased in their expression, while many are worker-biased. As with the desaturases, a group of M. natalensis elongases also reveal significant signals of positive selection (codeml branch-site test; P = 4 × 10−4), further indicating a greater diversification of CHC production in this higher termite.
The last step of CHC biosynthesis, the production of hydrocarbons from long-chain fatty aldehydes, is catalysed by a P450 gene, CYP4G1, in D. melanogaster35. We found one copy of CYP4G1 in B. germanica, Z. nevadensis and C. secundus, but three copies in M. natalensis, reinforcing the greater importance of CHC synthesis in this higher termite. Corroborating the known importance of maternal CHCs in B. germanica37, CYP4G1 is overexpressed in female adults compared to nymphs (Fig. 2 and Supplementary Table 15). In each of the termites, however, CYP4G1 is more highly expressed in workers (or kings in C. secundus) compared to queens (Fig. 2 and Supplementary Table 15), adding support that, compared to cockroach nymphs, a change in the dynamics and turnover of CHCs in termite workers has taken place.
Changes in gene regulation in termites
The development of distinct castes underlying division of labour is achieved via differential gene expression. Major changes in gene regulation have been reported as being central to the transition to eusociality in bees9 and ants10. Accordingly, we found major changes in putative DNA methylation patterns (levels per 1-to-1 orthologue) among the termites compared to four other hemimetabolous insect species (Fig. 4a). This is revealed by CpG depletion patterns (CpGo/e, observed versus expected number of CpGs), a reliable predictor of DNA methylation38,39, correlating more strongly between the termites than among any of the other analysed hemimetabolous insects (Fig. 4). In other words, within orthologous genes, predicted DNA methylation levels differ greatly between termites and other hemimetabolous species but remain conserved among termite species.
a, Principal component analysis (PCA) of predicted DNA methylation patterns among 2,664 1-to-1 orthologues, estimated via CpGo/e. The spheres represent the positions of the species within the 3D PCA, with the distance between the spheres representing the similarity of CpGo/e between species at each orthologue; the curves are the distribution of CpGo/e, with the dotted line showing CpGo/e = 1. b, Tag clouds of enriched (P < 0.05; Fisher test, weight algorithm, topGO96) GO terms (biological processes) among the lower (left) and the higher quartile (right) of CpGo/e within termites (top) and B. germanica (bottom). For termites, genes were merged from all three species for analysing GO term enrichment. Number of enriched genes and total number of genes in topGO enrichment tests (low CpG o/e /high CpGo/e/gene universe): B. germanica (3,291/1,842/11,409); termites (6,754/4,600/25,910). High CpGo/e indicates a low level of DNA methylation and vice versa.
The predicted levels of DNA methylation correlated negatively with the caste-specificity of expression for each of the termites. This is confirmed by a positive correlation between CpGo/e (negative association with level of DNA methylation) and absolute log2(fold change) of expression between queens and workers (Pearson’s r = 0.32 to 0.36; P < 2.2 × 10−16). The caste-specific expression of putatively unmethylated genes in termites is reflected in the enrichment of GO terms related to sensory perception, regulation of transcription, signalling and development, whereas methylated genes are mainly related to general metabolic processes (Fig. 4b and Supplementary Table 16). These results show strong parallels to findings for eusocial Hymenoptera40,41,42,43. This is in stark contrast to the non-eusocial cockroach, B. germanica, where there was only a very weak relationship between CpGo/e and differential expression between nymphs and adult females (r = 0.14), nor were any large differences apparent in enriched GO terms between putatively methylated and non-methylated genes (Fig. 4b).
Our results argue in favour of a diminished role of DNA methylation in caste-specific expression within eusocial insects, as recently shown38,44. In fact, DNA methylation appears to be important for the regulation of housekeeping genes because predicted methylated genes are related to general biological processes (further supported by lower CpGo/e within 1-to-1 orthologues than in non-conserved genes)45, while caste-specific genes are ‘released’ from this type of gene regulation. However, a recent study linked caste-specific DNA methylation to alternative splicing in Z. nevadensis46.
Major biological transitions are often accompanied by expansions of transcription factor (TF) families, such as genes containing zinc-finger (ZF) domains47. We also observed large differences in ZF families within the termites compared to B. germanica. Many ZF families were reduced or absent in termites, while different, unrelated ZF gene families were significantly expanded (Supplementary Tables 2–6). Queen-biased genes were significantly over-represented among ZF genes for each of the termites (P < 2 × 10−10; χ2 test; Supplementary Table 17), indicating an important role of ZF genes in the regulation of genes related to caste-specific tasks and colony organization in the termites. This is in contrast to the significant under-representation of differentially expressed ZF genes within B. germanica (P = 4.8 × 10−5; χ2-test). Interestingly, two other important TF families (bHLH and bZIP)47, which were not expanded in the termites, showed no caste-specific expression pattern (P > 0.05), except bZIP genes, in which queen-biased genes were marginally over-represented for M. natalensis (P = 0.049). These major upheavals in ZF gene families and their caste-specific expression show that major changes in TFs accompanied the evolution of termites, strikingly similar to the evolution of ants10.
