Abstract
Rice paddies are a major source of methane emissions. To meet the food demand of the growing population and to cope with global warming, reducing greenhouse gases and enhancing yields are critical. Here we demonstrate that a loss-of-function rice allele, gs3, mitigates methane emissions from methanogens by allocating more photosynthates to the grain and less to the root and increases yield by enlarging grain size and weight.
Similar content being viewed by others
Main
Atmospheric methane is a potent greenhouse gas and is second only to carbon dioxide in contributing to global warming. Rice paddies are the largest anthropogenic source of methane, accounting for 7–17% of the total atmospheric methane1,2. This is due to the warm, waterlogged soil and soil nutrients, which facilitate the production of methane through microbe-mediated methanogenesis2. In addition, with increasing demand for food, sustainable rice production technologies are urgently needed. Currently, efforts to mitigate methane emissions from rice paddies are focused largely on farming practices and measures, which can be challenging to implement effectively3. Beyond farming practices, many previous studies demonstrated that photosynthate allocation modulation (grain versus root) can mitigate methane emissions through genetic trait modulation4,5,6,7,8. For example, rice SUSIBA2, which harbours a transcription factor for sugar signalling in barley2, emits less methane by increasing starch content and spikelet number4. In this line, removing spikelet has been reported to notably increase methane emissions compared with wild type6,7,8,9. While the development of new rice varieties with reduced methane emissions is crucial, there is currently a lack of research on utilizing breeding strategies to harness the natural variation in rice that does not involve additional laborious manpower and can get over numerous obstacles from genetic modification. In this study, we developed a near-isogenic rice line, Milany360, by introgressing the gs3 mutant allele from a rice variety Shindongjin to increase grain yield and decrease methane flux by shifting the allocation of photosynthates away from the root and towards the grain by enlarging grain size.
The gene GS3, which was obtained through quantitative trait locus cloning as a gene on chromosome 3 controlling grain size, negatively regulates organ size via its organ-size-regulation domain9,10,11. In this study, we investigated the natural loss-of-function gs3 mutant allele, which harbours an A-to-stop-codon mutation at the 165th nucleotide position in the second exon in the organ-size-regulation domain9,12 (Extended Data Fig. 1). Because introgression of gs3 in rice varieties increases grain yield13, we hypothesized that the gs3 variety would show lower allocation of photosynthates to the root, resulting in low methane production by methanogenic archaea and higher allocation of photosynthates to the sink organ contributing the enhanced grain size.
Two Korean rice varieties, Saeilmi and Shindongjin, were crossed in 2012. Then a near-isogenic line (NIL) Milayang360 was selected from BC1F1 to BC2F1 progeny using marker-assisted backcrosses. Subsequently, a chip array revealed 99.9% genetic similarity between Milayang360 and Saeilmi (Fig. 1a and Extended Data Figs. 1, 2a).
To determine the effect of gs3 on methane emissions, we cultivated Milyang360 and Saeilmi for two years at two locations, Miryang and JinJu field stations of Rural Development Administration, South Korea, and measured the methane emission rate and the cumulative methane emissions (Fig. 1b). The average seasonal cumulative methane emissions of Milyang360 (130 kg ha–1) were 16% lower than those of Saeilmi (156 kg ha–1). Intriguingly, after the heading stage, the cumulative methane emissions of Milyang360 were 50 kg ha–1, a 37.5% reduction compared with Saeilmi (81 kg ha–1) (Fig. 1b). Most previous studies targeted the early growth stage for reducing methane emissions because the methane emissions peak was detected right after irrigation. However, recent emerging data also detected the peak at the heading stages, when photosynthate allocation modulation (grain versus root) may occur6.
