Abstract
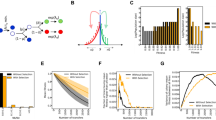
Mutations provide the variation that drives evolution, yet their effects on fitness remain poorly understood. Here we explore how mutations in the essential enzyme adenylate kinase (Adk) of Escherichia coli affect multiple phases of population growth. We introduce a biophysical fitness landscape for these phases, showing how they depend on molecular and cellular properties of Adk. We find that Adk catalytic capacity in the cell (the product of activity and abundance) is the major determinant of mutational fitness effects. We show that bacterial lag times are at a well-defined optimum with respect to Adk’s catalytic capacity, while exponential growth rates are only weakly affected by variation in Adk. Direct pairwise competitions between strains show how environmental conditions modulate the outcome of a competition where growth rates and lag times have a tradeoff, shedding light on the multidimensional nature of fitness and its importance in the evolutionary optimization of enzymes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Sanjuan, R., Moya, A. & Elena, S. F. The distribution of fitness effects caused by single-nucleotide substitutions in an RNA virus. Proc. Natl Acad. Sci. USA 101, 8396–8401 (2004).
Bershtein, S., Segal, M., Bekerman, R., Tokuriki, N. & Tawfik, D. S. Robustness-epistasis link shapes the fitness landscape of a randomly drifting protein. Nature 444, 929–932 (2006).
Bershtein, S., Mu, W. & Shakhnovich, E. I. Soluble oligomerization provides a beneficial fitness effect on destabilizing mutations. Proc. Natl Acad. Sci. USA 109, 4857–4862 (2012).
Bershtein, S., Mu, W., Serohijos, A. W., Zhou, J. & Shakhnovich, E. I. Protein quality control acts on folding intermediates to shape the effects of mutations on organismal fitness. Mol. Cell 49, 133–144 (2013).
Bershtein, S. et al. Protein homeostasis imposes a barrier on functional integration of horizontally transferred genes in bacteria. PLoS Genet. 11, e1005612 (2015).
Rodrigues, J. V. et al. Biophysical principles predict fitness landscapes of drug resistance. Proc. Natl Acad. Sci. USA 113, E1470–E1478 (2016).
Gong, L. I., Suchard, M. A. & Bloom, J. D. Stability-mediated epistasis constrains the evolution of an influenza protein. eLife 2, e00631 (2013).
Orr, H. A. Fitness and its role in evolutionary genetics. Nat. Rev. Genet. 10, 531–539 (2009).
Elena, S. F. & Lenski, R. E. Evolution experiments with microorganisms: the dynamics and genetic bases of adaptation. Nat. Rev. Genet. 4, 457–469 (2003).
Dykhuizen, D. E. & Hartl, D. L. Selection in chemostats. Microbiol. Rev. 47, 150–168 (1983).
Biel, S. W. & Hartl, D. L. Evolution of transposons: natural selection for Tn5 in Escherichia coli K12. Genetics 103, 581–592 (1983).
Fridman, O., Goldberg, A., Ronin, I., Shoresh, N. & Balaban, N. Q. Optimization of lag time underlies antibiotic tolerance in evolved bacterial populations. Nature 513, 418–421 (2014).
Bajaj, K., Chakrabarti, P. & Varadarajan, R. Mutagenesis-based definitions and probes of residue burial in proteins. Proc. Natl Acad. Sci. USA 102, 16221–16226 (2005).
Semisotnov, G. V. et al. Study of the “molten globule” intermediate state in protein folding by a hydrophobic fluorescent probe. Biopolymers 31, 119–128 (1991).
Navarro, S. & Ventura, S. Fluorescent dye ProteoStat to detect and discriminate intracellular amyloid-like aggregates in Escherichia coli. Biotechnol. J. 9, 1259–1266 (2014).
Dykhuizen, D. E., Dean, A. M. & Hartl, D. L. Metabolic flux and fitness. Genetics 115, 25–31 (1987).
Peters, J. M. et al. A comprehensive, CRISPR-based functional analysis of essential genes in bacteria. Cell 165, 1493–1506 (2016).