Evolution of genes related to moulting and metamorphosis
Hemimetabolous eusociality is characterized by differentiated castes, which represent different developmental stages. This is in contrast to eusocial Hymenoptera, in which workers and reproductives are adults. While cockroaches develop directly through several nymphal stages before becoming reproductive adults, termite development is more phenotypically plastic, and workers are essentially immatures (Fig. 2). In wood-dwelling termites, such as C. secundus and Z. nevadensis, worker castes are non-reproductive immatures that are totipotent to develop into other castes, while in the higher termite M. natalensis, workers can be irreversibly defined instars. It is therefore clear that a major change during the evolution of termites occurred within developmental pathways. Accordingly, we found changes in expression and gene family size of several genes related both to moulting and metamorphosis.
In the synthesis of the moulting hormone, 20-hydroxyecdysone, the six Halloween genes (five cytochrome P450s and a Rieske-domain oxygenase) play a key role48,49. Only one Halloween gene, Shade (Shd; CYP314A1), which mediates the final step of 20-hydroxyecdysone synthesis, is differentially expressed between the final nymphal stages and adult females in B. germanica (Fig. 2 and Supplementary Table 18), consistent with its role in the nymphal or imaginal moult. In the three termites, the Halloween genes show varying caste-specific expression (Fig. 2 and Supplementary Table 18), showing that ecdysone plays a significant role in the regulation of caste differences. Ecdysteroid kinase genes (EcK), which convert the insect moulting hormone into its inactive state, ecdysone 22-phosphate, for storage50, are only overexpressed in female adults compared to nymphs in B. germanica (16/51 genes, Fig. 2 and Supplementary Table 19). In termites, however, where the gene copy number is reduced (18 to 20 per species), these important moulting genes appear to have evolved worker-specific functions (Fig. 2 and Supplementary Table 19).
Whereas 20-hydroxyecdysone promotes moulting, juvenile hormone (JH) represses imaginal development in pre-adult instars51. JH is important in caste differentiation in eusocial insects, including termites12,52. Haemolymph JH-binding proteins (JHBPs), which transport JH to its target tissues53, are reduced within the termites (21 to 33 genes) but significantly expanded in B. germanica (51 copies; P = 0018; Supplementary Table 6). Thirteen of the JHBP genes are overexpressed in adult females and only 8 in nymphs in B. germanica (Fig. 2 and Supplementary Table 20). In both Z. nevadensis and M. natalensis, on the other hand, JHBPs are significantly more worker-biased (P < 0.01, χ2 test; Supplementary Table 20 and Fig. 2). In C. secundus, expression is more varied, with four worker-biased, seven king-biased and two queen-biased genes (Fig. 2 and Supplementary Table 20).
These changes in copy number and caste-specific expression of genes involved in moulting and metamorphosis within termites compared to the German cockroach demonstrate that changes occurred in the control of the developmental pathway along with the evolution of castes. However, this interpretation needs to be experimentally verified.
Conclusions
These results, considered alongside many studies on eusociality in Hymenoptera9,10,14,36, provide evidence that major changes in gene regulation and the evolution of sophisticated chemical communication are fundamental to the transition to eusociality in insects. Strong changes in DNA methylation patterns correlated with broad-scale modifications of expression patterns. Many of these modified expression patterns remained consistent among the three studied termite species and occurred within protein pathways essential for eusocial life, such as CHC production, chemoperception, ecdysteroid synthesis and JH transport. The stronger patterns we observe for M. natalensis, especially within genes linked to chemical communication, such as the expansion of Desat B and CYP4G1 genes and significant positive selection in desaturases, elongases and IRs, may be associated with this termite’s higher level of eusociality and its status as a superoganism13. The analysis of further higher and lower termites would shed light on the generality of these patterns and possibly assist in the distinction between the influences of ecological and eusocial traits.
Many of the mechanisms implicated in the evolution of eusociality in the termites occurred convergently around 50 Myr later in the phylogenetically distant Hymenoptera. However, several details are unique due to the distinct conditions within which eusociality arose. One important difference is the higher TE content within cockroaches and termites, which probably facilitated changes in gene family sizes, supporting the transition to eusociality. However, the most striking difference is the apparent importance of IRs for chemical communication in the termites, compared to ORs in Hymenoptera. According to our results, the non-eusocial ancestors of termites possessed a broad repertoire of IRs, which favoured the evolution of important functions for colony communication in these chemoreceptors within the termites, whereas in the solitary ancestors of eusocial hymenopterans ORs were most abundant14,25. The parallel expansions of different chemoreceptor families in these two independent origins of eusociality indicate that convergent selection pressures existed during the evolution of colony communication in both lineages.
Methods
Genome sequencing and assembly
Genomic DNA from a single Blattella germanica male from an inbred line (strain: American Cyanamid = Orlando Normal) was used to construct two paired-end (180-bp and 500-bp inserts) and one of the two mate-pair libraries (2-kb inserts). An 8-kb mate-pair library was constructed from a single female. The libraries were sequenced on an Illumina HiSeq2000 sequencing platform. The 413 Gb of raw sequence data were assembled with Allpaths LG54, and then scaffolded and gap-filled using the in-house tools Atlas-Link v.1.0 (https://www.hgsc.bcm.edu/software/atlas-link) and Atlas gap-fill v.2.2. For Cryptotermes secundus, three paired-end libraries (250-bp, 500-bp and 800-bp inserts) and three mate-pair libraries (2-kb, 5-kb and 10-kb inserts) were constructed from genomic DNA that was extracted from the head and thorax of 1,000 individuals, originating from a single, inbred field colony. The libraries were sequenced on an Illumina HiSeq2000 sequencing platform. The C. secundus genome was assembled using SOAPdenovo (v.2.04)55 with optimized parameters, followed by gapcloser (v1.10, released with SOAPdenovo) and kgf (v1.18, released with SOAPdenovo).