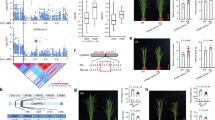
a, Phenotype of Milyang360 and Saeilmi plants in the rice paddy field. b, Seasonal methane emission and methane flux rates measured in triplicate in 2021 and 2022. Error bars represent standard error. Asterisks indicate significant differences (*P < 0.05; **P < 0.01; one-way analysis of variance (ANOVA)); P = 0.008 after heading stage of CH4 emission rate; P value of cumulative methane flux was 0.035 and 0.040 in 2021 and 2022, respectively. The data are shown as mean ± s.d.; n = 12 for average CH4 emissions and n = 6 for cumulative CH4 emissions in 2021 and 2022. c, Soluble sucrose contents of the different parts of two gs3 alleles at different growth stages. The data are shown as mean ± s.d. and n = 10. At the heading stage, P = 0.034 and 0.007 on panicle and root, respectively. At the grain-filling stage, P = 0.008 and 0.007 on panicle and root, respectively. d, Heatmap showing the abundances of methanogens and fermentation-related microbes. At the tillering stage, P = 0.152; however, after the heading stage, the relative abundance of microbes was significantly lower in Milyang360 rhizosphere than in Saeilmi rhizosphere. Relative abundance data were transformed to centred-log ratio, which involves dividing each element in the count vector by the geometric mean of all elements and then taking the natural logarithm of each ratio. This transformation enables the conversion of data from the simplex space to real co-ordinate space, and n = 15. e, Microbial community and beta-diversity analyses of the rhizospheres of five plants in 2021 and ten plants in 2022 by metagenome sequencing. Arrows indicate methanogenesis. PC1: methanogens. PC2: fermentation-related microbes. f, Quantification of methanogens and methanogenesis-related enzymes based on the number of mcrA gene copies in different gs3 alleles at different growth stages. Asterisks indicate significant differences (*P < 0.05; **P < 0.01; one-way ANOVA). At the heading stage, P = 0.035 and 0.0007 for DNA and RNA, respectively. At the grain-filling stage, P = 0.027 and 0.006 for DNA and RNA, respectively, and n = 10. FW, fresh weight.
To understand how gs3 reduces methane emissions, we measured the soluble sucrose content of different plant parts at different growth stages, analysed the relative expression of starch synthesis genes in grains and panicles, investigated root exudate contents and composition and their related gene expression and profiled microbial communities, including methanogens and fermentation-related microbes. After the heading stage, Milyang360 exhibited a 56% reduction in soluble sucrose content in roots compared with Saeilmi. However, the soluble sucrose content was 61% higher in the panicle of Milyang360 than in that of Saeilmi (Fig. 1c). However, there was no difference in photosynthesis rate between the two varieties (Extended Data Table 1). This result agrees with a previous study13 that showed that photosynthates are translocated more to above-ground than to underground organs in gs3 allele rice. The overall relative abundances of rhizosphere microbiota of Milyang360 and Saeilmi were not significantly different at the tillering (vegetative) stage; however, after the heading (reproductive) stage, the relative abundances of methanogens and fermentation-inducing microbes were significantly lower in Milyang360. The number of mcrA gene copies was significantly lower in Milyang360 than in Saeilmi (Fig. 1f). In Milyang360 rhizosphere, Proteobacteria and Verrucomicrobia were relatively highly abundant, whereas Firmicutes and Actinobacteria (putative fermentation-related bacteria) had relatively low abundances (Extended Data Fig. 3a). The relative abundances of obligate methanogens (Methanobacteriaceae and Methanotrichaceae) and putative fermentation-dependent microbes (Peptococcaceae, Melioribacteriaceae and Micrococcaceae), which ferment root exudates to produce organic acids and methane14, were significantly lower on the basis of their relative abundances in the Milyang360 rhizosphere after the heading stage. Beta-diversity analysis showed that rhizosphere microbial communities were clustered on the basis of genomic differences (Fig. 1d–f).
To validate the rhizosphere release of photosynthates, which provide the food source for methanogens, the root exudate content in the rhizosphere and the expression of exudate transporter genes in rice root were measured. After the heading stage, the dissolved organic carbon, carbohydrate and total organic acid contents of Milyang360 were significantly lower than those of Saeilmi by 21%, 26% and 32%, respectively (Fig. 2a). In addition, the expression of root exudate transporter genes, including OsALMT1 (which exports organic acids), OsSWEET11, OsSWEET14 and OsMFS1 (which export sugars such as glucose and sucrose), were significantly lower in Milyang360 after the heading stage (Fig. 2b).
