Abstract
The tobacco cutworm, Spodoptera litura, is among the most widespread and destructive agricultural pests, feeding on over 100 crops throughout tropical and subtropical Asia. By genome sequencing, physical mapping and transcriptome analysis, we found that the gene families encoding receptors for bitter or toxic substances and detoxification enzymes, such as cytochrome P450, carboxylesterase and glutathione-S-transferase, were massively expanded in this polyphagous species, enabling its extraordinary ability to detect and detoxify many plant secondary compounds. Larval exposure to insecticidal toxins induced expression of detoxification genes, and knockdown of representative genes using short interfering RNA (siRNA) reduced larval survival, consistent with their contribution to the insect’s natural pesticide tolerance. A population genetics study indicated that this species expanded throughout southeast Asia by migrating along a South India–South China–Japan axis, adapting to wide-ranging ecological conditions with diverse host plants and insecticides, surviving and adapting with the aid of its expanded detoxification systems. The findings of this study will enable the development of new pest management strategies for the control of major agricultural pests such as S. litura.
Similar content being viewed by others
Introduction
The tobacco cutworm, Spodoptera litura (Lepidoptera, Noctuidae), is an important polyphagous pest; its larvae feed on over 100 crops1. This pest is widely distributed throughout tropical and subtropical areas of Asia including India, China and Japan. In India particularly, S. litura causes heavy yield loss varying between 10 and 30%1. High fecundity and a short life cycle under tropical conditions result in a high rate of population increase and subsequent population outbreaks. In addition, it has evolved high resistance to every class of pesticide used against it2,3, including the biopesticide Bt4. Few complete genome sequences have been reported for noctuids, which include many serious agricultural pests. Asian researchers launched the S. litura genome project as an international collaboration in cooperation with the Fall armyworm International Public Consortium (FAW-IPC), for which a genome project is coordinately underway5. By comparative genomic studies with the monophagous species Bombyx mori and other Spodoptera species such as S. frugiperda (which has a different geographical distribution), S. litura genome information can provide new insights into mechanisms of evolution, host plant specialization and ecological adaptation, which can serve as a reference for noctuids and lead to selective targets for innovative pest control.
Results and discussion
Genome structure and linkage map of S. litura.
We sequenced and assembled a genome for S. litura comprising 438.32 Mb, which contains 15,317 predicted protein-coding genes analysed by GLEAN6 and 31.8% repetitive elements (Supplementary Tables 1–4). Among four representative lepidopteran species with complete genome sequences7,8,9, S. litura harbours the smallest number of species-specific gene families (Supplementary Fig. 1a and Supplementary Table 9). A phylogenetic tree constructed by single-copy orthologous groups showed that S. litura separated from B. mori and Danaus plexippus about 104.7 Myr ago (Ma), and diverged approximately 147 Ma from the more basal Plutella xylostella, whereas Lepidoptera as a whole separated from Diptera about 258 Ma, consistent with reported divergence time estimates10 (Supplementary Fig. 1b). To construct a linkage map, a heterozygous male F1 backcross (BC1) population was established between Japanese and Indian inbred strains. The resulting genetic analysis used 6088 RAD-tags as markers to anchor 639 scaffolds covering 380.89 Mb onto 31 chromosomes, which corresponded to 87% of the genome (Supplementary Section 2). Genomic syntenies from S. litura to B. mori and to Heliconius melpomene revealed two modes of chromosomal fusion (Supplementary Tables 10 and 11 and Supplementary Fig. 2). In one, six S. litura chromosomes (haploid chromosome number N = 31) were fused to form three B. mori chromosomes (N = 28). In the other, six sets of S. litura chromosomes were fused, corresponding to six H. melpomene chromosomes (N = 21)11, and another eight S. litura chromosomes were fused, corresponding to four other H. melpomene chromosomes. These changes were consistent with previous reports on chromosome evolution among butterflies including Melitaea cinxia 12 and the moth Manduca sexta 13 (Supplementary Section 2).
Massive expansion of bitter gustatory receptor and detoxification-related gene families associated with polyphagy of Noctuidae
To elucidate key genome changes associated with host plant specialization and adaptation in Lepidoptera, we compared chemosensory and detoxification-related gene families between the extremely polyphagous lepidopteran pest S. litura and the almost monophagous lepidopteran model organism B. mori. We found large expansions of the gustatory receptor (GR), cytochrome P450 (P450), carboxylesterase (COE) and glutathione-S-transferase (GST) gene families in S. litura (Table 1). Chemosensory genes play an essential role in host plant recognition of herbivores. GRs, especially, are highly variable among species, which could be a major factor for host plant adaptation. GRs are categorized into three classes—CO2 receptors, sugar receptors and bitter receptors—among which bitter receptors are most variable, while CO2 and sugar receptors are conserved14,15,16,17,18. Manual annotation identified 237 GR genes in the S. litura genome (Table 2, Fig. 1a and Supplementary Table 13), whereas in the other lepidopteran species investigated to date, most of which are mono- and oligophagous, only about 45–80 GRs are reported8,11,14,16,19,20. Since large expansions of GR genes were also reported recently in S. frugiperda 5 and in another polyphagous noctuid, Helicoverpa armigera 21, the expansion of GRs may be a unique adaptation mechanism for polyphagous Noctuidae to feed on a wide variety of host plants (Table 2). Phylogenetic analysis including GR genes of B. mori, M. sexta, H. melpomene and S. frugiperda showed clearly that greatly expanded bitter GR clades were composed of SlituGRs and SfruGRs exclusively (Supplementary Fig. 3), supporting a strong association of a major expansion of bitter receptor genes with the appearance of polyphagy in the Noctuidae. GR expansions mainly occurred by duplications, as many structurally similar GR genes are located in clusters on the same scaffold/chromosome (for example, Chr 12, 14 and 25; Fig. 1a–c). Interestingly, while many H. armigera GR genes have been identified as intronless21, especially in the bitter GR clade, here we found that almost all S. litura GR genes possessed introns. This suggests that different mechanisms led to GR expansion in these two species.
a, Comparison of chemosensory and detoxification related gene families between the extremely polyphagous pest S. litura and almost monophagous B. mori. Black thick bars denote the largest bitter GR cluster on Chr 12. R represents receptor. b, There is a large expansion of bitter GR genes on S. litura Chr 14. Thirteen bitter GR genes clustered on S. litura Chr 14 were mainly expressed in moth proboscis and larval maxilla, whereas the corresponding BmorGR gene cluster on Chr 10 composed of BmorGR55-57 was expressed in larval chemoreception organs16. c, Expansion of S. litura single-exon bitter GR genes on Chr 25 mainly expressed in moth proboscis. The corresponding BmorGR53, which is also a single-exon gene, was expressed in larval maxilla. d, Heatmap of S. litura GR expression in various tissues by RNA-Seq. L.Ant., larval antenna; L.Epi., larval epipharynx; L.Leg, larval legs; L.Max., larval maxilla; L.Mid., larval midgut; M.Ant., moth antenna; M.Leg, moth legs; M.P.G., moth pheromone glands; M.Pro., moth proboscis. The vertical red two-way arrow indicates the largest bitter GR cluster on Chr 12, which was mainly expressed in larval maxilla. Thick blue bars represent GR gene clusters on Chr 14 and Chr 25, which were mainly expressed in moth proboscis. R denotes receptor.