Transcriptome sequencing and assembly
For annotation purposes, 22 whole-body RNA-sequencing (RNA-Seq) samples from various developmental stages were obtained for B. germanica. For C. secundus, RNA-Seq libraries were obtained for three workers, four queens and four kings, based on degutted, whole-body extracts. In addition, we sequenced ten Macrotermes natalensis RNA-Seq libraries from three queens, one king and six pools of workers. All libraries were constructed using the Illumina (TruSeq) RNA-Seq kit.
For protein-coding gene annotation, B. germanica reads were assembled with de novo Trinity (version r2014-04-13)56. The C. secundus reads were assembled using Cufflinks on reads mapped with TopHat (version2.2.1)57,58, de novo Trinity56 and genome-guided Trinity on reads mapped with TopHat.
Repeat annotation
A custom C. secundus and B. germanica repeat library was constructed using a combination of homology-based and de novo approaches, including RepeatModeler/RepeatClassifier (http://www.repeatmasker.org/RepeatModeler/), LTRharvest/LTRdigest59 and TransposonPSI (http://transposonpsi.sourceforge.net/). The ab initio repeat library was complemented with the RepBase (update 29 August 2016)60 and SINE repeat databases, filtered for redundancy with CD-hit and classified with RepeatClassifier. RepeatMasker (version open-4.0.6, http://www.repeatmasker.org) was used to mask the C. secundus and B. germanica genome. Repeat content for the other studied species (Fig. 1) was obtained from the literature61,62,63,64,65,66,67.
Protein-coding gene annotation
The B. germanica genome was annotated with Maker (version 2.31.8)68, using the species-specific repeat library, B. germanica transcriptome data (22 whole-body RNA-Seq samples) and the Swiss-Prot/UniProt database (last accessed: 21 January 2016) plus the C. secundus and Zootermopsis nevadensis protein sequences for evidence-based gene model predictions. AUGUSTUS (version 3.2)69, GeneMark-ES Suite (version 4.21)70 and SNAP71 were used for ab initio predictions. C. secundus protein-coding genes were predicted using homology-based, ab initio and expression-based methods, and integrated into a final gene set (see Supplementary Information). Gene structures were predicted by GeneWise72. The ab initio annotations were predicted with AUGUSTUS73 and SNAP71, retained if supported by both methods and integrated with the homology-based predictions using GLEAN74. Transcriptome-based gene models were merged with PASA75 and tested for coding potential with CPC76 and OrfPredictor77. PASA gene models were merged with the homology-based and ab initio gene set, retaining the PASA models in case of overlap. Desaturases, elongases, chemosensory receptors, cytochrome P450s and genes involved in the juvenile hormone pathway were manually curated in Blattodea.
Differential gene expression
The C. secundus and M. natalensis RNA-Seq libraries were complemented with nine published Z. nevadensis libraries, yielding two to six libraries from workers, queens and kings for each termite. These were compared to six of the B. germanica libraries: two from fifth instar nymphs, two from sixth instar nymphs and two from adult females. Reads were mapped to the genome using HiSat278. Read counts per gene were obtained using htseq-count and DESeq279 was used for differential expression analysis. Differential expression analysis between kings (males), queens (females) and workers (majors and minors combined for M. natalensis) was assessed for the termites. For B. germanica we evaluated the differential expression between adults and the two last nymphal stages combined, with the assumption that the final nymphal stages are homologous to termite workers and the adult females are homologous to termite queens. Genes were considered significantly differentially expressed if P < 0.05 and log2(fold change) > |1| in order to account for allometric differences as recommended in a previous study80.
Protein orthology
In addition to B. germanica, C. secundus, Z. nevadensis and M. natalensis, 16 other insect proteomes were included in our analyses: Locusta migratoria, Rhodnius prolixus, Ephemera danica, Drosophila melanogaster, Aedes aegypti, Tribolium castaneum, Nasonia vitripennis, Polistes canadensis, Apis mellifera, Harpegnathos saltator, Linepithema humile, Camponotus floridanus, Pogonomyrmex barbatus, Solenopsis invicta, Acromyrmex echinatior and Atta cephalotes; as well as for the centipede Strigamia maritima as an outgroup (for sources, see Supplementary Table 22). These proteomes were grouped into orthologous clusters with OrthoMCL81, with a granularity of 1.5.
IR and OR identification, phylogeny and structure
Ionotropic receptors (IRs) were identified using two custom hidden Markov models (HMMs) obtained with hmmbuild and hmmpress of the HMMER suite82. The first HMM comprises the IR’s ion channel and ligand-binding domain based on a MAFFT83 protein alignment of 76 IRs from 15 species (Supplementary Table 23). The second HMM was built to distinguish IRs from iGluRs, IR8a and IR25a, which have an additional amino-terminal domain24. For this we built an HMM from 48 protein sequences (Supplementary Table 23). The proteomes were scanned with pfam_scan and the two custom HMMs, where proteins that matched the IR HMM, but not the amino-terminal domain HMM were annotated as IRs. Odorant receptors (ORs) were identified on the basis of the Pfam domain PF02949 (7tm OR).