a, Analysis of root exudates, including dissolved organic carbon, carbohydrates and total organic acids. One-way ANOVA was used for statistical analysis. At the heading stage, the P values for dissolved organic acid, carbohydrate and total organic acid were 0.038, 0.007 and 0.008, respectively. And at the grain-filling stage, P values were 0.041, 0.008 and 0.008, respectively. Error bars represent standard error. The data are shown as mean ± s.d. and n = 10. b, Relative expression of genes encoding root exudate transporters in the root. Green, pink and yellow lines indicate tillering, heading and grain-filling stages, respectively. One-way ANOVA was used for statistical analysis (*P < 0.05; **P < 0.01). The data are shown as mean ± s.d. and n = 10. At the heading stage, the P values of OsALMT1, OsSWEET11, OsSWEET14 and OsMFS1 were 0.008, 0.042, 0.035 and 0.037, respectively. At the grain-filling stage, the P values were 0.008, 0.007, 0.008 and 0.007, respectively. c, Seasonal methane flux and methane emission rates of different gs3 alleles grown under different nitrogen levels. One-way ANOVA was used for statistical analysis. The data are shown as mean ± s.d. and n = 12. For the total methane flux, the P value was 0.026 between L-Milyang360 and L-Saeilmi, 0.035 between N-Milyang360 and N-Saeilmi, 0.045 between L-Milyang360 and N-Milyang360 and 0.007 between L-Milyang360 and N-Saeilmi. For the CH4 emission rate, the P value was 0.008 between L-Milyang360 and L-Saeilmi, 0.006 between N-Milyang360 and N-Saeilmi, 0.042 between L-Milyang360 and N-Milyang360 and 0.005 between L-Milyang360 and N-Saeilmi. d, Agronomic traits of different gs3 alleles under different nitrogen levels were analysed using Kruskal–Wallis test (P < 0.05). Letters represent significant differences between gs3 alleles under different nitrogen levels (P < 0.05; Tukey’s honest significant difference test). The data are shown as mean ± s.d. and n = 10. The P values of the milling rice yield, thousand-grain weight and grain length were 0.042, 0.020 and 0.023, respectively. For the thousand-grain weight, the P value between L-Milyang360 and N-Saeilmi was 0.008 and for grain length was 0.007. e, Relative expression of starch synthesis genes in the two gs3 alleles under different nitrogen levels. Most genes were highly expressed in Milyang360. The degree of statistically significant difference in relative gene expression is indicated by the circle size. The data are shown as mean ± s.d. and n = 10. f, Schematic model depicting the reduction in methane emissions through the introduction of the gs3 allele, which has an additive effect with low nitrogen input. *P < 0.05; **P < 0.01. LN, low nitrogen; NN, normal nitrogen; NS, non-significant.
The process of starch synthesis in grains requires sucrose import and transition15,16. To evaluate whether photosynthates allocated to the sink contribute to starch accumulation in grain, contributing yield increase in Milyang360, the relative expression of nine genes related to starch biosynthesis was quantified in different plant parts and under different nitrogen levels. As expected, OsSUT1, OsSUT5 and OsDOF11, which regulate sucrose import into the cell and sucrose transport to the grain, were highly expressed in Milyang360. In addition, sucrose transition-related and starch synthesis-related genes were highly expressed in the grains and panicles of Milyang360, regardless of the nitrogen level (Fig. 2e).
Decreasing nitrogen input is also known to reduce methane flux in rice paddies17,18. To investigate the additive effect of photosynthate allocation and low nitrogen input on methane emissions, we conducted experiments on rice paddies that had been maintained for two decades under low and normal nitrogen conditions. Our results showed that reducing the nitrogen level and increasing the sink strength had an additive effect on mitigating methane emissions. The nitrogen effect involved in the mitigation of methane flux was approximately 10% regardless of the variety, which was statistically significant (Extended Data Table 2). Interestingly, compared with conventional fertilization (nitrogen (N)-treated) Saeilmi (N-Saeilmi), low-fertilization (L-treated) Milayng360 (L-Milyang360) plants exhibited 24% less methane emission during whole rice cultivation and 63% lower methane flux after heading (Fig. 2c). Milyang360 showed 6% higher milling grain yield following N treatment and 14% higher milling grain yield following low nitrogen input compared with those of Saeilmi by enlarging grain size (Fig. 2d).
In this study, we demonstrated that reducing nitrogen input and introducing the gs3 allele have an additive effect on the reduction in methane emissions. Root exudate secretion was affected by nitrogen levels as well as by an increase in sink strength. The root exudate content of L-Milyang360 plants was significantly lower than that of N-Saeilmi plants. In addition, the dissolved organic carbon, carbohydrate and organic acid contents of L-Milyang360 plants were approximately 30%, 46% and 47% lower than those of N-Saeilmi plants, respectively (Extended Data Fig. 4). Moreover, compared with N-Saeilmi, the soluble sucrose and starch contents of L-Milyang360 were higher in above-ground parts and lower in underground parts (Extended Data Table 1). The abundance of methanogens in L-Milyang360 was significantly lower than that in L-Saeilmi and similar to that in L-Saeilmi and N-Milayng360 (Extended Data Fig. 3b). Beta-diversity analysis showed that the metagenome sequencing data were grouped by the nitrogen level at the tillering stage but by both the nitrogen level and the gs3 allele at the heading stage (Extended Data Fig. 3a). Root exudate release, microbial community formation and methane emissions were affected by nitrogen input and photosynthate allocation.