Hegreness, M., Shoresh, N., Hartl, D. & Kishony, R. An equivalence principle for the incorporation of favorable mutations in asexual populations. Science 311, 1615–1617 (2006).
Cha, R. S., Zarbl, H., Keohavong, P. & Thilly, W. G. Mismatch amplification mutation assay (MAMA): application to the c-H-ras gene. PCR Meth. Appl. 2, 14–20 (1992).
Jiang, L., Mishra, P., Hietpas, R. T., Zeldovich, K. B. & Bolon, D. N. Latent effects of Hsp90 mutants revealed at reduced expression levels. PLoS Genet. 9, e1003600 (2013).
Serohijos, A. W. & Shakhnovich, E. I. Merging molecular mechanism and evolution: theory and computation at the interface of biophysics and evolutionary population genetics. Curr. Opin. Struct. Biol. 26, 84–91 (2014).
Serohijos, A. W., Lee, S. Y. & Shakhnovich, E. I. Highly abundant proteins favor more stable 3D structures in yeast. Biophys. J. 104, L1–L3 (2013).
Hartl, D. L. & Clark, A.G. Principles of population genetics 4th ed. (Sinauer, 2007).
Manhart, M., Adkar, B. V. & Shakhnovich, E. I. Tradeoffs between microbial growth phases lead to frequency-dependent and non-transitive selection. Preprint at http://bioRxiv.org/content/early/2016/12/23/096453 (2016).
Lambert, G., Liao, D., Vyawahare, S. & Austin, R. H. Anomalous spatial redistribution of competing bacteria under starvation conditions. J. Bacteriol. 193, 1878–1883 (2011).
Palkova, Z. Multicellular microorganisms: laboratory versus nature. EMBO Rep. 5, 470–476 (2004).
Hibbing, M. E., Fuqua, C., Parsek, M. R. & Peterson, S. B. Bacterial competition: surviving and thriving in the microbial jungle. Nat. Rev. Microbiol. 8, 15–25 (2010).
Todar, K. Online Textbook of Bacteriology (2015); http://www.textbookofbacteriology.net/
Adkar, B. V. et al. Protein model discrimination using mutational sensitivity derived from deep sequencing. Structure 20, 371–381 (2012).
Muller, C. W., Schlauderer, G. J., Reinstein, J. & Schulz, G. E. Adenylate kinase motions during catalysis: an energetic counterweight balancing substrate binding. Structure 4, 147–156 (1996).
Acknowledgements
This work was supported by NIH award R01 GM068670 to E.I.S. M.Ma. was supported by NIH award F32 GM116217. We thank S. Bershtein and A. Serohijos for helpful discussions.
Author information
Authors and Affiliations
Contributions
B.V.A. and E.I.S. designed the research; B.V.A., S.B., J.T. and M.Mu. performed experiments; B.V.A., M.Ma., S.B. and E.I.S. analysed the data; B.V.A., M.Ma., S.B. and E.I.S. wrote the paper. All authors edited and approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Methods; Supplementary Figures 1–12; Supplementary Tables 1–3 (PDF 2072 kb)
Supplementary Dataset 1
Growth curve data for Adk in E. coli. (XLS 207 kb)
Rights and permissions
About this article
Cite this article
Adkar, B., Manhart, M., Bhattacharyya, S. et al. Optimization of lag phase shapes the evolution of a bacterial enzyme. Nat Ecol Evol 1, 0149 (2017). https://doi.org/10.1038/s41559-017-0149
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41559-017-0149
This article is cited by
-
Decreased thermal niche breadth as a trade-off of antibiotic resistance
The ISME Journal (2022)
-
Pervasive cooperative mutational effects on multiple catalytic enzyme traits emerge via long-range conformational dynamics
Nature Communications (2021)
-
Taxon-specific microbial growth and mortality patterns reveal distinct temporal population responses to rewetting in a California grassland soil
The ISME Journal (2020)
-
Growth tradeoffs produce complex microbial communities on a single limiting resource
Nature Communications (2018)
-
Fermentation trip: amazing microbes, amazing metabolisms
Annals of Microbiology (2018)