Transcriptome and phylogenetic analyses of expanded bitter GR genes in S. litura
Transcriptome analysis revealed that at least 109 of the predicted bitter GR genes were expressed, mostly in larval palps and adult proboscis, but a large number were also expressed in other chemoreception organs such as antennae, legs and the pheromone gland (Fig. 1d). These observations are similar to GR expression patterns reported in adult tissues of H. melpomene 14 and in diverse developmental stages and tissues in H. armigera 21. Intriguingly, four bitter GR genes on Chr 25 and 14 bitter GR genes on Chr 14 were mainly expressed in moth proboscis (Fig. 1d), which S. litura uses to suck flower nectar to obtain energy for flying. Comparison with the silkmoth, which does not feed, showed that the expansion of these gene clusters could represent an adaptation to detect toxic plant secondary metabolites present in flower nectar (Fig. 1b,c). From our phylogenetic analysis (Supplementary Fig. 3), expansion of the biggest cluster of bitter GR genes on Chr 12 was Spodoptera-specific. These genes were mainly expressed in larval maxilla, consistent with the idea that a large expansion of bitter GR genes supports the polyphagy of Spodoptera and an ability to detect a large number of toxic metabolites in host plants (Fig. 1d). The mechanisms by which perception of bitter substances result in specific behaviours are complex, and those underlying bitter receptor function in Lepidoptera have not yet been elucidated.
Association of major expansions of SlituP450 genes with intensified detoxification
Detoxification of xenobiotics is crucial for ecological adaptation of highly polyphagous pest species to different host plants. This process usually involves several distinct detoxification pathways, from active metabolism of toxins22 to enhanced excretion activity by ABC transporters23,24. We annotated 138 P450 genes in the S. litura genome, among which P450 clans 3 and 4 showed large expansions (Fig. 2a, Supplementary Fig. 4 and Supplementary Table 14). CYP9a especially was greatly expanded on S. litura Chr 29 compared to the corresponding chromosome of B. mori (Fig. 2a, upper panel). Transcriptome analysis showed that some of the expanded S. litura CYP9a genes were inducible by treatment with xanthotoxin, imidacloprid or ricin (P450-100, 103 and 105; Fig. 2a, middle panel). CYP9a is reported to be inducible by xanthotoxin in S. litura 25 and S. exigua 26. Other P450 clan 3 expansions (CYP337a1 and a2, CYP6ae9 and CYP6b29, and CYP321b1) were also induced by the toxin treatments (Supplementary Fig. 5a), suggesting a link between P450 clan 3 expansions and an increase of tolerance to toxin in this pest. To test this hypothesis, we selected P450-74, 88, 92 and 98 as members of P450 clan 3 for knockdown experiments. We injected each siRNA of the corresponding P450 into fifth-instar larvae. After feeding with an artificial diet containing imidacloprid, we observed an increase in sensitivity to the insecticide in the treated larvae compared to controls (Supplementary Fig. 7a-d). Recently, the role of SlituCYP321b1 in insecticide resistance was confirmed by showing that it is overexpressed in the midgut after induction by several pesticides, and that RNAi-mediated silencing of SlituCYP321b1 significantly increased mortality of S. litura larvae exposed to the same pesticides27.
a, A comparison of the Cyp9a gene cluster on B. mori Chr 17 with S. litura Chr 29. Top: genomic organization. Cyp9a gene clusters contain four ADH genes in both species, while two GR genes are present only in S. litura. Cyp9a, red; ADH, yellow; GR, blue. Middle: expression heatmap of Cyp9a genes induced by toxin treatment in three tissues. Bottom: diversity of genes associated with the Cyp9a cluster domain including the ADH gene cluster among 16 local populations of S. litura (see also Fig. 4a). b, Expanded lepidopteran esterase gene cluster on S. litura Chr 2. Top: genomic organization in S. litura and B. mori. COE, red; ACE (acetylcholinesterase), green. Bottom: expression heatmap of COE induced by toxin treatment. Toxins: imidacloprid (Imid), ricin and xanthotoxin (Xan). Expression was measured in fat body (fb), midgut (mg) and Malpighian tubule (mp).
Major expansions of SlituGST genes enhance insecticide tolerance of this pest
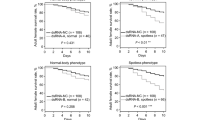
Expansions of SlituGST genes were derived from epsilon classes on Chr 9 and Chr 14; the expression of these genes was also induced by toxin treatment (Fig. 3a–c and Supplementary Table 16). We chose SlituGST07 and SlituGST20 as representatives of the expanded clusters on Chr 14 and Chr 9, respectively, for knockdown and imidacloprid pesticide binding assays. We injected the siRNAs into fifth-instar larvae, then fed them an artificial diet containing imidacloprid (50 µg g−1). This treatment resulted in lethality in siRNA-injected larvae, while controls remained alive (Fig. 3d,e), consistent with the idea that expansion of the GSTε class conferred an increase in detoxification ability. Figure 3f,g shows the inhibitory effects of imidacloprid on SlituGST07 and SlituGST20 in a competitive binding assay (Supplementary Section 6). These observations confirmed that expansion of GSTε contributes to the detoxification ability of this pest.
a, Phylogenetic tree of GSTs of S. litura (magenta) and B. mori (blue). Arrows show representative GSTε genes, SlituGST07 and GST20, for each GSTε cluster used for knockdown and binding assays. We excluded SlituGST45-49 from the phylogenetic tree, since these microsomal GSTs are very short compared with other GSTs and their amino acid sequences are fairly distant from other classes. b,c, Organization and expression of expanded S. litura GSTε genes on Chr 9 and Chr 14 with toxin treatment. Toxins and tissues are the same as Fig. 2a,b. d,e, Increased sensitivity to the pesticide imidacloprid caused by knockdown of GST07 and GST20. d, Knockdown of fifth-instar larvae, performed by siRNA injection and confirmed by RT-qPCR. At 24 hr after siRNA injection, larvae were fed an artificial diet containing imidacloprid (50 µg g−1). The percentage of larvae affected by imidacloprid (dead and almost dead; see Methods) is shown. Ten larvae were used per experiment in three independent replicates and the results are presented with the standard deviation (SD). e, Knockdown reduction rates of GST07 and GST20 (31% and 57%, respectively). Control larvae were injected with siGFP (see Methods). The relative expression is shown as mean + SD of three independent replicates of 10 larvae each, using a Student’s t-test, *P<0.05, **P<0.01. f,g, Binding assay of SlituGST07 (f) and SlituGST20 (g) with imidacloprid. The inhibitory effect of imidacloprid on SlituGST07 and SlituGST20 was determined using CDNB and GSH as substrates (see Methods). Enzymatic activity of SlituGST07 and SlituGST20 was measured in the presence of various concentrations of imidacloprid. The value from the assay with 1 × 10−4 mM of imidacloprid was set to 100%. Error bars denote SEM from 3 independent experiments (10 larvae per treatment).
