Abstract
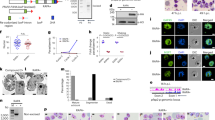
Success in eliminating malaria will depend on whether parasite evolution outpaces control efforts. Here, we show that Plasmodium falciparum parasites (the deadliest of the species causing human malaria) found in low-transmission-intensity areas have evolved to invest more in transmission to new hosts (reproduction) and less in within-host replication (growth) than parasites found in high-transmission areas. At the cellular level, this adaptation manifests as increased production of reproductive forms (gametocytes) early in the infection at the expense of processes associated with multiplication inside red blood cells, especially membrane transport and protein trafficking. At the molecular level, this manifests as changes in the expression levels of genes encoding epigenetic and translational machinery. Specifically, expression levels of the gene encoding AP2-G—the transcription factor that initiates reproduction—increase as transmission intensity decreases. This is accompanied by downregulation and upregulation of genes encoding HDAC1 and HDA1—two histone deacetylases that epigenetically regulate the parasite’s replicative and reproductive life-stage programmes, respectively. Parasites in reproductive mode show increased reliance on the prokaryotic translation machinery found inside the plastid-derived organelles. Thus, our dissection of the parasite’s adaptive regulatory architecture has identified new potential molecular targets for malaria control.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
26 August 2020
In the HTML version of the article, the hyperlink for Supplementary Data 4 incorrectly directed to Supplementary Data 6. The PDF version was unaffected. This has now been corrected.
References
Stearns, S. C. The Evolution of Life Histories (Oxford Univ. Press, New York, 1992).
Reece, S. E., Ramiro, R. S. & Nussey, D. H. Plastic parasites: sophisticated strategies for survival and reproduction? Evol. Appl. 2, 11–23 (2009).
Mideo, N. & Day, T. On the evolution of reproductive restraint in malaria. Proc. R. Soc. B 275, 1217–1224 (2008).
Gandon, S., Mackinnon, M. J., Nee, S. & Read, A. F. Imperfect vaccines and the evolution of parasite virulence. Nature 414, 751–755 (2001).
Noor, A. M. et al. The changing risk of Plasmodium falciparum malaria infection in Africa: 2000–10: a spatial and temporal analysis of transmission intensity. Lancet 383, 1739–1747 (2014).
Kirk, K. & Lehane, A. M. Membrane transport in the malaria parasite and its host erythrocyte. Biochem. J. 457, 1–18 (2014).
El Bissati, K. et al. The plasma membrane permease PfNT1 is essential for purine salvage in the human malaria parasite Plasmodium falciparum. Proc. Natl Acad. Sci. USA 103, 9286–9291 (2006).
Saliba, K. J. & Kirk, K. pH regulation in the intracellular malaria parasite, Plasmodium falciparum. H(+) extrusion via a V-type H(+)-ATPase. J. Biol. Chem. 274, 33213–33219 (1999).
Silvestrini, F. et al. Protein export marks the early phase of gametocytogenesis of the human malaria parasite Plasmodium falciparum. Mol. Cell. Proteom. 9, 1437–1448 (2010).
Kafsack, B. F. et al. A transcriptional switch underlies commitment to sexual development in malaria parasites. Nature 507, 248–252 (2014).
Painter, H. J., Campbell, T. L. & Llinas, M. The Apicomplexan AP2 family: integral factors regulating Plasmodium development. Mol. Biochem. Parasitol. 176, 1–7 (2011).
Coleman, B. I. et al. A Plasmodium falciparum histone deacetylase regulates antigenic variation and gametocyte conversion. Cell Host Microbe 16, 177–186 (2014).
Smith, J. D. et al. Switches in expression of Plasmodium falciparumvar genes correlate with changes in antigenic and cytoadherent phenotypes of infected erythrocytes. Cell 82, 101–110 (1995).
Recker, M. et al. Transient cross-reactive immune responses can orchestrate antigenic variation in malaria. Nature 429, 555–558 (2004).
Zhang, M., Joyce, B. R., Sullivan, W. J. Jr & Nussenzweig, V. Translational control in Plasmodium and Toxoplasma parasites. Eukaryot. Cell 12, 161–167 (2013).