In conclusion, the introduction of the gs3 mutant allele in rice decreased methane emissions and showed an additive effect with low nitrogen input. The null gs3 allele increased sink strength by enhancing the allocation of photosynthates to above-ground (grain) rather than to below-ground (root), contributing to a decrease in root exudate release. Collectively, our results suggest that Milyang360 harbouring the gs3 allele could serve as a model plant for managing methane emissions through conventional breeding programmes. In 2020, rice paddies emitted 29.8 Tg of methane for Food and Agriculture Organization Tier 1 (http://www.fao.org/faostat) around the world. We suggest that if the gs3 allele effect is applied on scale factor for rice cultivar (SFr) of methane emissions calculating formula, it can reduce global methane by 4.8 Tg (29.8 Tg × 0.16 = 4.8 Tg). Taken together, introgression of the gs3 allele in rice germ plasm can serve to not only reduce greenhouse gas emissions and nitrogen fertilizer input but also meet the escalating global food demand.
Methods
Breeding of the NIL Milyang360
In 2019, following the method outlined by ref. 19, Milyang360 was generated through a marker-assisted backcrossing process using the gs3 KASP marker. The NIL Milyang360 was created using foreground selection on BC1F1, BC2F1 and BC2F2. The generation process was accelerated in the greenhouse during the winter season. To determine the introgression ratio and select the NIL, background selection was performed on BC2F5 plants. The selected line was then tested through yield trials in 2017, replicated yield trials in 2018 and regional adaptability yield trials in 15 regions of South Korea over a period of 3 years. The introgression ratio of Milyang360 was determined using the KNU Affymetrix Axiom Oryza 580K Genotyping Array (Thermo Fisher Scientific). Milyang360 and Saeilmi seeds were deposited in the gene bank of the National Institute of Agricultural Science, Rural Development Administration (accession numbers: K278471 and IT318798, respectively).
Field experimental design and cultivation management
The experimental fields were located in Miryang, South Korea (35° 29′ 32.2872″ N, 128° 44′ 32.1972″ E) and Jinju, South Korea (35° 9′ 12.14268″ N, 128° 6′ 3.73284″ E) during the period 2021–2022. Before transplanting, conventional tillage was performed, and no straw or organic matter was applied. The rice varieties used were Milyang360 and Saeilmi, which were transplanted after 30 days of sowing. The seedlings were transplanted simultaneously in both experimental fields (Miryang and Jinju) on 5 Jun, with a spacing of 30 cm × 15 cm. Each field (8 m × 70 m) had three replicate plots. Chemical fertilizer (N-P2O5-K2O: 21-17-17) was applied at a total rate of 90 kg ha−1 N during rice cultivation. This fertilizer was split into three applications: 54 kg ha−1 N as basal application before transplanting, 18 kg ha−1 N at 20 days after transplanting (DAT) and another 18 kg ha−1 N at 65 DAT under normal nitrogen conditions. In the low-nitrogen plots, only half the amount of nitrogen used in normal plots was applied. The normal-nitrogen and low-nitrogen fields have been maintained for 20 years. Both sites were continuously flooded until 110 DAT as part of field management practices. During the rice growing season, continuous flooding was applied so that a water level of 5–7 cm above the soil surface was consistently maintained .
Measurement of methane emissions and soil redox potential on rice paddy
A closed-chamber experiment20,21 was performed to analyse methane emissions from the seasonal rice crops cultivated in Miryang. Another closed-chamber experiment was performed in Jinju for two years under low and normal nitrogen levels. Gas samples were collected from each treatment and replicated every 15 min for a total of 30 min using 50 ml plastic syringes with a three-way stopcock in each of three glass chambers (62 cm diameter, 112 cm height) in each plot. The chambers were placed on flooded soil, covering eight rice hills each, and had four holes at the bottom to maintain a water level of 5–7 cm above the soil–water interface. Gas samples were collected three times a day (between 8:00 and 12:00 lt) to calculate the daily methane flux twice a week during the rice cultivation period. The chambers were equipped with a fan for air circulation and a thermometer for monitoring the air temperature during sampling. The chambers were kept open except during sampling, and gas samples were immediately transferred to air-evacuated glass vials for analysis in the laboratory. Methane concentration was measured using Agilent 7890A gas chromatograph equipped with a stainless steel Porapak NQ column (Q80-100 mesh, cat. no. 1002-11105) and a flame ionization detector. The temperatures of the column, injector and detector were set at 80 °C, 100 °C and 110 °C, respectively, and helium and hydrogen were used as the carrier and burning gases, respectively. To calculate methane flux, the rate of change in methane concentration per unit surface area of the chamber within a specified time interval was calculated using the following equation:
The equation considers factors such as methane flux (F, mg m–2 h–1 CH4), chamber volume (V), chamber area (A), rate of methane accumulation (∆/∆t) and absolute temperature during gas sampling (T). Linear regression was applied to concentration versus time data (collected at 0, 15 and 30 min) to calculate the slope. If a sample deviated from the regression line, or if the correlation coefficient of the regression was not significant at the 95% confidence limit, the sample was either recalculated without the outlier or rejected. Finally, the total methane flux for the entire cropping period was calculated using the following equation:
where Ri indicates the methane emission rate and was expressed as grams of methane per square metre per day for each sampling interval represented by the index I, Di represents the number of days within each sampling interval and n is the total number of sampling intervals and was calculated to determine the overall methane emission rate. Continuous monitoring of the soil redox potential (Eh) was conducted at a depth of 20 cm concurrently with methane sampling.