Associating large expansions of SlituCOE genes with intensified detoxification
COE genes, which play an important role in the metabolism of a wide range of xenobiotics associated with plants and insecticides22,28,29,30, also showed large expansions of lepidopteran and α classes (Table 1, Supplementary Fig. 6a and Supplementary Table 15). RNA-Seq analysis showed that the expanded COE genes were inducible with toxin treatment, suggesting again that their expansion is linked to an increase in detoxification ability (Fig. 2b, lower panel). These results supported knockdown experiments for COE-57 and COE-58 whereby injected larvae fed with an artificial diet containing imidacloprid showed a 60–80% increase in sensitivity compared to controls (Supplementary Fig. 7e,f). Taken together with our knockdown experiments, transcript induction by imidacloprid indicates that expansion of the P450, GST and COE families is linked to tolerance of this insecticide.
Roles of non-expanded detoxification gene families
Although the APN and ABC gene families did not exhibit significant expansion, they were highly induced by ricin treatment (Supplementary Figs. 8 and 9 and Supplementary Tables 17 and 18). APN 31, ABCC2 32 and ABCA2 33 have been shown to function as Cry protein receptors32,33 (see Supplementary Sections 7 and 8). Thus, APN and ABC transport proteins may be involved in the response to different classes of xenobiotics. Altogether our results suggest that S. litura probably achieves its impressive polyphagy by adopting a strategy of large expansions of diverse sensory and detoxification-related genes, with probable cross-talk in their regulation, to adapt to a great variety of host plants.
Genetic population structure reveals extensive long-distance migration of this pest
We analysed the genetic diversity and gene flow of S. litura sampled from 3 locations in India, 11 locations in China and 2 locations in Japan (Supplementary Table 21). This yielded a clear geographical map of the genetic diversity of the surveyed local populations and genetic population structure in these countries. We observed extremely high genetic similarity between Hyderabad (central India), Fujian (the southeast coast of mainland China) and Okinawa/Tsukuba (Japan) (F ST < 0.01, Fig. 4a and Supplementary Table 23). The model-based structure analysis34 provided a predicted population structure consistent with an F ST-based cluster analysis (Fig. 4b and Supplementary Fig. 10a,b). By incorporating the estimated allele frequency divergence between the ancestral populations, we obtained a very stable picture of population structure relative to the assumed number of ancestral populations (K). Here, again, we observed extremely high genetic similarity between central India (Hyderabad and Matsyapuri), the southeast coast of mainland China (Zhejiang, Guangzhou and Fujian) and Japan (Okinawa and Tsukuba). The assignment of individual genomes to the ancestral populations provided a detailed picture of the gene flow (Fig. 4b). These results are consistent with the study of DNA sequence variation among populations of S. litura in China and Korea35. An additional factor affecting population dispersal is oversea migration from southern China to western Japan driven by typhoons36,37. Geographical data on the Asian monsoon in July–August38 may support our results, enabling S. litura to undertake a trip of even longer distance from southern India to China and Japan.
a, F ST-based cluster analysis of local populations. Structure results (b and Supplementary Fig. 10) suggest that one of the individuals from the Hunan2 sample belongs to a migration population (Hunan2-4), while the other 3 individuals belong to a local population (Hunan2-1, Hunan2-2 and Hunan2-3). Here, we treated the Hunan2 samples as a mixed population with both migration and local populations. b, Assignment of the individual genomes in the samples to the ancestral populations predicted by structure. We obtained the predicted allele frequency divergences between individual genomes by the predicted allele frequency divergence between the ancestral populations and the membership coefficients of the individual genomes (see Methods). c, Two-dimensional allele frequency spectra in the paired population groups. d, Global picture of the migration route predicted by ∂a∂i. The inset shows the number of migrating chromosomes per generation. The four closed ropes represent the migrating population in India, local populations in China, migrating populations in China, and the populations in Japan. The size of the circles represents the genetic diversity (π).
To understand the global pattern of migration routes, we analysed the joint allele frequency spectrums (Fig. 4c) by ∂a∂i (diffusion approximation for demographic inference)39. ∂a∂i fits the solution of the Fokker–Planck–Kolmogorov equation to the data of the joint allele frequency spectrum, and the estimated values of the coefficients provide direct information on the population histories and migration rates. Based on the F ST-based population structure and the model-based assignment of the individual genomes, we constructed six population groups: two groups in India (India_local and India_migrate), three in China (China_isolate, China_local, and China_migrate), and one in Japan. By applying the isolation with migration model40 to each of the pairs of population groups, we identified a global route from the Indian migrating population through the Chinese local population, which ranges from the south at Hainan to the north at Hubei (Fig. 4d). This Chinese local population has a large number of migrants to and from the Chinese migrating population. We observed moderate numbers of migrants from China to Japan and from China to India. ∂a∂i also implied that the local populations in India and China have been shrinking significantly for the past 2000–3000 years. In contrast, the Japanese population has been expanding for the past 5000 years (Supplementary Figs. 11 and 12 and Supplementary Table 24). It would be of interest to investigate the extent to which these local populations are also pests and have insecticide resistance.
Conclusion
This study provides strong evidence on how this polyphagous insect has evolved to become a deleterious and powerful global pest through adaptative changes and subsequent selection of gene expansions. It also provides an explanation for the genetic basis for its high tolerance to pesticides, which involves mechanisms similar to plant allelochemical detoxification. The population genetic analysis revealed the extensive migratory ability of S. litura. Such a deeper understanding through genomics and transcriptomics will enable us to develop novel pest management strategies for the control of major agricultural pests like S. litura and its near relatives, and to design new classes of insecticide molecules.
Methods
Genome sequencing and assembly
An inbred strain of S. litura (the Ishihara strain) was developed by successive single-pair sib matings for 24 generations and reared on an artificial diet at 25 °C. Male moths were used to extract genomic DNA for sequencing. Shotgun libraries with insert sizes of 170, 300, 500 and 800 bp (short insert sizes) and 2, 5 and 10 kb (large insert sizes) were constructed by following the manufacturer’s protocol (http://www.illumina.com). After quality control of DNA libraries, ssDNA fragments were hybridized and amplified to form clusters on flow cells. Paired-end sequencing was performed following the standard Illumina protocol.
The S. litura genome was assembled using the software program ALLPATHS-LG build 4775841. The assembly used default parameters with the exception of using a ploidy setting of 2 (PLOIDY = 2), as recommended for a diploid organism, in the data preparation stage, and a minimum contig size set to 200 bp (MIN_CONTIG = 200) in the running stage (running the RunAllPathsLG command). Gaps within the scaffolds were filled based on the short insert size libraries, using the GapCloser in the SOAPdenovo package42. Assembled scaffolds were assigned to chromosomes by the order and orientation of a linkage map combined with a synteny analysis between S. litura and B. mori. The sequencing depth and GC content distribution of the assembled genome sequence were evaluated by mapping the short insert size reads back to the scaffolds using SOAP243.
Genome annotation
Three methods were used for S. litura gene prediction including ab initio, homology-based and transcript-based methods; the GLEAN program6 was used to derive consensus gene predictions. For ab initio prediction, AUGUSTUS44 and SNAP45 were used to predict protein-coding genes. For homology-based prediction, proteins from five insect genomes (Anopheles gambiae, Drosophila melanogaster, B. mori, Acyrthosiphon pisum and D. plexippus) were first mapped to the S. litura genome using TBLASTN (E-value ≤ 0.00001), and then accurate splicing patterns were built with GeneWise (version 2.0)46. In the transcript-based method, the assembled transcriptome results were mapped onto the genome by BLAT with identity ≥99% and coverage ≥95%. We used TopHat to identify exon–intron splice junctions and refine the alignment of the RNA-Seq reads to the genome47, and Cufflinks (version 1.2.0 release) to define a final set of predicted genes48. Finally, we integrated the three kinds of gene predictions to produce a comprehensive and non-redundant reference gene set using GLEAN. Gene function information was assigned based on the best hits derived from the alignments to proteins annotated in the SwissProt, TrEMBL49 and KEGG50 databases using BLASTP51. Motifs and domains of proteins were annotated using InterPro52 by searching public databases, including Pfam, PRINTS, PROSITE, ProDom and SMART. We also described gene functions using Gene Ontology (GO)53.
