Abstract
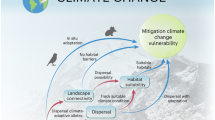
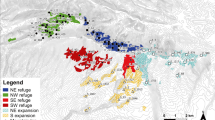
Accounting for population-level adaptation and migration remains a central challenge to predicting climate change effects on biodiversity. Assessing how climate change could disrupt local climate adaptation, resulting in maladaptation and possibly extirpation, can inform where climate change poses the greatest risks across species ranges. For the forest tree species balsam poplar (Populus balsamifera), we used climate-associated genetic loci to predict population maladaptation with and without migration, the distance to sites that minimize maladaptation, and the emergence of novel genotype–climate associations. We show that the greatest disruptions to contemporary genotype–climate associations occur along the longitudinal edges of the range, where populations are predicted to be maladapted to all future North American climates, rescue via migration is most limited and novel genotype–climate associations emerge. Our work advances beyond species-level range modelling towards the long-held goal of simultaneously estimating the contributions of maladaptation and migration to understanding the risks that populations may face from shifting climates.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data are publicly available. The allele frequencies are available in ref. 14, and the climate data are available at https://www.worldclim.org.
Code availability
The R code and genetic data to calculate local, forward and reverse offsets are available at github.com/agougher/poplarAdaptiveOffset.
References
Aitken, S. N., Yeaman, S., Holliday, J. A., Wang, T. & Curtis-McLane, S. Adaptation, migration or extirpation: climate change outcomes for tree populations. Evol. Appl. 1, 95–111 (2008).
Davis, M. B. & Shaw, R. G. Range shifts and adaptive responses to Quaternary climate change. Science 292, 673–679 (2001).
Ikeda, D. H. et al. Genetically informed ecological niche models improve climate change predictions. Glob. Change Biol. 23, 164–176 (2017).
Maguire, K. C., Shinneman, D. J., Potter, K. M. & Hipkins, V. D. Intraspecific niche models for ponderosa pine (Pinus ponderosa) suggest potential variability in population-level response to climate change. Syst. Biol. https://doi.org/10.1093/sysbio/syy017 (2018).
Smith, A. B., Godsoe, W., Rodríguez-Sánchez, F., Wang, H.-H. & Warren, D. Niche estimation above and below the species level. Trends Ecol. Evol. 34, 260–273 (2019).
Razgour, O. et al. Considering adaptive genetic variation in climate change vulnerability assessment reduces species range loss projections. Proc. Natl Acad. Sci. USA 116, 10418–10423 (2019).
Wang, T., O’Neill, G. A. & Aitken, S. N. Integrating environmental and genetic effects to predict responses of tree populations to climate. Ecol. Appl. 20, 153–163 (2010).
Radeloff, V. C. et al. The rise of novelty in ecosystems. Ecol. Appl. 25, 2051–2068 (2015).
Williams, J. W. & Jackson, S. T. Novel climates, no-analog communities, and ecological surprises. Front. Ecol. Environ. 5, 475–482 (2007).
Aitken, S. N. & Whitlock, M. C. Assisted gene flow to facilitate local adaptation to climate change. Annu. Rev. Ecol. Evol. Syst. 44, 367–388 (2013).
Vitt, P., Havens, K., Kramer, A. T., Sollenberger, D. & Yates, E. Assisted migration of plants: changes in latitudes, changes in attitudes. Biol. Conserv. 143, 18–27 (2010).
Williams, M. I. & Dumroese, R. K. Preparing for climate change: forestry and assisted migration. J. For. 111, 287–297 (2013).
Keller, S. R., Levsen, N., Olson, M. S. & Tiffin, P. Local adaptation in the flowering-time gene network of balsam poplar, Populus balsamifera L. Mol. Biol. Evol. 29, 3143–3152 (2012).
Keller, S. R., Chhatre, V. E. & Fitzpatrick, M. C. Influence of range position on locally adaptive gene–environment associations in Populus flowering time genes. J. Hered. 109, 47–58 (2018).
Chuine, I. Why does phenology drive species distribution? Phil. Trans. R. Soc. B 365, 3149–3160 (2010).
Morin, X., Viner, D. & Chuine, I. Tree species range shifts at a continental scale: new predictive insights from a process-based model. J. Ecol. 96, 784–794 (2008).
Ferrier, S., Manion, G., Elith, J. & Richardson, K. Using generalized dissimilarity modelling to analyse and predict patterns of beta diversity in regional biodiversity assessment. Divers. Distrib. 13, 252–264 (2007).