Analysis of the soluble sucrose and nitrogen contents of rice plants
To determine the levels of soluble sucrose and nitrogen in rice plants, ten plants were selected on the basis of their parts, growth stage and nitrogen levels. The plants were collected and analysed on the basis of their fresh weight.
The determination of the soluble sugar and nitrogen content was conducted using the phenol and salicylic acid methods, as described previously22. The starch content of rice plant samples was determined by first extracting the soluble sucrose by heating the samples in perchloric acid to convert starch into soluble sucrose. The soluble sucrose content was determined, which was then multiplied by 0.9 to calculate the starch content. To determine the leaf chlorophyll and carotenoid contents, pigments were extracted using 95% ethanol and measured using a spectrophotometer (Agilent Carry 7000 UMS).
Analysis of the rhizosphere and endosphere microbiota of Milyang360 and Saeilmi by metagenome sequencing
Rhizosphere soil and root samples (1 g) were collected from each of five plants in 2021 and ten plants in 2022 belonging to 24 treatments at tillering, heading and grain-filling stages. Soil and endophytic DNA were extracted from each sample using the FastDNA Spin Kit (MP Bio) following the manufacturer’s protocol. The DNA was then polymerase chain reaction (PCR) amplified using universal primers (Bioneer) for 16S ribosomal RNA genes, which target the bacterial 16S sequence (V3–V4 region, V3-F: TCGTCGGCAGCGTCAGATGTGTATAAGAGACAGCCTACGGGNGGCWGCAG; V4-R: GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAGGACTACHVGGGTATCTAATCC)23. The amplified DNA was purified, quantified, pooled and subjected to emulsion PCR using the Herculase II Fusion DNA Polymerase Nextera XT Index Kit V2 from the Illumina MiSeq (Macrogen). The libraries were purified, quantified and then loaded onto the Illumina sequencer. Raw reads were processed using the CD-HIT-OTU software to filter errors and remove noise and were sorted on the basis of barcode sequences. The raw signals of sequences were obtained using a PicoTiter plate. The sequence reads were validated for both nucleotide quality (average Phred score >20) and read length (>300 bp). The data were visualized using Mothur (version 1.33.0), and a phylogenetic tree was generated using similarity thresholds of >75%, >80%, >85%, >90%, >94% and >97% at the phylum, class, order, family, genus and species levels. The rhizo- and endomicrobiome sequence data have been deposited in the NCBI Sequence Read Archive database under the NCBI BioProjects PRJNA917217 in 2021 (accession numbers SAMN32529381–SAMN32529506) and PRJNA917456 in 2022 (accession numbers SAMN32538821–SAMN32539090).
Quantification of methanogens and methanotrophs in the rhizosphere and endosphere of Milyang360 and Saeilmi by qPCR
Soil DNA/RNA and endophytic DNA/RNA were extracted from soil and root samples (0.5 g), respectively, using the FastDNA Spin Kit for Soil (MP Bio) and RNeasy Kit (Qiagen). The copy number of genes related to methanogens (mcrA) and methanotrophs (pmoA), and type I and type II methanotrophs were quantified by real-time PCR with Quantstudio5 (Thermo Fisher Scientific). Absolute quantification for soil and root DNA/RNA samples was performed by quantitative PCR (qPCR). A 1/10 serial dilution of the initial standard mass particle solution was used to generate a standard template, which was then amplified to determine the threshold cycle (Ct) for a known concentration. The obtained R2 values were greater than 0.99. The primers used for the analysis are listed in Extended Data Table 3.
