Abstract
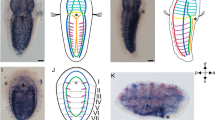
In amniote embryos, skeletal muscles in the trunk are derived from epithelial dermomyotomes, the ventral margin of which extends ventrally to form body wall muscles. At limb levels, ventral dermomyotomes also generate limb-muscle precursors, an Lbx1-positive cell population that originates from the dermomyotome and migrates distally into the limb bud. In elasmobranchs, however, muscles in the paired fins were believed to be formed by direct somitic extension, a developmental pattern used by the amniote body wall muscles. Here we re-examined the development of pectoral fin muscles in catsharks, Scyliorhinus, and found that chondrichthyan fin muscles are indeed formed from Lbx-positive muscle precursors. Furthermore, these precursors originate from the ventral edge of the dermomyotome, the rest of which extends towards the ventral midline to form body wall muscles. Therefore, the Lbx1-positive, de-epithelialized appendicular muscle precursors appear to have been established in the body plan before the divergence of Chondrichthyes and Osteichthyes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
17 October 2017
In Fig. 2 of this Article originally published, some erroneous lines appeared on the left side of the images in panels c, e and g. The figure should have appeared as shown below. These errors have now been corrected in all versions of the Article.
References
Jagla, K. et al. Mouse Lbx1 and human LBX1 define a novel mammalian homeobox gene family related to the Drosophila lady bird genes. Mech. Dev. 53, 345–356 (1995).
Schafer, K. & Braun, T. Early specification of limb muscle precursor cells by the homeobox gene Lbx1h. Nat. Genet. 23, 213–216 (1999).
Brohmann, H., Jagla, K. & Birchmeier, C. The role of Lbx1 in migration of muscle precursor cells. Development 127, 437–445 (2000).
Gross, M. K. et al. Lbx1 is required for muscle precursor migration along a lateral pathway into the limb. Development 127, 413–424 (2000).
Cinnamon, Y., Kahane, N. & Kalcheim, C. Characterization of the early development of specific hypaxial muscles from the ventrolateral myotome. Development 126, 4305–4315 (1999).
Neyt, C. et al. Evolutionary origins of vertebrate appendicular muscle. Nature 408, 82–86 (2000).
Ochi, H. & Westerfield, M. Lbx2 regulates formation of myofibrils. BMC Dev. Biol. 9, 13 (2009).
Goodrich, E. Studies on the Structure and Development of Vertebrates (Macmillan, London, 1930).
Braus, H. Beitrage zur entwicklung der musculatur unddes peripheren nervensystems der selachier. Morphologisches Jahrbuch 27, 501–629 (1899).
Dohrn, A. Die paarigen und unpaaren Flossen der Selachier. Mittheilungen aus der Zoologischen Station zu Neapel 5, 161–195 (1884).
Ballard, W. W., Mellinger, J. & Lechenault, H. A series of normal stages for development of Scyliornius canicula, the lesser spotted dogfish (Chondrichthyes: Scyliohnidae). J. Exp. Zool. 267, 318–336 (1993).
Mennerich, D., Schafer, K. & Braun, T. Pax-3 is necessary but not sufficient for lbx1 expression in myogenic precursor cells of the limb. Mech. Dev. 73, 147–158 (1998).
Kusakabe, R., Kuraku, S. & Kuratani, S. Expression and interaction of muscle-related genes in the lamprey imply the evolutionary scenario for vertebrate skeletal muscle, in association with the acquisition of the neck and fins. Dev. Biol. 350, 217–227 (2011).
Kusakabe, R. & Kuratani, S. Evolution and developmental patterning of the vertebrate skeletal muscles: perspectives from the lamprey. Dev. Dynam. 234, 824–834 (2005).
Minchin, J. E. et al. Oesophageal and sternohyal muscle fibres are novel Pax3-dependent migratory somite derivatives essential for ingestion. Development 140, 2972–2984 (2013).
Chen, F., Liu, K. C. & Epstein, J. A. Lbx2, a novel murine homeobox gene related to the Drosophila ladybird genes is expressed in the developing urogenital system, eye and brain. Mech. Dev. 84, 181–184 (1999).
Lou, Q., He, J., Hu, L. & Yin, Z. Role of lbx2 in the noncanonical Wnt signaling pathway for convergence and extension movements and hypaxial myogenesis in zebrafish. Biochim. Biophys. Acta 1823, 1024–1032 (2012).
Jacob, M., Christ, B. & Jacob, H. J. The migration of myogenic cells from the somites into the leg region of avian embryos. An ultrastructural study. Anat. Embryol. 157, 291–309 (1979).
Brand-Saberi, B., Muller, T. S., Wilting, J., Christ, B. & Birchmeier, C. Scatter factor/hepatocyte growth factor (SF/HGF) induces emigration of myogenic cells at interlimb level in vivo. Dev. Biol. 179, 303–308 (1996).