Sheiner, L., Vaidya, A. B. & McFadden, G. I. The metabolic roles of the endosymbiotic organelles of Toxoplasma and Plasmodium spp. Curr. Opin. Microbiol. 16, 452–458 (2013).
Wilson, R. J. M., Gardner, M. J., Feagin, J. E. & Williamson, D. H. Have malaria parasites three genomes? Parasitol. Today 7, 134–136 (1991).
Chaubey, S., Kumar, A., Singh, D. & Habib, S. The apicoplast of Plasmodium falciparum is translationally active. Mol. Microbiol. 56, 81–89 (2005).
Pino, P. et al. Mitochondrial translation in absence of local tRNA aminoacylation and methionyl tRNA Met formylation in Apicomplexa. Mol. Microbiol. 76, 706–718 (2010).
Yeh, E. & DeRisi, J. L. Chemical rescue of malaria parasites lacking an apicoplast defines organelle function in blood-stage Plasmodium falciparum. PLoS Biol. 9, e1001138 (2011).
Painter, H. J., Morrisey, J. M., Mather, M. W. & Vaidya, A. B. Specific role of mitochondrial electron transport in blood-stage Plasmodium falciparum. Nature 446, 88–91 (2007).
Ke, H. et al. The heme biosynthesis pathway is essential for Plasmodium falciparum development in mosquito stage but not in blood stages. J. Biol. Chem. 289, 34827–34837 (2014).
Van Schaijk, B. C. et al. Type II fatty acid biosynthesis is essential for Plasmodium falciparum sporozoite development in the midgut of Anopheles mosquitoes. Eukaryot. Cell 13, 550–559 (2014).
Wiley, J. D. et al. Isoprenoid precursor biosynthesis is the essential metabolic role of the apicoplast during gametocytogenesis in Plasmodium falciparum. Eukaryot. Cell 14, 128–139 (2015).
Gisselberg, J. E., Dellibovi-Ragheb, T. A., Matthews, K. A., Bosch, G. & Prigge, S. T. The suf iron-sulfur cluster synthesis pathway is required for apicoplast maintenance in malaria parasites. PLoS Pathog. 9, e1003655 (2013).
Jacot, D., Waller, R. F., Soldati-Favre, D., MacPherson, D. A. & MacRae, J. I. Apicomplexan energy metabolism: carbon source promiscuity and the quiescence hyperbole. Trends Parasitol. 32, 56–70 (2016).
Lang-Unnasch, N. & Murphy, A. D. Metabolic changes of the malaria parasite during the transition from the human to the mosquito host. Annu. Rev. Microbiol. 52, 561–590 (1998).
MacRae, J. I. et al. Mitochondrial metabolism of sexual and asexual blood stages of the malaria parasite Plasmodium falciparum. BMC Biol. 11, 67 (2013).
Delves, M. et al. The activities of current antimalarial drugs on the life cycle stages of Plasmodium: a comparative study with human and rodent parasites. PLoS Med. 9, e1001169 (2012).
Goodman, C. D. et al. Parasites resistant to the antimalarial atovaquone fail to transmit by mosquitoes. Science 352, 349–353 (2016).
Gunderson, J. H. et al. Structurally distinct, stage-specific ribosomes occur in Plasmodium. Science 238, 933–937 (1987).
Démbéle, L. et al. Persistence and activation of malaria hypnozoites in long-term primary hepatocyte cultures. Nat. Med. 20, 307–312 (2014).
Mancio-Silva, L., Lopez-Rubio, J. J., Claes, A. & Scherf, A. Sir2a regulates rDNA transcription and multiplication rate in the human malaria parasite Plasmodium falciparum. Nat. Commun. 4, 1530 (2013).
Zhang, M. et al. PK4, a eukaryotic initiation factor 2alpha(eIF2alpha) kinase, is essential for the development of the erythrocytic cycle of Plasmodium. Proc. Natl Acad. Sci. USA 109, 3956–3961 (2012).
Zhang, M. et al. The Plasmodium eukaryotic initiation factor-2alpha kinase IK2 controls the latency of sporozoites in the mosquito salivary glands. J. Exp. Med. 207, 1465–1474 (2010).