Root exudate analysis
Root exudates were collected as described previously24 and filtered through a 0.45 μm filter paper. The filtered samples were lyophilized to concentrate the exudates and analysed for organic carbon using a total organic carbon analyser (Analytik Jena). The contents of total organic carbon, carbohydrates and total organic acids were analysed using kits from Cominbio, while the identity of organic acids was determined by high-performance liquid chromatography (Agilent). Ten plants were used for this analysis.
Gene expression analysis
Total RNA was extracted from various plant parts using the RNeasy Kit (Qiagen), according to the manufacturer’s instructions. Then complementary DNA was synthesized using the ProtoScript II First Strand cDNA Synthesis Kit (NEB). Subsequently, qPCR reactions were performed using the amfiSure qGreen qPCR Master Mix (2X) with low ROX (GenDEPOT). Gene expression was analysed by real-time PCR on the Quantstudio5 instrument (Thermo Fisher Scientific), with 40 cycles. Transcript levels of genes were normalized relative to the level of rice Actin1 gene. The primer sequences used to perform real-time PCR are listed in Extended Data Table 3.
Assessment of soil physiochemical characteristics and measurement of plant agronomic traits
Experimental plots maintained for two decades under normal (9 kg per 10 acres) and low (4.5 kg per 10 acres) nitrogen levels in the Department of Southern Area Crop Science, National Institute of Crop Science, Rural Development Administration (35° 29′ 32.2872″ N, 128° 44′ 32.1972″ E) were used for the assessment of soil physiochemical properties: soil pH, electrical conductivity, total nitrogen (N) and extractable phosphorous (P) and potassium (K) as described previously25. Soil pH was measured using a 1/2.5 soil to deionized water suspension. Electrical conductivity was measured in a 1/5 soil to deionized water suspension. Total N was analysed using the Kjeldahl method, and total oxidizable organic carbon was determined using the Walkley and Black wet-oxidation method26. The total oxidizable organic carbon content was multiplied by a factor of 1.72 to calculate organic matter content. Extractable P and K were determined according to the Egner–Riehm extraction procedure using a solution containing 0.1 M ammonium lactate and 0.4 M acetic acid (pH = 3.65–3.75), which is suitable for use in acidic soils.
Plant agronomic traits, including photosynthetic rate, tiller number per plant, spikelet number per panicle, plant height, panicle length, heading date and milled grain yield, were examined. All agronomic traits were measured in triplicate from ten randomly selected plants in each experimental plot; grain yield was measured in triplicate using 100 plants.
Plant photosynthetic rate was quantified using the LI-6800 instrument (LI-COR). Measurements were conducted on different parts of ten plants between 10:00 lt and 13:00 lt under varying levels of nitrogen27,28,29.
Methane emissions calculation formula from 2006 IPCC guideline
The formula for calculating methane emissions from every country paddy type and cultivation management is as follows.
where EFi = EFc × SFw × SFp × SFo × SFs,r, EFijk is the emission factor for rice paddy fields in each category (kg ha−1 d−1 CH4), Tijk is the number of rice cultivation days in rice paddy fields of each category (day), Aijk is the harvest area of rice paddy fields in each category (ha yr−1), EFc is the baseline emission factor (kg ha−1 d−1 CH4), SFw is the scale factor used for rice paddy type and water management during the growing season, SFp is the scale factor used for water management before the growing season, SFo is the scale factor used for application of organic amendments and SFs,r is the scale factor used for soil type and rice cultivar (only when applicable).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The rhizo- and endomicrobiome sequence data have been deposited in the NCBI Sequence Read Archive database under the NCBI BioProjects PRJNA917217 in 2021 (accession numbers SAMN32529381–SAMN32529506) and PRJNA917456 in 2022 (accession numbers: SAMN32538821–SAMN32539090).
References
Bridgham, S. D., Cadillo‐Quiroz, H., Keller, J. K. & Zhuang, Q. Methane emissions from wetlands: biogeochemical, microbial, and modeling perspectives from local to global scales. Glob. Change Biol. 19, 1325–1346 (2013).
Liu, Y. & Whitman, W. B. Metabolic, phylogenetic, and ecological diversity of the methanogenic archaea. Ann. N. Y. Acad. Sci. 1125, 171–189 (2008).
Qiu, J. China cuts methane emissions from rice fields. Nature https://doi.org/10.1038/news.2009.833 (2009).
Su, J. et al. Expression of barley SUSIBA2 transcription factor yields high-starch low-methane rice. Nature 523, 602–606 (2015).