Multiple sequence alignments of IRs and ORs were obtained with hmmalign82, using the Pfam OR HMM PF02949 and custom IR HMM to guide the alignment. Gene trees were computed with FastTree84 (options: -pseudo -spr 4 -mlacc 2 -slownni) and visualized with iTOL v385. Putative IR ligand-binding residues and structural regions were identified on the basis of the alignments with D. melanogaster IRs and iGluRs of known structure86.
Gene family expansions and contractions
For the analyses of gene family expansions and contractions, the hierarchical clustering algorithm MC-UPGMA87 was used, with a ProtoLevel cutoff of 80 (ref. 88). Protein families were further divided into sub-families if they contained more than 100 proteins in a single species, or more than an average of 35 proteins per species. Proteins were blasted against the RepeatMasker TE database (E-value < 10−5) and clusters where > 50% of the proteins were identified as transposable elements were discarded. Clade- and species-specific protein family expansions and contractions, were identified with CAFE v3.089 using the same protocol as in previous studies9,10 (see also Supplementary Information).
TE-facilitated expansions
The repeat content in the 10-kb flanking regions of B. germanica, C. secundus, Z. nevadensis and M. natalensis genes was calculated using bedtools90. Coding DNA sequences (CDSs) from neighbouring genes were removed and the repeat content was analysed using generalized linear mixed models (glmmPQL implemented in the R91 package MASS92) with binomial error distribution. Fixed predictors included gene family expansion, species ID and their interaction. Cluster ID was fitted as a random factor to avoid pseudo-replication. Significance was assessed on the basis of the Wald t-test (R package aod93) at α < 0.05. Main and interaction effects for each of the genomic regions are listed in Supplementary Table 8. Model parameters are listed in Supplementary Table 8.
Tests for positive selection
To test for positive selection within gene families of interest, site model tests 7 and 8 were performed (model = 0; NSsites = 7 8) on species-specific CDS alignments, or branch-site test (model = 2; NSsites = 2; fix_omega = 1 for null model and 0 for alternative model) on multi-species alignments. Protein sequences were aligned using MAFFT83 with the E-INS-i strategy, and CDS alignments were created using pal2nal.pl94. Phylogenetic trees were created with FastTree84. Alignments were trimmed using Gblocks (settings: -b2 = 21; -b3 = 20; -b4 = 5; -b5 = a). Models were compared using likelihood-ratio test and where P < 0.05, Bayes empirical Bayes results were consulted for codon positions under positive selection (P < 0.05).
CpG depletion patterns and GO enrichment
To estimate DNA methylation, we compared observed to expected CpG counts within CDS sequences38,39. A low CpGo/e indicates a high level of DNA methylation, as the cytosines of methylated CpGs often mutate to thymines. Expected CpG counts were calculated by dividing the product of cytosine and guanine counts by the sequence length. The principal component analysis in Fig. 4 was created using the R function prcomp on log-transformed CpGo/e values for all 1-to-1 orthologues for the seven hemimetabolous species. These orthologues were extracted from the OrthoMCL results. The three-dimensional (3D) plot was created with the plot3d command from the R package rgl.
CpG-depleted (first quartile) and -enriched (fourth quartile) genes were tested for enrichment of Gene Ontology terms. Pfam protein domains were obtained for B. germanica, Z. nevadensis, C. secundus and M. natalensis protein sequences using PfamScan95. Corresponding GO terms were obtained with Pfam2GO. GO-term over-representation was assessed using the TopGO96 package in R. Enrichment analysis was performed using the weight algorithm selecting nodesize = 10 to remove terms with fewer than ten annotated GO terms. After that, GO terms classified as significant (topGOFisher < 0.05) were visualized using the R package tagcloud (https://cran.r-project.org/web/packages/tagcloud/).
Life Science Reporting Summary
Further information on experimental design is available in the Life Sciences Reporting Summary.
Code availability
All custom-made scripts used in these analyses are available at the following repository: https://github.com/ebbgroup/Genomic-comparisons-in-Blattodea.
Data availability
The genome assembly of Blattella germanica is archived on NCBI under the accession PRJNA203136. The genome assembly of Cryptotermes secundus is available on NCBI under the accession PRJNA381866. The additionally annotated genes for Z. nevadensis and M. natalensis are available from the Dryad Digital Repository: https://doi.org/10.5061/dryad.51d4r. Transcriptomic reads generated in this study are available in SRA (B. germanica: PRJNA382128; C. secundus: PRJNA382129; M. natalensis: PRJNA382034).
References
Szathmáry, E. & Maynard Smith, J. The major evolutionary transitions. Nature 374, 227–232 (1995).
Andersson, M. The evolution of eusociality. Annu. Rev. Ecol. Syst. 15, 165–189 (1984).
Wilson, E. O. The Insect Societies (Harvard University Press, Cambridge, 1971).