Bladt, F., Riethmacher, D., Isenmann, S., Aguzzi, A. & Birchmeier, C. Essential role for the c-met receptor in the migration of myogenic precursor cells into the limb bud. Nature 376, 768–771 (1995).
Dietrich, S. et al. The role of SF/HGF and c-Met in the development of skeletal muscle. Development 126, 1621–1629 (1999).
Mackenzie, S., Walsh, F. S. & Graham, A. Migration of hypoglossal myoblast precursors. Dev. Dynam. 213, 349–358 (1998).
Lours-Calet, C. et al. Evolutionarily conserved morphogenetic movements at the vertebrate head–trunk interface coordinate the transport and assembly of hypopharyngeal structures. Dev. Biol. 390, 231–246 (2014).
Takagi, W. et al. Hepatic and extrahepatic distribution of ornithine urea cycle enzymes in holocephalan elephant fish (Callorhinchus milii). Comp. Biochem. Physiol. B 161, 331–340 (2012).
Didier, D. A., Leclair, E. E. & Vanbuskirk, D. R. Embryonic staging and external features of development of the chimaeroid fish, Callorhinchus milii (Holocephali, Callorhinchidae). J. Morphol. 236, 25–47 (1998).
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Tamura, K. et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011).
Sharpe, J. et al. Optical projection tomography as a tool for 3D microscopy and gene expression studies. Science 296, 541–545 (2002).
Kawamoto, T. Use of a new adhesive film for the preparation of multi-purpose fresh-frozen sections from hard tissues, whole-animals, insects and plants. Arch. Histol. Cytol. 66, 123–143 (2003).
Oisi, H., Ota, K. G., Kuraku, S. & Kuratani, S. Craniofacial development of hagfishes and the evolution of vertebrates. Nature 10, 175–180 (2013).
Kaneko, H., Nakatani, Y., Fujimura, K. & Tanaka, M. Development of the lateral plate mesoderm in medaka Oryzias latipes and Nile tilapia Oreochromis niloticus: |insight into the diversification of pelvic fin position. J. Anat. 225, 659–674 (2014).
Nakaya, Y., Sukowati, E. W., Wu, Y. & Sheng, G. RhoA and microtubule dynamics control cell-basement membrane interaction in EMT during gastrulation. Nat. Cell Biol. 10, 765–774 (2008).
Acknowledgements
We thank A. Tweedale and staff of the Station Biologique de Roscoff for collecting S. canicula embryos, J. D. Bell for collecting C. milii embryos, Y. Yamamoto and K. Ikeda for electron microscopy studies and Y. Oisi and S. Higuchi for technical support. This work was supported in part by a Grant-in-Aid for Scientific Research (B) (25291086) (16H04828) and the Inamori Foundation to M.T., a Grant-in-Aid for Scientific Research (C) (16K07384) to R.K., the Japan–Australia Research Cooperative Program to S.H., the Spanish Ministry of Economy and Competitiveness, ‘Centro de Excelencia Severo Ochoa 2013–2017’, SEV-2012-0208 to J.S., and a Grant-in-Aid for Scientific Research (A) (15H02416) to S.Kurat.
Author information
Authors and Affiliations
Contributions
M.T., E.O., R.K. and S.Kurat. designed the project and wrote the manuscript, J.S. supervised OPT analyses, and E.O. performed most experiments, except the following experiments. S.Kurak. assisted with sequence identification and molecular phylogenetic analyses, S.H. collected C. milii embryos and provided related materials, A.R.-M. performed OPT analyses, and K.O. assisted with cloning and phylogenetic analyses.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplemental Figures
Supplementary Figures 1-5.
Supplementary Video 1
3D-reconstructed S. canicula embryos at stage 27.
Supplementary Video 2
3D-reconstructed S. canicula embryos at stage 28.
Supplementary Video 3
OPT-scanned S. canicula embryos at stage 26.
Supplementary Video 4
OPT-scanned S. canicula embryos at early stage 27.
Supplementary Video 5
OPT-scanned S. canicula embryos at early stage 28.
Rights and permissions
About this article
Cite this article
Okamoto, E., Kusakabe, R., Kuraku, S. et al. Migratory appendicular muscles precursor cells in the common ancestor to all vertebrates. Nat Ecol Evol 1, 1731–1736 (2017). https://doi.org/10.1038/s41559-017-0330-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-017-0330-4
This article is cited by
-
Novel developmental bases for the evolution of hypobranchial muscles in vertebrates
BMC Biology (2020)
-
Stepwise participation of HGF/MET signaling in the development of migratory muscle precursors during vertebrate evolution
Zoological Letters (2018)
-
Evolution of the muscular system in tetrapod limbs
Zoological Letters (2018)
-
Developmental mechanisms of migratory muscle precursors in medaka pectoral fin formation
Development Genes and Evolution (2018)