Jiang, Y. et al. Effect of rice panicle size on paddy field CH4 emissions. Biol. Fertil. Soils 52, 389–399 (2016).
Sass, R. L. & Cicerone, R. J. Photosynthate allocations in rice plants: food production or atmospheric methane? Proc. Natl Acad. Sci. USA 99, 11993–11995 (2002).
Chen, Y. et al. OsRGA1 optimizes photosynthate allocation for roots to reduce methane emissions and improve yield in paddy ecosystems. Soil Biol. Biochem. 160, 108344 (2021).
Jiang, Y. et al. Limited potential of harvest index improvement to reduce methane emissions from rice paddies. Glob. Change Biol. 25, 686–698 (2019).
Mao, H. et al. Linking differential domain functions of the GS3 protein to natural variation of grain size in rice. Proc. Natl Acad. Sci. USA 107, 19579–19584 (2010).
Fan, C. et al. GS3, a major QTL for grain length and weight and minor QTL for grain width and thickness in rice, encodes a putative transmembrane protein. Theor. Appl. Genet. 112, 1164–1171 (2006).
Zhang, H. et al. A Gγ protein regulates alkaline sensitivity in crops. Science 379, eade8416 (2023).
Takano-Kai, N. et al. Evolutionary history of GS3, a gene conferring grain length in rice. Genetics 182, 1323–1334 (2009).
Yoon, D. K. et al. The gs3 allele from a large‐grain rice cultivar, Akita 63, increases yield and improves nitrogen‐use efficiency. Plant Direct 6, e417 (2022).
Evans, P. N. et al. An evolving view of methane metabolism in the Archaea. Nat. Rev. Microbiol. 17, 219–232 (2019).
Wu, Y. et al. Rice transcription factor OsDOF11 modulates sugar transport by promoting expression of sucrose transporter and SWEET genes. Mol. plant 11, 833–845 (2018).
Hirose, T., Ohdan, T., Nakamura, Y. & Terao, T. Expression profiling of genes related to starch synthesis in rice leaf sheaths during the heading period. Physiol. Plant. 128, 425–435 (2006).
Banger, K., Tian, H. & Lu, C. Do nitrogen fertilizers stimulate or inhibit methane emissions from rice fields? Glob. Change Biol. 18, 3259–3267 (2012).
Kim, G. W., Gutierrez-Suson, J. & Kim, P. J. Optimum N rate for grain yield coincides with minimum greenhouse gas intensity in flooded rice fields. Field Crops Res. 237, 23–31 (2019).
Kang, J.-W. et al. Accelerated development of rice stripe virus-resistant, near-isogenic rice lines through marker-assisted backcrossing. PLoS ONE 14, e0225974 (2019).
Rolston, D. Gas diffusivity. Methods Soil Anal. Part 1 5, 1089–1102 (1986).
Gutierrez, J., Atulba, S. L., Kim, G. & Kim, P. J. Importance of rice root oxidation potential as a regulator of CH4 production under waterlogged conditions. Biol. Fertil. Soils 50, 861–868 (2014).
Wang, X. & Huang, J. Principles and Techniques of Plant Physiological Biochemical Experiment (Higher Education Pres, 2006).
Klindworth, A. et al. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. https://doi.org/10.1093/nar/gks808 (2013).
Chen, Y. et al. Rice root morphological and physiological traits interaction with rhizosphere soil and its effect on methane emissions in paddy fields. Soil Biol. Biochem. 129, 191–200 (2019).
Alvarenga, P., Mourinha, C., Palma, P., Cruz, N. & Rodrigues, S. M. Assessment of soil physicochemical characteristics and As, Cu, Pb and Zn contamination in non-active mines at the Portuguese sector of the Iberian Pyrite Belt. Environments 9, 105 (2022).
Walkley, A. & Black, I. A. An examination of the Degtjareff method for determining soil organic matter, and a proposed modification of the chromic acid titration method. Soil Sci. 37, 29–38 (1934).
IPCC Guidelines for National Greenhouse Gas Inventories (eds Eggleston, H. S., Buendia, L., Miwa, K., Ngara, T. & Tanabe, K.) (IGES, 2006).
Holmes, A. J., Costello, A., Lidstrom, M. E. & Murrell, J. C. Evidence that participate methane monooxygenase and ammonia monooxygenase may be evolutionarily related. FEMS Microbiol. Lett. 132, 203–208 (1995).
Costello, A. M. & Lidstrom, M. E. Molecular characterization of functional and phylogenetic genes from natural populations of methanotrophs in lake sediments. Appl. Environ. Microbiol. 65, 5066–5074 (1999).