Repeats and transposable element families in the S. litura genome were first detected by the RepeatModeler (version open-1.0.7) pipeline, with rmblast-2.2.28 as a search engine. With the assistance of RECON54 and RepeatScout55, the pipeline employs complementary computational methods to build and classify consensus models of putative repeats. tRNAs were annotated by tRNAscan-SE with default parameters. rRNAs were annotated by RNAmmer prediction and homology-based search of published rRNA sequences in insects (deposited in the Rfam database). snRNAs and miRNAs were sought using a two-step method: after aligning with BLAST, INFERNAL was used to search for putative sequences in the Rfam database (release 9.1).
Gene family clustering and phylogenetic tree construction
Protein sequences longer than 30 amino acids were collected from nine sequenced arthropod species (B. mori, P. xylostella, D. plexippus, D. melanogaster, A. darlingi, Apis mellifera, Harpegnathos saltator, Tribolium castaneum and Tetranychus urticae) and S. litura for gene family clustering using Treefam56. We aligned all-to-all using BLASTP with an E-value cut-off of 0.0000001, and assigned a connection (edge) between two nodes (genes) if more than a third of a region was aligned in both genes. An H-score ranging from 0 to 100 was used to weigh the similarity (edge). For two genes, G1 and G2, the H-score was defined as score (G1G2)/max (score(G1G1),score(G2G2)), where ‘score’ is the raw BLAST score. The average distance was used for the hierarchical clustering algorithm, requiring the minimum edge weight (H-score) to be larger than 10 and the minimum edge density (total number of edges/theoretical number of edges) to be larger than 1/3.
386 single-copy genes from the 10 species were aligned by MUSCLE57. We used MODELTEST58 to select the best substitution model (GTR) and MRBAYES59 to construct the phylogenetic tree. Then we estimated divergence time and neutral substitution rate per year (branch/divergence time) among species. The PAML mcmctree60 used to estimate the species divergence time referred to two fossil calibrations, including the divergence time of D. melanogaster and Culicidae (238.5–295.4 million years ago) and the divergence time of D. melanogaster and Hymenoptera (238.5–307.2 million years ago)61,62. T. urticae (Arachnida) was used as an outgroup, and a bootstrap value was set as 1000. In addition, the evolutionary changes in the protein family size (expansion or contraction) were analysed using the CAFÉ program63, which assesses the protein family expansion or contraction based on the topology of the phylogenetic tree.
Linkage map
Two genetically contrasting strains of S. litura, one developed at the University of Delhi, India (called the India strain) and another available at the National Institute of Agrobiological Sciences, Japan (the Ishihara strain), were employed to generate a mapping population. F1 offspring were obtained by crossing an India male and an Ishihara female. An F1 male was crossed with an Ishihara female as back cross (BC1), and these BC1 offspring were used to develop a RAD library64. Genomic DNA was isolated from 116 BC1 individuals, Ishihara male, India female and F1 male, and RAD sequencing libraries were constructed following a standard protocol. Sequencing was carried out using an Illumina HiSeq2000 platform. RAD-seq reads were aligned to the reference genome sequence using Short Oligonucleotide Analysis Package 2 (SOAP2)43 to analyse the genotypes of each individual at every genomic site. Polymorphic loci relative to the reference sequence were selected and then filtered. SNP markers were recorded if they were supported by at least 5 reads with quality value greater than 20, and ambiguous SNPs (SNP = N) were eliminated. Only SNP markers that were homozygous and polymorphic between parents, heterozygous in the F1 and followed a Mendelian segregation pattern were selected for linkage map construction. This resulted in the identification of a total of 87,120 RAD markers. Further filtering was done by selecting only SNP markers with a missing rate of <0.09 that were separated by at least 2000 bp. After such stringent filtering, a total of 6088 SNP markers were obtained and subsequently used to develop a linkage map using JoinMap 4.165. The limit of detection (LOD) score = Z = log(probability of sequence with linkage/probability of sequence with no linkage) for the occurrence of linkage was set to 4–20 (start–end). By applying the indicated parameters, we narrowed down the map to 31 linkage groups (Supplementary Fig. 2b).
Syntenic comparison
We obtained peptides and genome sequences for B. mori 66, Papilio xuthus 67 and H. melpomene 11. If a gene had more than one transcript, only the first transcript in the annotation was used. To search for homology, protein-coding genes of S. litura were compared to those of B. mori, P. xuthus and H. melpomene using BLASTP51. For a protein sequence, the best five non-self hits in each target genome that met an E-value threshold of 0.00001 were reported. Whole-genome BLASTP results and the genome annotation file were used to compute collinear blocks for all possible pairs of chromosomes using MCScan software68. A region with at least 5 syntenic genes and no more than 15 gapped genes was called a syntenic block.
Annotation of the gustatory receptor (GR) gene family
A set of described Lepidoptera gustatory receptors (GRs) was used to search the S. litura genome by TBLASTN. Additionally, a combination approach of HMMER69 and Genewise46 was used to identify additional GR sequences. Scaffolds that were found to contain candidate GR genes were aligned to protein sequences to define intron/exon boundaries using Scipio70 and Exonerate71. The GR classification and the integrity of the deduced proteins were verified using BLASTP against the non-redundant GenBank database. When genes were split in different scaffolds, the protein sequences were merged for further analyses.
Annotation and phylogenetic study of the cytochrome P450 (CYP) gene family
Identity between two CYP proteins can be as low as 25% but the conserved motifs distributed along the sequence allow clear identification of CYP sequences. Conserved CYP protein structure is featured by a four-helix bundle (D, E, I and L), helices J and K, two sets of β sheets and a coil called the ‘meander’. The conserved motifs include WXXXR in the C helix, the conserved Thr of helix I, EXXR of helix K and the PERF motif followed by a haeme-binding region FXXGXXXCXG around the axial Cys ligand72. All the scaffolds containing candidate CYPs were manually annotated to identify intron/exon boundaries. Protein CYP sequences were compared by phylogenetic studies to the S. frugiperda CYPome73 for name attribution.
Annotation of carboxylesterase (COE), glutathione-S-transferase (GST), aminopeptidase N (APN) and ATP-binding cassette (ABC) transporter gene families
Sets of lepidopteran amino acid sequences for each gene family were collected from KAIKObase (http://sgp.dna.affrc.go.jp/KAIKObase/) and the NCBI Reference Sequence database. Each gene family was then searched in the S. litura genome assembly and predicted gene set by TBLASTN and BLASTP using each set of lepidopteran amino acid sequences. Identified genes were further examined by HMMER3 search (cutoff E-value = 0.001) using the Pfam database to confirm conserved domains in each gene family. In addition, the classification of each gene family was performed with BLASTP in the non-redundant GenBank database.