Ellis, N., Smith, S. J. & Pitcher, C. R. Gradient forests: calculating importance gradients on physical predictors. Ecology 93, 156–168 (2012).
Fitzpatrick, M. C. & Keller, S. R. Ecological genomics meets community-level modelling of biodiversity: mapping the genomic landscape of current and future environmental adaptation. Ecol. Lett. 18, 1–16 (2015).
Fei, S. et al. Divergence of species responses to climate change. Sci. Adv. 3, e1603055 (2017).
VanDerWal, J. et al. Focus on poleward shifts in species’ distribution underestimates the fingerprint of climate change. Nat. Clim. Change 3, 239–243 (2013).
Shaw, R. G. From the past to the future: considering the value and limits of evolutionary prediction. Am. Nat. 193, 1–10 (2018).
Hampe, A. & Petit, R. J. Conserving biodiversity under climate change: the rear edge matters. Ecol. Lett. 8, 461–467 (2005).
Yun, J. et al. Influence of winter precipitation on spring phenology in boreal forests. Glob. Change Biol. 24, 5176–5187 (2018).
Fu, Y. H. et al. Unexpected role of winter precipitation in determining heat requirement for spring vegetation green-up at northern middle and high latitudes. Glob. Change Biol. 20, 3743–3755 (2014).
Peterson, M. L., Doak, D. F. & Morris, W. F. Incorporating local adaptation into forecasts of species’ distribution and abundance under climate change. Glob. Change Biol. 25, 775–793 (2019).
Atkins, K. E. & Travis, J. M. J. Local adaptation and the evolution of species’ ranges under climate change. J. Theor. Biol. 266, 449–457 (2010).
Chen, I.-C., Hill, J. K., Ohlemuller, R., Roy, D. B. & Thomas, C. D. Rapid range shifts of species associated with high levels of climate warming. Science 333, 1024–1026 (2011).
Groom, Q. J. Some poleward movement of British native vascular plants is occurring, but the fingerprint of climate change is not evident. PeerJ 1, e77 (2013).
Olson, M. S. et al. The adaptive potential of Populus balsamifera L. to phenology requirements in a warmer global climate. Mol. Ecol. 22, 1214–1230 (2013).
Fitzpatrick, M., Chhatre, V., Soolanayakanahally, R. & Keller, S. Experimental support for genomic prediction of climate maladaptation using the machine learning approach Gradient Forests. Preprint at https://doi.org/10.22541/au.159863198.86187354 (2020).
Blois, J. L., Zarnetske, P. L., Fitzpatrick, M. C. & Finnegan, S. Climate change and the past, present, and future of biotic interactions. Science 341, 499–504 (2013).
Keller, S. R. et al. Climate-driven local adaptation of ecophysiology and phenology in balsam poplar, Populus balsamifera L. (Salicaceae). Am. J. Bot. 98, 99–108 (2011).
Alberto, F. J. et al. Potential for evolutionary responses to climate change—evidence from tree populations. Glob. Change Biol. 19, 1645–1661 (2013).
Little, E. L. Atlas of United States Trees (US Dept of Agriculture, Forest Service, 1971).
Romero-Lankao, P. et al. in Climate Change 2014: Impacts, Adaptation, and Vulnerability. Part B: Regional Aspects (eds Barros, V. R. et al.) 1439–1498 (Cambridge Univ. Press, 2014).
Fetter, K. C., Gugger, P. F. & Keller, S. R. in Comparative and Evolutionary Genomics of Angiosperm Trees (eds Groover, A. & Cronk, Q.) 303–333 (Springer International, 2017); https://doi.org/10.1007/7397_2016_19
Soolanayakanahally, R. Y., Guy, R. D., Silim, S. N., Drewes, E. C. & Schroeder, W. R. Enhanced assimilation rate and water use efficiency with latitude through increased photosynthetic capacity and internal conductance in balsam poplar (Populus balsamifera L.). Plant Cell Environ. 32, 1821–1832 (2009).
Chhatre, V. E. et al. Climatic niche predicts the landscape structure of locally adaptive standing genetic variation. Preprint at https://doi.org/10.1101/817411 (2019).
Günther, T. & Coop, G. Robust identification of local adaptation from allele frequencies. Genetics 195, 205–220 (2013).
Frichot, E., Schoville, S. D., Bouchard, G. & François, O. Testing for associations between loci and environmental gradients using latent factor mixed models. Mol. Biol. Evol. 30, 1687–1699 (2013).
Hijmans, R. J., Cameron, S. E., Parra, J. L., Jones, P. G. & Jarvis, A. Very high resolution interpolated climate surfaces for global land areas. Int. J. Climatol. 25, 1965–1978 (2005).