Babbitt, S. E. et al. Plasmodium falciparum responds to amino acid starvation by entering into a hibernatory state. Proc. Natl Acad. Sci. USA 109, E3278–E3287 (2012).
Bunnik, E. M. et al. Polysome profiling reveals translational control of gene expression in the human malaria parasite Plasmodium falciparum. Genome Biol. 14, R128 (2013).
Caro, F., Ahyong, V., Betegon, M. & DeRisi, J. L. Genome-wide regulatory dynamics of translation in the Plasmodium falciparum asexual blood stages. eLife 3, e04106 (2014).
Beilsten-Edmands, V. et al. eIF2 interactions with initiator tRNA and eIF2B are regulated by post-translational modifications and conformational dynamics. Cell Discov. 1, 15020 (2015).
Greischar, M. A., Mideo, N., Read, A. F. & Bjornstad, O. N. Predicting optimal transmission investment in malaria parasites. Evolution 70, 1542–1558 (2016).
Mackinnon, M. J. & Read, A. F. Virulence in malaria: an evolutionary viewpoint. Phil. Trans. R. Soc. Lond. B 359, 965–986 (2004).
Mackinnon, M. J. & Read, A. F. Immunity promotes virulence evolution in a malaria model. PLoS Biol. 2, E230 (2004).
Mackinnon, M. J. & Marsh, K. The selection landscape of malaria parasites. Science 328, 866–871 (2010).
Greischar, M. A., Mideo, N., Read, A. F. & Bjornstad, O. N. Quantifying transmission investment in malaria parasites. PLoS Comput. Biol. 12, e1004718 (2016).
Buckling, A. G. L., Crooks, L. & Read, A. F. Plasmodium chabaudi: effect of antimalairal drugs on gametocytogenesis. Exp. Parasitol. 93, 45–54 (1999).
Reece, S. E., Duncan, A. B., West, S. A. & Read, A. F. Host cell preference and variable transmission strategies in malaria parasites. Proc. R. Soc. B 272, 511–517 (2005).
Ke, H. et al. Variation among Plasmodium falciparum strains in their reliance on mitochondrial electron transport chain function. Eukaryot. Cell 10, 1053–1061 (2011).
Daily, J. P. et al. In vivo transcriptional profiling of Plasmodium falciparum. Malar. J. 3, 30 (2004).
Mobegi, V. A. et al. Genome-wide analysis of selection on the malaria parasite Plasmodium falciparum in West African populations of differing infection endemicity. Mol. Biol. Evol. 31, 1490–1499 (2014).
Taylor, L. H. & Read, A. F. Why so few transmission stages? Reproductive restraint by malaria parasites. Parasitol. Today 13, 135–140 (1997).
Mackinnon, M. J. et al. Comparative transcriptional and genomic analysis of Plasmodium falciparum field isolates. PLoS Pathog. 5, e1000644 (2009).
Bozdech, Z. et al. The transcriptome of the intraerythrocytic developmental cycle of Plasmodium falciparum. PLoS Biol. 1, 85–100 (2003).
Mok, S. et al. Artemisinin resistance in Plasmodium falciparum is associated with an altered temporal pattern of transcription. BMC Genom. 12, 391 (2011).
Smyth, G. K. in Bioinformatics and Computational Biology Solutions Using R and Bioconductor (eds Gentleman, R. et al.) 397–420 (Springer, New York, 2005).
R Development Core Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, 2015).
Glynn, E. F., Chen, J. & Mushegian, A. R. Detecting periodic patterns in unevenly spaced gene expression time series using Lomb–Scargle periodograms. Bioinformatics 22, 310–316 (2006).
Smyth, G. K., Michaud, J. & Scott, H. The use of within-array replicate spots for assessing differential expression in microarray experiments. Bioinformatics 21, 2067–2075 (2005).
Lopez-Barragan, M. J. et al. Directional gene expression and antisense transcripts in sexual and asexual stages of Plasmodium falciparum. BMC Genom. 12, 587 (2011).