Construction of a phylogenetic tree of CYP, COE, GST, APN and ABC transporter gene families
Amino acid sequences of each lepidopteran gene family were automatically aligned by Mafft program version 7 (http://mafft.cbrc.jp/alignment/software/algorithms/algorithms.html), using an E-INS-i strategy74. When the alignment showed highly conservative and non-conservative regions, only the conservative regions were retained for further analysis. Model selection was conducted by MEGA version 675 and the LG+Gamma+I mode76,77,78. The maximum likelihood tree was inferred by RaxML version 879 using the LG+Gamma+I model. To evaluate the confidence of the tree topology, the bootstrap method80 was applied with 1000 replications using the rapid bootstrap algorithm81.
Illumina sequencing (RNA-Seq analysis)
Total RNA (1 μg) was used to make cDNA libraries using a TruSeq RNA sample preparation kit (Illumina, San Diego, CA). A total of 78 individual cDNA libraries were prepared by ligating sequencing adaptors to cDNA fragments synthesized using random hexamer primers. Raw sequencing data were generated using an Illumina HiSeq4000 system (Illumina, USA). The average length of the sequenced fragments was 260 bp. Raw reads were filtered by removal of adaptors and low-quality sequences before mapping. Reads containing sequencing adaptors, more than 5% unknown nucleotides or more than 50% bases of quality value less than 10, were eliminated. This output was termed ‘clean reads’. For analysis of gene expression, clean reads of each sample were mapped to S. litura gene sets using Bowtie2 (version 2.2.5), and then RSEM (v1.2.12) was used to count the number of mapped reads and estimate the FPKM (fragments per kilobase per million mapped fragments) value of each gene. Significant differential expression of genes was determined using the criteria that the false discovery rate was <0.01 and the ratio of intensity against control was >2 for induction or <0.5 for reduction.
Toxin treatment of S. litura larvae for transcriptome analysis
Fifth-instar larvae of the inbred strain were each fed with 1 g of artificial diet supplemented with 1 mg g−1 xanthotoxin. Control larvae were fed an artificial diet without xanthotoxin. For the ricin and imidacloprid treatments, the artificial diet was supplemented with either ground Ricinus communis seeds at a concentration of 50 mg g−1 or imidacloprid at a concentration of 50 µg g−1, respectively. Ten individuals were used for each treatment and three independent replicates were performed. Whole larvae were used for RNA extraction at 48 h post toxin treatment. Fat body, midgut and malpighian tubule were dissected from the toxin-treated larvae for RNA preparation. Total RNA was extracted from the tissues using Trizol reagent according to the manufacturer’s instructions (Invitrogen, USA) and contaminating DNA was digested with RNase-free DNase I (Takara, China). The integrity and quality of the mRNA samples were confirmed using an Agilent Bioanalyzer 2100 (Agilent Technologies, Santa Clara, CA).
GR transcriptome analysis
Larval antenna, thoracic legs, ephipharynx, maxilla and midgut were dissected from sixth-instar larvae, while antenna, legs, pheromone glands and proboscis were from moths. Due to very low GR expression levels, we used 100 larvae for RNA preparation. For expression profiling, we recorded all GR genes with expression levels higher than 0.1 FPKM in any tissue (Fig. 1d; red).
Quantitative PCR with reverse transcription (RT-qPCR)
Total RNA was subjected to reverse transcription using a PrimeScript™ RT Master Mix (Perfect Real Time) (TaKaRa) in 50 μl reaction volumes (2500 ng total RNA) and then diluted 5-fold. 1 μl cDNA was used per 10 μl PCR reaction volume. PCR was carried out with the following program: 94 °C for 2 min followed by 30 cycles of 94 °C for 10 sec, 50 °C for 15 sec, and 72 °C for 30 sec with rTaq DNA polymerase (TaKaRa) using pairs of gene-specific primers (Supplementary Table 19). RT-qPCR of each gene was repeated at least three times in two independent samples. BmActin3 was used as a control for each set of RT-qPCR reactions and for gel loading.
siRNA injection for knockdown of SlituGST, SlituP450 and SlituCOE genes
4 µl of siRNA (100 pm µl−1) were injected into the haemolymph of each fifth-instar larva, while injection of the same amount (4 µl) of GFP siRNA was used for controls. After 24 h post injection, larvae were reared on an artificial diet supplemented with imidacloprid at 50 µg g−1 until bioassay. siRNA sequences are listed in Supplementary Table 20.
To determine the effect of imidacloprid ingestion, larval condition was scored at 2, 6, 12, 18, 24, 36 and 48 post feeding. ‘Affected’ means that larvae rounded up and did not move after a couple of hours when touched, as if dead (suspended animation). However, several hours later, many affected larvae recovered from their suspended state, probably due to detoxification of ingested imidacloprid. The GST knockdown experiment used 3 replicates of 10 larvae. Post feeding replicates were scored independently for SlituGST-7 and -20; the remaining knockdowns (SlituP450-0740, -088, -092 and -098, and SlituCOE-057 and -058) were conducted as preliminary trials without replicates using 30 larvae per gene.
Overexpression and purification of recombinant SlituGST07 and SlituGST20 proteins
Competent Escherichia coli Rosetta (DE3) pLysS cells (Novagen; EMD Millipore) were transformed with expression vectors harbouring SlituGST07 cDNA (pET32.M3) or SlituGST20 cDNA (pCold_SUMO) and grown at 37 °C on Luria-Bertani (LB) medium containing 100 µg ml−1 ampicillin. After cells transformed with SlituGST07 cDNA reached a density of 0.7 OD600, isopropyl 1-thio-ß-D-galactoside (IPTG) was added to a final concentration of 1 mM to induce the production of recombinant protein and cultured overnight at 30 °C. Cells were then harvested by centrifugation, homogenized in 20 mM Tris-HCl buffer (pH 8.0) containing 0.5 M NaCl, 4 mg ml−1 of lysozyme, and disrupted by sonication. Cells transformed with SlituGST20 cDNA were grown to a density of 0.5 OD600, and stored on ice for 30 min before addition of IPTG to a final concentration of 1 mM, followed by a further incubation overnight at 18 °C before harvesting and disruption. Unless otherwise noted, all of the operations described below were conducted at 4 °C. The supernatant was clarified by centrifugation at 10,000g for 15 min and subjected to Ni2+-affinity chromatography equilibrated with 20 mM Tris-HCl buffer (pH 8.0) containing 0.2 M NaCl. After washing with the same buffer, the samples were eluted with a linear gradient of 0–0.5 M imidazole. The enzyme-containing fractions, assayed as described below, were pooled, concentrated using a centrifugal filter (Millipore, Billerica, MA, USA), and applied to a Superdex 200 column (GE Healthcare Bio-Sciences, Buckinghamshire, UK) equilibrated with the same buffer plus 0.2 M NaCl. Each fraction was assayed and analysed by SDS-PAGE using a 15% polyacrylamide slab gel containing 0.1% SDS, according to the method of Laemmli82. Protein bands were visualized by Coomassie Brilliant Blue R250 staining.