Rubenstein, D. R. & Abbot, P. (eds) Comparative Social Evolution (Cambridge University Press, Cambridge, 2017).
Misof, B. et al. Phylogenomics resolves the timing and pattern of insect evolution. Science 346, 763–767 (2014).
Legendre, F. et al. Phylogeny of Dictyoptera: dating the origin of cockroaches, praying mantises and termites with molecular data and controlled fossil evidence. PLoS One 10, e0130127 (2015).
Bourguignon, T. et al. The evolutionary history of termites as inferred from 66 mitochondrial genomes. Mol. Biol. Evol. 32, 406–421 (2015).
Elsner, D., Kremer, L. P., Arning, N. & Bornberg-Bauer, E. Comparative genomic approaches to investigate molecular traits specific to social insects. Curr. Opin. Insect Sci. 16, 87–94 (2016).
Kapheim, K. M. et al. Genomic signatures of evolutionary transitions from solitary to group living. Science 348, 1139–1143 (2015).
Simola, D. F. et al. Social insect genomes exhibit dramatic evolution in gene composition and regulation while preserving regulatory features linked to sociality. Genome Res. 23, 1235–1247 (2013).
Woodard, S. H. et al. Genes involved in convergent evolution of eusociality in bees. Proc. Natl. Acad. Sci. USA 108, 7472–7477 (2011).
Korb, J. & Hartfelder, K. Life history and development - a framework for understanding developmental plasticity in lower termites. Biol. Rev. 83, 295–313 (2008).
Boomsma, J. J. & Gawne, R. Superorganismality and caste differentiation as points of no return: how the major evolutionary transitions were lost in translation. Biol. Rev. 93, 28–54 (2018).
Zhou, X. et al. Chemoreceptor evolution in Hymenoptera and its implications for the evolution of eusociality. Genome Biol. Evol. 7, 2407–2416 (2015).
Trible, W. et al. Orco mutagenesis causes loss of antennal lobe glomeruli and impaired social behavior in ants. Cell 170, 727–735.e10 (2017).
Poulsen, M. et al. Complementary symbiont contributions to plant decomposition in a fungus-farming termite. Proc. Natl Acad. Sci. USA 111, 14500–14505 (2014).
Terrapon, N. et al. Molecular traces of alternative social organization in a termite genome. Nat. Commun. 5, 3636 (2014).
Gregory, T. R. Animal Genome Size Database (accessed 25 November 2017); http://www.genomesize.com/.
Ferreira, P. G. et al. Transcriptome analyses of primitively eusocial wasps reveal novel insights into the evolution of sociality and the origin of alternative phenotypes. Genome Biol. 14, R20 (2013).
Korb, J. et al. A genomic comparison of two termites with different social complexity. Front. Genet. 6, 9 (2015).
Kazazian, H. H. Mobile elements: drivers of genome evolution. Science 303, 1626–1632 (2004).
Joseph, R. M. & Carlson, J. R. Drosophila chemoreceptors: a molecular interface between the chemical world and the brain. Trends Genet. 31, 683–695 (2015).
Brand, P. & Ramírez, S. R. The evolutionary dynamics of the odorant receptor gene family in corbiculate bees. Genome Biol. Evol. 9, 2023–2036 (2017).
Croset, V. et al. Ancient protostome origin of chemosensory ionotropic glutamate receptors and the evolution of insect taste and olfaction. PLoS. Genet. 6, e1001064 (2010).
Robertson, H. M., Gadau, J. & Wanner, K. W. The insect chemoreceptor superfamily of the parasitoid jewel wasp Nasonia vitripennis. Insect Mol. Biol. 19, 121–136 (2010).
Chen, Y., He, M., Li, Z.-Q., Zhang, Y.-N. & He, P. Identification and tissue expression profile of genes from three chemoreceptor families in an urban pest, Periplaneta americana. Sci. Rep. 6, 27495 (2016).
Koh, T.-W. et al. The Drosophila IR20a clade of ionotropic receptors are candidate taste and pheromone receptors. Neuron 83, 850–865 (2014).
Pellegrino, M., Steinbach, N., Stensmyr, M. C., Hansson, B. S. & Vosshall, L. B. A natural polymorphism alters odour and DEET sensitivity in an insect odorant receptor. Nature 478, 511–514 (2011).
Nichols, A. S. & Luetje, C. W. Transmembrane segment 3 of Drosophila melanogaster odorant receptor subunit 85b contributes to ligand-receptor interactions. J. Biol. Chem. 285, 11854–11862 (2010).
Oystaeyen, A. V. et al. Conserved class of queen pheromones stops social insect workers from reproducing. Science 343, 287–290 (2014).
Weil, T., Hoffmann, K., Kroiss, J., Strohm, E. & Korb, J. Scent of a queen–cuticular hydrocarbons specific for female reproductives in lower termites. Naturwissenschaften 96, 315–319 (2009).
Dietemann, V., Peeters, C., Liebig, J., Thivet, V. &Hölldobler, B. Cuticular hydrocarbons mediate discrimination of reproductives and nonreproductives in the ant Myrmecia gulosa. Proc. Natl Acad. Sci. USA 100, 10341–10346 (2003).