Acknowledgements
This work was carried out with the support of Cooperative Research Program for Agriculture Science and Technology Development Project no. PJ01700801 (RS-2022-RD010405) and PJ01699002 (RS-2022-RD010288) Rural Development Administration, Republic of Korea, and the KRIBB Initiative Program (KGM2112335).
Author information
Authors and Affiliations
Contributions
Y.K. wrote the manuscript and performed most experiments. J.-Y.L., S.-M.L., J.-K.C., H.P., J.-W.K., N.R.K. and K.-W.O evaluated agronomic traits. J.C., H.G.C., T.H.K. and P.J.K. measured methane emissions. D.K. and Y.-S.K. analysed the microbial community. J.-H.L. and C.-M.R. wrote and managed the manuscript and correspondence.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Climate Change thanks Paul Bodelier and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Breeding strategy of near isogenic line; Milyang360.
Saeilmi and Shindongjin are 96% genetically similar. Chip array analysis revealed that Milyang360 is 99.9% genetically similar to Saeilmi. MABC indicates the marker-assisted backcross.
Extended Data Fig. 2 Plant agronomic traits of Milyang360 and Saeilmi.
a, Marker-assisted backcross of Milyang360 using the KASP marker for the identification of plants harboring the gs3 allele. b, Plant phenotypes under different nitrogen levels. c, Agronomic traits of Milyang360 and Saeilmi plants under different nitrogen levels. Ten plants were used for the investigation of agronomic traits. The agronomic traits were analyzed using Kruskal-Wallis test (p < 0.05). Letters represent significant differences between gs3 alleles under different nitrogen levels (p < 0.05; Tukey’s HSD test. The data are shown as means ± SD and n = 10.
Extended Data Fig. 3 Abundances of methanotroph on rhizosphere and endosphere.
a, Microbiota and beta-diversity of rhizosphere of gs3 alleles under different nitrogen levels. b, Quantification of methanogen abundance and mcrA gene copy number in the rhizosphere of the two gs3 alleles under different nitrogen levels. Kruskal-Wallis test was used to determine significant differences (p < 0.05) in mcrA copy number. Letters represent significant differences between gs3 alleles under different nitrogen levels (p < 0.05; Tukey’s HSD test). The data are shown minima is the lowest data point, maxima is the highest data point, median is the middle value in the data set, first quartile is median of the lower half of the dataset, third quartile is the median of the upper half of the dataset and n = 10. c, Heatmap showing the abundance of endophyte methanotroph on different growth stages. Relative abundance data were transformed to centered-log ratio (CLR), which involves dividing each element in the count vector by the geometric mean of all elements and then taking the natural logarithm of each ratio. This transformation enables the conversion of data from the simplex space to real co-ordinate space. d, Quantification of methanotrophs in the rhizosphere and endosphere, based on pmoA gene copy number. The relative abundance of endosphere microbiota was analyzed. Significant differences in pmoA gene copy number were analyzed using Kruskal-Wallis test (p < 0.05). Letters represent significant differences between gs3 alleles under different nitrogen levels (p < 0.05; Tukey’s HSD test. The data are shown minima is the lowest data point, maxima is the highest data point, median is the middle value in the data set, first quartile is median of the lower half of the dataset, third quartile is the median of the upper half of the dataset and n = 10.
Extended Data Fig. 4 Root exudate contents of Milyang360 and Saeilmi.
Root exudates of Milyang360 and Saeilmi plants at different growth stages and under different nitrogen levels. Significant differences in root exudates were analyzed using Kruskal-Wallis test (p < 0.05). Letters represent significant differences between gs3 alleles under different nitrogen levels (p < 0.05; Tukey’s HSD test). At the tillering stage, p-value of dissolved organic carbon, carbohydrate, total organic acid, acetic acid, oxalic acid and maleic acid were 0.045, 0.041, 0.038, 0.045, 0.039 and 0.040 respectively. At the heading stage, p-value were 0.041, 0.032, 0.035, 0.035, 0.030 and 0.027. At the grain filling stage, p-value were 0.038, 0.032, 0.025, 0.034, 0.033 and 0.027. The data are shown as means ± SD and n = 10.
Supplementary information
Supplementary Information
Supplementary Fig. 1 and Tables 1–3.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kwon, Y., Lee, JY., Choi, J. et al. Loss-of-function gs3 allele decreases methane emissions and increases grain yield in rice. Nat. Clim. Chang. 13, 1329–1333 (2023). https://doi.org/10.1038/s41558-023-01872-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41558-023-01872-5