Measurement of GST enzyme activity
GST activity was measured spectrophotometrically using 1-chloro-2,4-dinitrobenzene (CDNB) and glutathione (GSH) as standard substrates83. Briefly, 1 µl of a test solution was added to 0.1 ml of a citrate-phosphate-borate buffer (pH 7.0) containing 5 mM CDNB and 5 mM GSH. Increase in absorbance at 340 nm min−1 was monitored at 30 °C and expressed as moles of CDNB conjugated with GSH per min per mg of protein using the molar extinction coefficient of the resultant 2,4-dinitrophenyl-glutathione: ε340 = 9600 M−1 cm−1.
Sampling and sequencing for population genetics study
S. litura was sampled from three locations in India (Delhi, Hyderabad and Matsyapuri), 11 locations in China, including Fujian, Guanxi, 2 locations in Guangzhou (Guangzhou and South China Normal University), Hainan, Hubei, Shanxi, Zhejiang, 3 locations in Hunan (Hunan1, Hunan2 and Hunan3), and 2 locations in Japan (Tsukuba and Okinawa). Four individuals were sampled from each location, except for Hunan1 (3 individuals). A total of 63 individuals were used in this study.
Mapping and SNP calling
First, mapping of reads of each individual to the reference genome was conducted. The proper mapping rate was about 70% for 56 individuals except for 7 individuals (Supplementary Table 21). Since the proper mapping rates for four individuals from the Shanxi population and three individuals from Fujian were extremely low, they were excluded from the population genomics analysis. SNP calling was conducted by comparing 56 genomes with the reference genome. Finally, a multiple VCF file was generated including 56 individuals. Sites with missing values or quality values below 20 were screened by VCFtools software84. In total, 46,595,432 SNPs were identified and included in this analysis.
Genetic diversity, population structure and balancing selection
The nucleotide diversity (π) of 14 local populations and pairwise F ST values were calculated using VCFtools software with window size 5000 bp, step 2500 bp. The genomic nucleotide diversity was obtained by averaging over the values of windows. The weighted F ST was calculated using the Weir and Cockerham estimator85. Based on the pairwise F ST, hierarchical cluster analysis was conducted using R software. Because of the small sample size in each sampling location, interpretation of population genomic analysis needs careful evaluation of the precision. The precision of π and F ST values were evaluated by parametric bootstrap with coalescent simulation86. Haplotypes of windows were generated using the population-specific π values multiplied by 5000 and 4 Nms calculated as 1/F ST−1. Two haplotypes were generated for each window. A thousand sets of haplotypes were generated independently and concatenated to make a bootstrap sample. For each of 100 bootstrap samples, the π values and pairwise F ST were calculated to estimate the standard errors. The adopted number of sets was less than the number of the scaffolds. Because the genome size of S. litura was about 4 × 108 bp, we mimicked the subsampling of windows that were separated by bp on average so that we could estimate approximate independence between the sub-sampled windows.
To confirm the observed population structure, we conducted a model-based structure analysis34,87. Based on the allele frequency divergence among the ancestral populations (P) and the membership coefficients that assign the populations to the ancestral populations (Q), we calculated the predicted allele frequency divergence between the population (QPQ t). We also analysed individual-level membership coefficients and the allele frequency divergence.
We further estimated the global pattern of migration by analysing the joint allele frequency spectrums in terms of the population histories and the migration patterns by ∂a∂i (diffusion approximation for demographic inference)39. To avoid the complex effect of selection, we analysed SNPs in introns. Out of ~20 million intronic SNPs, we randomly sampled 2 million SNPs. Based on the multi-dimensional scaling of F ST and the assignment of the individual genomes by structure, we constructed six population groups: the Indian local population (with the sample from Delhi), Indian migratory population (with the samples from Hyderabad and Matsyapuri), Chinese isolated population (with the samples from Guangzhou2 and Hunan1), Chinese local population (with the samples from Hunan3, Guangxi, Hainan, three individuals of Hunan2 and Hainan), Chinese migratory population (with the samples from Fujian, and one individual each of Hunan2, Hunan3, Hunan4, Zhejiang and Guangzhou1), and Japanese migrating population (with the samples from Okinawa and Tsukuba). To each pair of population groups we applied the IM (isolation with migration) model40 with population expansion/shrinkage. The estimated migration rates represent the number of migrating chromosomes per generation. To obtain the population sizes and the time of population splitting from the estimated relative values, we followed a previous study88 that assumes the generation time of 0.3 year and uses the standard mutation rate of 8.4 × 10−9 (per site per generation) from Drosophila 89. The standard errors were obtained by parametric bootstrap of coalescent simulation86. Assuming the estimated scenarios of population history, we generated 100 bootstrap samples of 2 million SNPs. To reflect the correlation structure between SNP loci, we assumed that they were evenly distributed on 28 chromosomes. SNPs on different chromosomes are independent. Noting that the mean distance between the neighbouring SNP loci (in bp) was
we set the recombination rate to be ρ = 2.3 × 10−5. We also tested two alternative values, ρ = 0 and ρ = 0.01, and obtained similar standard errors.
References
Ferry, N., Edwards, M. G., Gatehouse, J. A. & Gatehouse, A. M. Plant-insect interactions: molecular approaches to insect resistance. Curr. Opin. Biotechnol. 15, 155–161 (2004).
Sparks, T. C. & Nauen, R. IRAC: mode of action classification and insecticide resistance management. Pestic. Biochem. Physiol. 121, 122–128 (2015).
Arthropod Pesticide Resistance Database (Michigan State Univ., 2017); https://www.pesticideresistance.org
Wan, P., Wu, K., Huang, M., Yu, D. & Wu, J. Population dynamics of Spodoptera litura (Lepidoptera: Noctuidae) on Bt cotton in the Yangtze River Valley of China. Environ. Entomol. 37, 1043–1048 (2008).
Gouin, A. et al. Two genomes of highly polyphagous lepidopteran pests (Spodoptera frugiperda, Noctuidae) with different host-plant ranges. Sci. Rep. http://dx.doi.org/10.1038/s41598-017-10461-4 (2017).
Elsik, C. G. et al. Creating a honey bee consensus gene set. Genome Biol. 8, R13 (2007).
International Silkworm Genome Consortium. The genome of a lepidopteran model insect, the silkworm Bombyx mori. Insect Biochem. Mol. Biol. 38, 1036–1045 (2008).
You, M. et al. A heterozygous moth genome provides insights into herbivory and detoxification. Nat. Genet. 45, 220–225 (2013).
Zhan, S., Merlin, C., Boore, J. L. & Reppert, S. M. The monarch butterfly genome yields insights into long-distance migration. Cell 147, 1171–1185 (2011).
Mutanen, M., Wahlberg, N. & Kaila, L. Comprehensive gene and taxon coverage elucidates radiation patterns in moths and butterflies. Proc. Biol. Sci. 277, 2839–2848 (2010).
Heliconius Genome Consortium. Butterfly genome reveals promiscuous exchange of mimicry adaptations among species. Nature 487, 94–98 (2012).
Ahola, V. et al. The Glanville fritillary genome retains an ancient karyotype and reveals selective chromosomal fusions in Lepidoptera. Nat. Commun. 5, 4737 (2014).
Kanost, M. R. et al. Multifaceted biological insights from a draft genome sequence of the tobacco hornworm moth, Manduca sexta. Insect Biochem. Mol. Bio. 76, 118–147 (2016).
Briscoe, A. D. et al. Female behaviour drives expression and evolution of gustatory receptors in butterflies. PLoS Genet. 9, e1003620 (2013).