Frichot, E. & François, O. LEA: an R package for landscape and ecological association studies. Methods Ecol. Evol. 6, 925–929 (2015).
Goudet, J. & Jombart, T. hierfstat: Estimation and tests of hierarchical F-statistics. R package version 0.04-22 (2015).
Manion, G., Lisk, M., Nieto-Lugilde, D., Mokany, K. & Fitzpatrick, M. gdm: Generalized dissimilarity modeling. R package version 1.3.11 (2018).
Hijmans, R. J. geosphere: Spherical trigonometry. R package version 1.5-10 (2019).
Gougherty, A. V., Chhatre, V. E., Keller, S. R. & Fitzpatrick, M. C. Contemporary range position predicts the range-wide pattern of genetic diversity in balsam poplar (Populus balsamifera L.). J. Biogeogr. 47, 1246–1257 (2020).
Vallejos, R., Osorio, F. & Bevilacqua, M. Spatial Relationships Between Two Georeferenced Variables: With Applications in R (Springer, 2018).
Acknowledgements
We thank V. E. Chhatre for providing comments on an earlier draft of this manuscript. This work was supported by National Science Foundation Plant Genome Research award no. 1461868 to S.R.K. and M.C.F., and an UMCES PhD fellowship to A.V.G.
Author information
Authors and Affiliations
Contributions
A.V.G. and M.C.F. conceived the study. S.R.K. processed and provided the genetic information. A.V.G. analysed the data and led the writing, with contributions and discussion from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Climate Change thanks Erin Landguth, Christian Rellstab and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Sampled population locations in geographic and climatic space.
a, Geographic locations of populations used in GDM and GF models (blue circles), within balsam poplar’s range (white polygon). b, Position of populations (blue circles), and cells in balsam poplar’s range (black dots) in current North American climate space (gray circles). Red circles show the composite future climate (RCP 8.5) of North America used in predictions. Climate space is shown as the first two principal components (PCs) of current North American climate (mean diurnal range, isothermality, mean summer temperature, mean winter temperature, summer precipitation, winter precipitation), with future climates predicted into the same PCA space. b, is shown only for visualization purposes.
Extended Data Fig. 2 Red-green-blue map of local (red), forward (green), and reverse (blue) offsets.
Offset values were calculated from Gradient Forest throughout the range of balsam poplar for 2070 and RCP 8.5. Brighter cells, closer to white, have relatively high values along each of the three axes while darker cells, closer to black, have relatively lower values. b-d, Bivariate scattergrams of (a), with 1:1 lines. Individual maps used in (a) are shown in Extended Data Fig. 4.
Extended Data Fig. 3 Local, forward, and reverse offsets from generalized dissimilarity models for balsam poplar.
a & b, Local genetic offset, (c & d) forward offset, and (e & f) reverse offset from a generalized dissimilarity model for RCP 4.5 (first column; a, c, e) and RCP 8.5 (second column; b, d, f) for 2070. Note the non-linear color scale.
Extended Data Fig. 4 Local, forward, and reverse offsets from Gradient Forest for balsam poplar.
a & b, Local genetic offset, (c & d) forward offset, and (e & f) reverse offset from a Gradient Forest model for RCP 4.5 (first column; a, c, e) and RCP 8.5 (second column; b, d, f) for 2070.
Extended Data Fig. 5 Distance and initial bearing to locations that minimizes forward offset.
Distance and (b) initial bearing were calculated from the focal cell to the location in future North American climate (2070, RCP 8.5) that minimizes predicted offset from a Gradient Forest model. Polar histogram in (b) shows the log10 number of cells in each bearing bin.
Supplementary information
Supplementary Information
Supplementary Figs. 1–12 and Tables 1 and 2.
Rights and permissions
About this article
Cite this article
Gougherty, A.V., Keller, S.R. & Fitzpatrick, M.C. Maladaptation, migration and extirpation fuel climate change risk in a forest tree species. Nat. Clim. Chang. 11, 166–171 (2021). https://doi.org/10.1038/s41558-020-00968-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41558-020-00968-6
This article is cited by
-
Genomic insights into local adaptation and vulnerability of Quercus longinux to climate change
BMC Plant Biology (2024)
-
Genomic signals of local adaptation in Picea crassifolia
BMC Plant Biology (2023)
-
Natural hybridization reduces vulnerability to climate change
Nature Climate Change (2023)
-
Climate-induced range shifts drive adaptive response via spatio-temporal sieving of alleles
Nature Communications (2023)
-
Local adaptation and future climate vulnerability in a wild rodent
Nature Communications (2023)