Jackson, A. L., Inger, R., Parnell, A. C. & Bearhop, S. Comparing isotopic niche widths among and within communities: SIBER–Stable Isotope Bayesian Ellipses in R. J. Anim. Ecol. 80, 595–602 (2011).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 559 (2008).
Langfelder, P. & Horvath, S. Tutorials for the WGCNA Package; https://labs.genetics.ucla.edu/horvath/CoexpressionNetwork/Rpackages/WGCNA/Tutorials/index.html.
Csardi, G. & Nepusz, T. The igraph software package for complex network research. InterJournal Complex Systems 1695, 1–9 (2006).
Vembar, S. S., Macpherson, C. R., Sismeiro, O., Coppee, J. Y. & Scherf, A. The PfAlba1 RNA-binding protein is an important regulator of translational timing in Plasmodium falciparum blood stages. Genome Biol. 16, 212 (2015).
Muller, K., Matuschewski, K. & Silvie, O. The Puf-family RNA-binding protein Puf2 controls sporozoite conversion to liver stages in the malaria parasite. PLoS ONE 6, e19860 (2011).
Miao, J. et al. The Puf-family RNA-binding protein PfPuf2 regulates sexual development and sex differentiation in the malaria parasite Plasmodium falciparum. J. Cell Sci. 123, 1039–1049 (2010).
Gu, Z., Gu, L., Eils, R., Schlesner, M. & Brors, B. circlize implements and enhances circular visualization in R. Bioinformatics 30, 2811–2812 (2014).
Acknowledgements
This paper is published with the permission of the director of the Kenya Medical Research Institute (KEMRI). The authors are grateful to the study participants and the parasite culture laboratory at the KEMRI–Wellcome Trust Research Programme, Kilifi, Kenya. We also thank G. McFadden, M. Greischar and A. Read for helpful comments and H. Ginsburg for assistance with the gene sets for the enrichment tests. This work was supported by the Wellcome Trust (grant numbers 088634 to M.J.M. and 092741 and 077176 to K.M.).
Author information
Authors and Affiliations
Contributions
M.K.R., M.A.N., J.J.S., J.M.N., M.M.E., A.S.A., M.M.K. and M.J.M. collected the data. Z.B. and S.M. provided the microarray materials. I.M.E., J.N.W., K.M. and M.J.M. organized the field work. M.J.M. and M.K.R. prepared the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–9; Supplementary Tables 1, 3 and 5; Legends for Supplementary Figures 10–11 and Supplementary Tables 2, 4, 6, 7 and 8; Supplementary references.
Supplementary Table 2
Genes showing significant differences in expression level between high (H) and low (L) transmission populations.
Supplementary Table 4
Summary of methods for adjusting for host, parasite and gene properties in the analyses.
Supplementary Table 6
Summary of network module properties.
Supplementary Table 7
Gene sets used for functional.
Supplementary Table 8
Abbreviated names for epigenetic and translational machinery genes used in the analyses.
Supplementary Figure 10
H–L differentiation among transport genes.
Supplementary Figure 11
H–L differentiation among trafficking genes.
Rights and permissions
About this article
Cite this article
Rono, M.K., Nyonda, M.A., Simam, J.J. et al. Adaptation of Plasmodium falciparum to its transmission environment. Nat Ecol Evol 2, 377–387 (2018). https://doi.org/10.1038/s41559-017-0419-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-017-0419-9
This article is cited by
-
Epigenetic regulation as a therapeutic target in the malaria parasite Plasmodium falciparum
Malaria Journal (2024)
-
MalariaSED: a deep learning framework to decipher the regulatory contributions of noncoding variants in malaria parasites
Genome Biology (2023)
-
Diagnostic performance of NxTek™ Eliminate Malaria-Pf test for the detection of Plasmodium falciparum in school children with asymptomatic malaria
Malaria Journal (2023)
-
Schoolchildren with asymptomatic malaria are potential hotspot for malaria reservoir in Ethiopia: implications for malaria control and elimination efforts
Malaria Journal (2023)
-
Plasmodium falciparum gametocyte carriage in longitudinally monitored incident infections is associated with duration of infection and human host factors
Scientific Reports (2023)