Gardiner, A., Barker, D., Butlin, R. K., Jordan, W. C. & Ritchie, M. G. Drosophila chemoreceptor gene evolution: selection, specialization and genome size. Mol. Ecol. 17, 1648–1657 (2008).
Guo, H. et al. Expression map of a complete set of gustatory receptor genes in chemosensory organs of Bombyx mori. Insect Biochem. Mol. Biol. 82, 74–82 (2017).
Kent, L. B. & Robertson, H. M. Evolution of the sugar receptors in insects. Evol. Biol. 9, 41 (2009).
Robertson, H. M. & Kent, L. B. Evolution of the gene lineage encoding the carbon dioxide receptor in insects. J. Insect Sci. 9, 19 (2009).
Koenig, C. et al. A reference gene set for chemosensory receptor genes of Manduca sexta. Insect Biochem. Mol. Biol. 66, 51–63 (2015).
Zhang, H. J. et al. Topological and functional characterization of an insect gustatory receptor. PLoS ONE 6, e24111 (2011).
Xu, W., Papanicolaou, A., Zhang, H. J. & Anderson, A. Expansion of a bitter taste receptor family in a polyphagous insect herbivore. Sci. Rep. 6, 23666 (2016).
Li, X., Schuler, M. A. & Berenbaum, M. R. Molecular mechanisms of metabolic resistance to synthetic and natural xenobiotics. Annu. Rev. Entomol. 52, 231–253 (2007).
Dermauw, W. & Van Leeuwen, T. The ABC gene family in arthropods: comparative genomics and role in insecticide transport and resistance. Insect Biochem. Mol. Biol. 45, 89–110 (2014).
Xie, X. et al. Genome-wide analysis of the ATP-binding cassette (ABC) transporter gene family in the silkworm, Bombyx mori. Mol. Biol. Rep. 39, 7281–7291 (2012).
Wang, R. L., Staehelin, C., Xia, Q. Q., Su, Y. J. & Zeng, R. S. Identification and characterization of CYP9A40 from the tobacco cutworm moth (Spodoptera litura), a cytochrome P450 gene induced by plant allelochemicals and insecticides. Int. J. Mol. Sci. 16, 22606–22620 (2015).
Zhao, C., Feng, X., Tang, T. & Qiu, L. Isolation and expression analysis of CYP9A11 and cytochrome P450 reductase gene in the beet armyworm (Lepidoptera: Noctuidae). J. Insect Sci. 15, 122 (2015).
Wang, R. L. et al. Identification of a novel cytochrome P450 CYP321B1 gene from tobacco cutworm (Spodoptera litura) and RNA interference to evaluate its role in commonly used insecticides. Insect Sci. 24, 235–247 (2017).
Claudianos, C. et al. A deficit of detoxification enzymes: pesticide sensitivity and environmental response in the honeybee. Insect Mol. Biol. 15, 615–636 (2006).
Hemingway, J. & Karunaratne, S. H. Mosquito carboxylesterases: a review of the molecular biology and biochemistry of a major insecticide resistance mechanism. Med. Vet. Entomol. 12, 1–12 (1998).
Tsubota, T. & Shiotsuki, T. Genomic and phylogenetic analysis of insect carboxyl/cholinesterase genes. J. Pestic. Sci. 35, 310–314 (2010).
Bravo, A., Likitvivatanavong, S., Gill, S. S. & Soberon, M. Bacillus thuringiensis: A story of a successful bioinsecticide. Insect Biochem. Mol. Biol. 41, 423–431 (2011).
Atsumi, S. et al. Single amino acid mutation in an ATP-binding cassette transporter gene causes resistance to Bt toxin Cry1Ab in the silkworm, Bombyx mori. Proc. Natl Acad. Sci. USA 109, E1591–1598 (2012).
Tay, W. T. et al. Insect resistance to Bacillus thuringiensis toxin Cry2Ab is conferred by mutations in an ABC transporter subfamily A protein. PLoS Genet. 11, e1005534 (2015).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Wan, X. et al. DNA sequence variation of the tobacco cutworm, Spodoptera litura (Lepidoptera: Noctuidae), determined by mitochondrial A+T-rich region and nuclear ITS2 sequences. Biochem. Genet. 49, 760–787 (2011).
Murata, M. & Tojo, S. Flight capability and fatty acid level in triacylglycerol of long-distance migratory adults of the common cutworm, Spodoptera litura. Zoolog. Sci. 21, 181–188 (2004).
Tojo, S. et al. Overseas migration of the common cutworm, Spodoptera litura (Lepidoptera: Noctuidae), from May to mid-July in East Asia. Appl. Entomol. Zool. 48, 141–140 (2013).
Wang, B., Yim, S. Y., Lee, J. Y., Liu, J. & Ha, K. J. Future change of Asian-Australian monsoon under RCP 4.5 anthropogenic warming scenario. Clim. Dynam. 42, 83–100 (2014).
Gutenkunst, R. N., Hernandez, R. D., Williamson, S. H. & Bustamante, C. D. Inferring the joint demographic history of multiple populations from multidimensional SNP frequency data. PLoS Genet. 5, e1000695 (2009).
Hey, J. & Nielsen, R. Multilocus methods for estimating population sizes, migration rates and divergence time, with applications to the divergence of Drosophila pseudoobscura and D. persimilis. Genetics 167, 747–760 (2004).
Gnerre, S. et al. High-quality draft assemblies of mammalian genomes from massively parallel sequence data. Proc. Natl Acad. Sci. USA 108, 1513–1518 (2011).
Li, R. et al. De novo assembly of human genomes with massively parallel short read sequencing. Genome Res. 20, 265–272 (2010).
Li, R. et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25, 1966–1967 (2009).
Stanke, M., Steinkamp, R., Waack, S. & Morgenstern, B. AUGUSTUS: a web server for gene finding in eukaryotes. Nucleic Acids Res. 32, W309–312 (2004).
Korf, I. Gene finding in novel genomes. Bioinformatics 5, 59 (2004).
Birney, E., Clamp, M. & Durbin, R. GeneWise and Genomewise. Genome Res. 14, 988–995 (2004).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Bairoch, A. & Apweiler, R. The SWISS-PROT protein sequence database and its supplement TrEMBL in 2000. Nucleic Acids Res. 28, 45–48 (2000).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Mount, D. W. Using the Basic Local Alignment Search Tool (BLAST). CSH Protoc. 2007, pdb.top17 (2007).
Mulder, N. & Apweiler, R. InterPro and InterProScan: tools for protein sequence classification and comparison. Methods Mol. Biol. 396, 59–70 (2007).
Ashburner, M. et al. Gene Ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Bao, Z. & Eddy, S. R. Automated de novo identification of repeat sequence families in sequenced genomes. Genome Res. 12, 1269–1276 (2002).
Price, A. L., Jones, N. C. & Pevzner, P. A. De novo identification of repeat families in large genomes. Bioinformatics 21, i351–358 (2005).
Li, H. et al. TreeFam: a curated database of phylogenetic trees of animal gene families. Nucleic Acids Res. 34, D572–580 (2006).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Posada, D. & Crandall, K. A. MODELTEST: testing the model of DNA substitution. Bioinformatics 14, 817–818 (1998).