Dallerac, R. et al. A Δ9 desaturase gene with a different substrate specificity is responsible for the cuticular diene hydrocarbon polymorphism in Drosophila melanogaster. Proc. Natl Acad. Sci. USA 97, 9449–9454 (2000).
Finck, J., Berdan, E. L., Mayer, F., Ronacher, B. & Geiselhardt, S. Divergence of cuticular hydrocarbons in two sympatric grasshopper species and the evolution of fatty acid synthases and elongases across insects. Sci. Rep. 6, 33695 (2016).
Qiu, Y. et al. An insect-specific P450 oxidative decarbonylase for cuticular hydrocarbon biosynthesis. Proc. Natl. Acad. Sci. USA 109, 14858–14863 (2012).
Helmkampf, M., Cash, E. & Gadau, J. Evolution of the insect desaturase gene family with an emphasis on social Hymenoptera. Mol. Biol. Evol. 32, 456–471 (2015).
Fan, Y., Eliyahu, D. & Schal, C. Cuticular hydrocarbons as maternal provisions in embryos and nymphs of the cockroach Blattella germanica. J. Exp. Biol. 211, 548–554 (2008).
Bewick, A. J., Vogel, K. J., Moore, A. J. & Schmitz, R. J. Evolution of DNA methylation across insects. Mol. Biol. Evol. 34, 654–655 (2017).
Park, J. et al. Comparative analyses of DNA methylation and sequence evolution using Nasonia genomes. Mol. Biol. Evol. 28, 3345–3354 (2011).
Elango, N., Hunt, B. G., Goodisman, M. A. D. & Yi, S. V. DNA methylation is widespread and associated with differential gene expression in castes of the honeybee, Apis mellifera. Proc. Natl. Acad. Sci. USA 106, 11206–11211 (2009).
Standage, D. S. et al. Genome, transcriptome and methylome sequencing of a primitively eusocial wasp reveal a greatly reduced DNA methylation system in a social insect. Mol. Ecol. 25, 1769–1784 (2016).
Patalano, S. et al. Molecular signatures of plastic phenotypes in two eusocial insect species with simple societies. Proc. Natl. Acad. Sci. USA 112, 13970–13975 (2015).
Rehan, S. M., Glastad, K. M., Lawson, S. P. & Hunt, B. G. The genome and methylome of a subsocial small carpenter bee, Ceratina calcarata. Genome Biol. Evol. 8, 1401–1410 (2016).
Libbrecht, R., Oxley, P. R., Keller, L. & Kronauer, D. J. C. Robust DNA methylation in the clonal raider ant brain. Curr. Biol. 26, 391–395 (2016).
Foret, S., Kucharski, R., Pittelkow, Y., Lockett, G. A. & Maleszka, R. Epigenetic regulation of the honey bee transcriptome: unravelling the nature of methylated genes. BMC Genom. 10, 472 (2009).
Glastad, K. M., Gokhale, K., Liebig, J. & Goodisman, M. A. D. The caste- and sex-specific DNA methylome of the termite Zootermopsis nevadensis. Sci. Rep. 6, 37110 (2016).
Schmitz, J. F., Zimmer, F. & Bornberg-Bauer, E. Mechanisms of transcription factor evolution in Metazoa. Nucleic Acids Res. 44, 6287–6297 (2016).
Rewitz, K. F., Rybczynski, R., Warren, J. T. & Gilbert, L. I. The Halloween genes code for cytochrome P450 enzymes mediating synthesis of the insect moulting hormone. Biochem. Soc. Trans. 34, 1256–1260 (2006).
Lang, M. et al. Mutations in the neverland gene turned Drosophila pachea into an obligate specialist species. Science 337, 1658–1661 (2012).
Sonobe, H. et al. Purification, kinetic characterization, and molecular cloning of a novel enzyme, ecdysteroid 22-kinase. J. Biol. Chem. 281, 29513–29524 (2006).
Jindra, M., Belles, X. & Shinoda, T. Molecular basis of juvenile hormone signaling. Curr. Opin. Insect Sci. 11, 39–46 (2015).
Korb, J. in Genomics, Physiology and Behaviour of Social Insects Vol. 48 (eds Zayed, A. & Kent, C. F.) 131–161 (Academic Press, 2015).
Kolodziejczyk, R. et al. Insect juvenile hormone binding protein shows ancestral fold present in human lipid-binding proteins. J. Mol. Biol. 377, 870–881 (2008).
Gnerre, S. et al. High-quality draft assemblies of mammalian genomes from massively parallel sequence data. Proc. Natl. Acad. Sci. USA 108, 1513–1518 (2011).
Li, Y., Hu, Y., Bolund, L. & Wang, J. State of the art de novo assembly of human genomes from massively parallel sequencing data. Hum. Genomics 4, 271–277 (2010).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Roberts, A., Trapnell, C., Donaghey, J., Rinn, J. L. & Pachter, L. Improving RNA-Seq expression estimates by correcting for fragment bias. Genome Biol. 12, R22 (2011).
Ellinghaus, D., Kurtz, S. & Willhoeft, U. LTRharvest, an efficient and flexible software for de novo detection of LTR retrotransposons. BMC Bioinforma. 9, 18 (2008).
Bao, W., Kojima, K. K. & Kohany, O. Repbase Update, a database of repetitive elements in eukaryotic genomes. Mob. DNA 6, 11 (2015).
Chipman, A. D. et al. The first myriapod genome sequence reveals conservative arthropod gene content and genome organisation in the centipede Strigamia maritima. PLoS Biol. 12, e1002005 (2014).
Mesquita, R. D. et al. Genome of Rhodnius prolixus, an insect vector of Chagas disease, reveals unique adaptations to hematophagy and parasite infection. Proc. Natl. Acad. Sci. USA 112, 14936–14941 (2015).
Nene, V. et al. Genome sequence of Aedes aegypti, a major arbovirus vector. Science 316, 1718–1723 (2007).
The Honeybee Genome Sequencing Consortium. Insights into social insects from the genome of the honeybee Apis mellifera. Nature 443, 931–949 (2006).
Gadau, J. et al. The genomic impact of 100 million years of social evolution in seven ant species. Trends Genet.. 28, 14–21 (2012).
Richards, S. et al. The genome of the model beetle and pest Tribolium castaneum. Nature 452, 949–955 (2008).
Wang, X. et al. The locust genome provides insight into swarm formation and long-distance flight. Nat. Commun. 5, 2957 (2014).
Holt, C. & Yandell, M. MAKER2: an annotation pipeline and genome-database management tool for second-generation genome projects. BMC Bioinforma. 12, 491 (2011).
Keller, O., Kollmar, M., Stanke, M. & Waack, S. A novel hybrid gene prediction method employing protein multiple sequence alignments. Bioinformatics 27, 757–763 (2011).
Lomsadze, A., Ter-Hovhannisyan, V., Chernoff, Y. O. & Borodovsky, M. Gene identification in novel eukaryotic genomes by self-training algorithm. Nucleic Acids Res. 33, 6494–6506 (2005).
Korf, I. Gene finding in novel genomes. BMC Bioinforma. 5, 59 (2004).
Birney, E., Clamp, M. & Durbin, R. GeneWise and Genomewise. Genome Res. 14, 988–995 (2004).
Stanke, M. et al. AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic Acids Res. 34, W435–W439 (2006).
Elsik, C. G. et al. Creating a honey bee consensus gene set. Genome Biol. 8, R13 (2007).
Campbell, M. A., Haas, B. J., Hamilton, J. P., Mount, S. M. & Buell, C. R. Comprehensive analysis of alternative splicing in rice and comparative analyses with Arabidopsis. BMC Genomics 7, 327 (2006).
Kong, L. et al. CPC: assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res. 35, W345–W349 (2007).
Min, X. J., Butler, G., Storms, R. & Tsang, A. OrfPredictor: predicting protein-coding regions in EST-derived sequences. Nucleic Acids Res. 33, W677–W680 (2005).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11, 1650–1667 (2016).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Montgomery, S. H. & Mank, J. E. Inferring regulatory change from gene expression: the confounding effects of tissue scaling. Mol. Ecol. 25, 5114–5128 (2016).
Li, L., Stoeckert, C. J. & Roos, D. S. OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res. 13, 2178–2189 (2003).
Eddy, S. R. Profile hidden Markov models. Bioinformatics 14, 755–763 (1998).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree: computing large minimum evolution trees with profiles instead of a distance matrix. Mol. Biol. Evol. 26, 1641–1650 (2009).
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 44, W242–W245 (2016).
Benton, R., Vannice, K. S., Gomez-Diaz, C. & Vosshall, L. B. Variant ionotropic glutamate receptors as chemosensory receptors in Drosophila. Cell 136, 149–162 (2009).
Loewenstein, Y., Portugaly, E., Fromer, M. & Linial, M. Efficient algorithms for accurate hierarchical clustering of huge datasets: tackling the entire protein space. Bioinformatics 24, i41–i49 (2008).
Rappoport, N., Linial, N. & Linial, M. ProtoNet: charting the expanding universe of protein sequences. Nat. Biotechnol. 31, 290–292 (2013).
Han, M. V., Thomas, G. W. C., Lugo-Martinez, J. & Hahn, M. W. Estimating gene gain and loss rates in the presence of error in genome assembly and annotation using CAFE 3. Mol. Biol. Evol. 30, 1987–1997 (2013).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, 2014).
Venables, W. & Ripley, B. Modern Applied Statistics with S, 4th edn (Springer, New York, 2002).
Lesnoff, M. & Lancelot, R. aod: Analysis of Overdispersed Data R Package Version 1.3 (2012); http://cran.r-project.org/package=aod
Suyama, M., Torrents, D. & Bork, P. PAL2nal: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res. 34, W609–W612 (2006).
Finn, R. D. et al. The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res. 44, D279–D285 (2016).
Alexa, A. & Rahnenführer, J. topGO: Enrichment Analysis for Gene Ontology R package version 2.30.0 (2016); http://bioconductor.org/packages/release/bioc/html/topGO.html
Bell, W. J., Roth, L. M. & Nalepa, C. A. Cockroaches: Ecology, Behavior, and Natural History (JHU Press, Baltimore, 2007).