Huelsenbeck, J. P. & Ronquist, F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17, 754–755 (2001).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Douzery, E. J., Snell, E. A., Bapteste, E., Delsuc, F. & Philippe, H. The timing of eukaryotic evolution: does a relaxed molecular clock reconcile proteins and fossils? Proc. Natl Acad. Sci. USA 101, 15386–15391 (2004).
Benton, M. J. & Donoghue, P. C. Paleontological evidence to date the tree of life. Mol. Biol. Evol. 24, 26–53 (2007).
De Bie, T., Cristianini, N., Demuth, J. P. & Hahn, M. W. CAFE: a computational tool for the study of gene family evolution. Bioinformatics 22, 1269–1271 (2006).
Baxter, S. W. et al. Linkage mapping and comparative genomics using next-generation RAD sequencing of a non-model organism. PLoS ONE 6, e19315 (2011).
Van Ooijen, J. W. Multipoint maximum likelihood mapping in a full-sib family of an outbreeding species. Genet. Res. 93, 343–349 (2011).
Shimomura, M. et al. KAIKObase: an integrated silkworm genome database and data mining tool. Genomics 10, 486 (2009).
Li, X. et al. Outbred genome sequencing and CRISPR/Cas9 gene editing in butterflies. Nat. Commun. 6, 8212 (2015).
Wang, Y. et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 40, e49 (2012).
Finn, R. D., Clements, J. & Eddy, S. R. HMMER web server: interactive sequence similarity searching. Nucleic Acids Res. 39, W29–37 (2011).
Keller, O., Odronitz, F., Stanke, M., Kollmar, M. & Waack, S. Scipio: using protein sequences to determine the precise exon/intron structures of genes and their orthologs in closely related species. Bioinformatics 9, 278 (2008).
Slater, G. S. & Birney, E. Automated generation of heuristics for biological sequence comparison. Bioinformatics 6, 31 (2005).
Werck-Reichhart, D. & Feyereisen, R. Cytochromes P450: a success story. Genome Biol. 1, 1–9 (2000).
Sezutsu, H., Le Goff, G. & Feyereisen, R. Origins of P450 diversity. Phil. Trans. R. Soc. Lond. B Biol. Sci. 368, 20120428 (2013).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013).
Hasegawa, M., Kishino, H. & Yano, T. Dating of the human-ape splitting by a molecular clock of mitochondrial DNA. J. Mol. Evol. 22, 160–174 (1985).
Le, S. Q. & Gascuel, O. An improved general amino acid replacement matrix. Mol. Biol. Evol. 25, 1307–1320 (2008).
Yang, Z. Maximum likelihood phylogenetic estimation from DNA sequences with variable rates over sites: approximate methods. J. Mol. Evol. 39, 306–314 (1994).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Sanderson, M. J. & Wojciechowski, M. F. Improved bootstrap confidence limits in large-scale phylogenies, with an example from Neo-Astragalus (Leguminosae). Syst. Biol. 49, 671–685 (2000).
Stamatakis, A., Hoover, P. & Rougemont, J. A rapid bootstrap algorithm for the RAxML Web servers. Syst. Biol. 57, 758–771 (2008).
Laemmli, U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227, 680–685 (1970).
Habig, W. H., Pabst, M. J. & Jakoby, W. B. Glutathione S-transferases: the first enzymatic step in mercapturic acid formation. J. Biol. Chem. 249, 7130–7139 (1974).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Weir, B. S. & Cokerham, C. C. Estimating F-statistics for the analysis of population structure. Evol. Int. J. Organic Evol. 38, 1358–1370 (1984).
Hudson, R. R. Generating samples under a Wright-Fisher neutral model of genetic variation. Bioinformatics 18, 337–338 (2002).
Raj, A., Stephens, M. & Pritchard, J. K. fastSTRUCTURE: variational inference of population structure in large SNP data sets. Genetics 197, 573–589 (2014).
Zhan, S. et al. The genetics of monarch butterfly migration and warning colouration. Nature 514, 317–321 (2014).
Haag-Liautard, C. et al. Direct estimation of per nucleotide and genomic deleterious mutation rates in Drosophila. Nature 445, 82–85 (2007).
i5K Consortium. The i5K Initiative: advancing arthropod genomics for knowledge, human health, agriculture, and the environment. J. Hered. 104, 595–600 (2013).
Acknowledgements
This paper was supported by the grant of the One Thousand Foreign Experts Recruitment Program of the Chinese Government (No. WO20125500074).
Author information
Authors and Affiliations
Contributions
K.M., K.K-O., K.P.A. and Q.F. designed the whole project. T.C., Y.W., L.P., Y.G., C.L., D.L. and B.F. performed sequencing, genome assembly and analysis. K.M., L.H., H.G., J.L., F.H., N.M., E.J-J., R.K.B., T.S. and A.J. worked on manual annotation. K.K-O. and R.K.S. established BC1 of S. litura inbred strains. R.V.C., A.T. and K.P.A. worked linkage analysis. K.Y., L.F., Z.C., H.G., X.W., Z.L., Y.C. and K.M. carried out pesticide tolerance experiments. W.L. and S.L. performed transcriptome analysis. K.M., J.L., A.P., K.P.A. and Q.F. collected S. litura samples for population genetics. J.W. and H.K. performed population genetics analysis. T.C., H.K., K.P.A., A.J., F.H., L.H. and K.M. wrote the manuscript. M.R.G., E.d’A., G.S., Q.X. and Q.F. revised manuscript. E.d’A. and M.R.G. coordinated with the S. frugiperda genome community and the i5k initiative90. T.C., J.W., Y.W., R.V.C., L.H., K.Y., L.F. and W.L. equally contributed to this work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This Whole Genome Shotgun project has been deposited at DDBJ/ENA/GenBank under the accession MTZO00000000 and the BioProject accession PRJNA344815. Sequence reads of genome assembly and resequencing (local populations for population genetics) have been deposited in the NCBI sequence read archive (SRA) under accessions SRP095695. RNA-Seq data used for genome assessment and gene prediction have been submitted to DDBJ DRA under accession numbers DRA005501. RNA-Seq data for GR transcriptome analysis and toxin treatment experiments have been submitted to NCBI SRA under BioProject accession PRJNA368925. The RAD-seq data generated to develop linkage and physical maps has been submitted to GenBank under the BioProject accession number PRJNA369226.
Electronic supplementary material
Supplementary Information
Supplementary Methods and Discussion; Supplementary Figures 1–12; Supplementary Tables 1–24
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cheng, T., Wu, J., Wu, Y. et al. Genomic adaptation to polyphagy and insecticides in a major East Asian noctuid pest. Nat Ecol Evol 1, 1747–1756 (2017). https://doi.org/10.1038/s41559-017-0314-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-017-0314-4
This article is cited by
-
Genomic identification and evolutionary analysis of chemosensory receptor gene families in two Phthorimaea pest species: insights into chemical ecology and host adaptation
BMC Genomics (2024)
-
Few chemoreceptor genes in the ambrosia beetle Trypodendron lineatum may reflect its specialized ecology
BMC Genomics (2024)
-
A chromosome-level genome assembly of Sesamia inferens
Scientific Data (2024)
-
Knock-Out of ACY-1 Like Gene in Spodoptera litura Supports the Notion that FACs Improve Nitrogen Metabolism
Journal of Chemical Ecology (2024)
-
Chromosome-level genome of black cutworm provides novel insights into polyphagy and seasonal migration in insects
BMC Biology (2023)