Acknowledgements
We thank O. Niehuis for allowing use of the unpublished E. danica genome, J. Gadau and C. Smith for comments and advice on the manuscript, and J. Schmitz for assistance with analyses and proofreading the manuscript. J.K. thanks Charles Darwin University (Australia), especially S. Garnett and the Horticulture and Aquaculture team, for providing logistic support to collect C. secundus. The Parks and Wildlife Commission, Northern Territory, the Department of the Environment, Water, Heritage and the Arts gave permission to collect (Permit number 36401) and export (Permit WT2010-6997) the termites. USDA is an equal opportunity provider and employer. M.C.H. and E.J. are supported by DFG grant BO2544/11-1 to E.B.-B. J.K. is supported by University of Osnabrück and DFG grant KO1895/16-1. X.B. and M.-D.P. are supported by Spanish Ministerio de Economía y Competitividad (CGL2012-36251 and CGL2015-64727-P to X.B., and CGL2016-76011-R to M.-D.P.), including FEDER funds, and by Catalan Government (2014 SGR 619). C.S. is supported by grants from the US Department of Housing and Urban Development (NCHHU-0017-13), the National Science Foundation (IOS-1557864), the Alfred P. Sloan Foundation (2013-5-35 MBE), the National Institute of Environmental Health Sciences (P30ES025128) to the Center for Human Health and the Environment, and the Blanton J. Whitmire Endowment. M.P. is supported by a Villum Kann Rasmussen Young Investigator Fellowship (VKR10101).
Author information
Authors and Affiliations
Contributions
E.B.-B. conceived, managed and coordinated the project; M.C.H., E.J. and H.M.R. are joint first authors. J.K. conceived and managed the C. secundus sequencing project, and coordinated termite-related analyses; S.R. conceived and managed the B. germanica sequencing project; S.R., S.D., S.L.L., H.C., H.V.D., H.D., Y.H., J.Q., S.C.M., D.S.T.H., K.C.W., D.M.M. and R.A.G. carried out B. germanica library construction, genome sequencing and assembly; C.S. and A.W-.K. provided biological material through full-sib mating for B. germanica; X.B. and C.S. co-managed the B. germanica analysis; M.P. and C.P.C. implemented Web Apollo data traces; S.O. and M.P. provided biological material for M. natalensis; C.G., J.G., J.M.M.-K., A.M., F.S., H.H. and J.K. coordinated and carried out DNA and RNA sequencing for C. secundus; M-D.P., X.B. and G.Y. coordinated transcriptome sequencing of B. germanica; L.M. performed automated gene prediction on C. secundus; E.J. and N.A. improved assembly and annotation for B. germanica & C. secundus, and compared and analysed genome sizes and quality. E.J., N.A. and L.P.M.K. analysed TEs; M.C.H. analysed CpG patterns and signatures of selection; T.B-F., E.J., C.K., L.P.M.K. and A.L-E. performed orthology and phylogenetic analyses; L.P.M.K., E.J., H.M.R. and M.C.H. analysed gene family evolution; A.L-E., E.J. and M.C.H. analysed transcriptomes and performed differential expression analyses; T.B.-F. and A.L-E. carried out orthoMCL clustering; H.M.R. corrected gene models for chemoreceptors; C.K. and E.J. corrected gene models for desaturases and elongases; A-K.H. and M.C.H. corrected gene models for cytochrome P450s; E.B.-B. and M.C.H drafted and wrote the manuscript; X.B., M.-D.P. and J.K. contributed to sections of the manuscript; E.J., L.P.M.K., A.L-E., C.K. and M.C.H. wrote and organized the Supplementary Information; L.P.M.K., N.A., A.L-E., M.C.H. and E.B.-B. prepared figures for the manuscript. All authors read, corrected and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Methods, Supplementary Figures 1-6, Supplementary Tables 1-23.
Supplementary Data 1
A1_mnat_blattodeaIRS_site_test: 3 Output of codeml site test on Macrotermes natalensis Blattodea-IR genes.A2_mnat_OR_site_test: 8 Output of codeml site test on Macrotermes natalensis Orange OR genes. A3_csec_OR_site_test: 13 Output of codeml site test on Cryptotermes secundus Orange OR genes. A4_mnat_desatB_site_test: 18 Output of codeml site test on Macrotermes natalensis Desat B genes. A5_mnat_elongase_branch_site_test: 23 Tree and output of codeml branch-site test on Macrotermes natalensis elongase genes. A6_IR_hidden_Markov_models.rar: 28 Hidden Markov models for IRs.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Harrison, M.C., Jongepier, E., Robertson, H.M. et al. Hemimetabolous genomes reveal molecular basis of termite eusociality. Nat Ecol Evol 2, 557–566 (2018). https://doi.org/10.1038/s41559-017-0459-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-017-0459-1
This article is cited by
-
A genetic toolkit underlying the queen phenotype in termites with totipotent workers
Scientific Reports (2024)
-
Oral-based nanoparticle-wrapped dsRNA delivery system: a promising approach for controlling an urban pest, Blattella germanica
Journal of Pest Science (2024)
-
The remoulding of dietary effects on the fecundity / longevity trade-off in a social insect
BMC Genomics (2023)
-
Pest status, molecular evolution, and epigenetic factors derived from the genome assembly of Frankliniella fusca, a thysanopteran phytovirus vector
BMC Genomics (2023)
-
Caste-specific expressions and diverse roles of takeout genes in the termite Reticulitermes speratus
Scientific Reports (2023)