Abstract
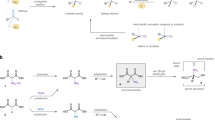
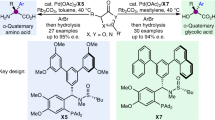
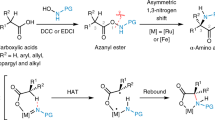
Stereoselective protonation is a challenge in asymmetric catalysis. The small size and high rate of transfer of protons mean that face-selective delivery to planar intermediates is hard to control, but it can unlock previously obscure asymmetric transformations. Particularly, when coupled with a preceding decarboxylation, enantioselective protonation can convert the abundant acid feedstocks into structurally diverse chiral molecules. Here an anchoring group strategy is demonstrated as a potential alternative and supplement to the conventional structural modification of catalysts by creating additional catalyst–substrate interactions. We show that a tailored benzamide group in aminomalonic acids can help build a coordinated network of non-covalent interactions, including hydrogen bonds, π–π interactions and dispersion forces, with a chiral acid catalyst. This allows enantioselective decarboxylative protonation to give α-amino acids. The malonate-based synthesis introduces side chains via a facile substitution of aminomalonic esters and thus can access structurally and functionally diverse amino acids.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data supporting the findings of this study are available within the paper and its Supplementary Information. Crystallographic data for compound (+)-6 have been deposited at the Cambridge Crystallographic Data Centre (CCDC) under deposition no. CCDC 2206851. These data can be obtained free of charge from the CCDC (http://www.ccdc.cam.ac.uk/data_request/cif). Source data are provided with this paper.
References
Fehr, C. Enantioselective protonation of enolates and enols. Angew. Chem. Int. Ed. 35, 2566–2587 (1996).
Duhamel, L., Duhamel, P. & Plaquevent, J. ‑C. Enantioselective protonations: fundamental insights and new concepts. Tetrahedron 15, 3653–3691 (2004).
Mohr, J. T., Hong, A. Y. & Stoltz, B. M. Enantioselective protonation. Nat. Chem. 1, 359–369 (2009).
Blanchet, J., Baudoux, J., Amere, M., Lasne, M.-C. & Rouden, J. Asymmetric malonic and acetoacetic acid syntheses – a century of enantioselective decarboxylative protonations. Eur. J. Org. Chem. 5493–5506 (2008).
Mohr, J. T., Nishimata, T., Behenna, D. C. & Stoltz, B. M. Catalytic enantioselective decarboxylative protonation. J. Am. Chem. Soc. 128, 11348–11349 (2006).
Marinescu, S. C., Nishimata, T., Mohr, J. T. & Stoltz, B. M. Homogeneous Pd-catalyzed enantioselective decarboxylative protonation. Org. Lett. 10, 1039–1042 (2008).
Morita, M. et al. Two methods for catalytic generation of reactive enolates promoted by a chiral poly Gd complex: application to catalytic enantioselective protonation reactions. J. Am. Chem. Soc. 131, 3858–3859 (2009).
Mitsuhashi, K., Ito, R., Arai, T. & Yanagisawa, A. Catalytic asymmetric protonation of lithium enolates using amino acid derivatives as chiral proton sources. Org. Lett. 8, 1721–1724 (2006).
Poisson, T., Oudeyer, S., Dalla, V., Marsais, F. & Levacher, V. Straightforward organocatalytic enantioselective protonation of silyl enolates by means of Cinchona alkaloids and carbxylic acids. Synlett 2447–2450 (2008).
Leow, D., Lin, S., Chittimalla, S. K., Fu, X. & Tan, C. ‑H. Enantioselective protonation catalyzed by a chiral bicyclic guanidine derivative. Angew. Chem. Int. Ed. 47, 5641–5645 (2008).
Dai, X., Nakai, T., Romero, J. A. C. & Fu, G. C. Enantioselective synthesis of protected amines by the catalytic asymmetric addition of hydrazoic acid to ketenes. Angew. Chem. Int. Ed. 46, 4367–4369 (2007).
Miyamoto, K. & Kourist, R. Arylmalonate decarboxylase—a highly selective bacterial biocatalyst with unknown function. Appl. Microbiol. Biotechnol. 100, 8621–8631 (2016).
Wilding, M., Goodall, M. & Micklefield, J. in Comprehensive Chirality 1st edn (eds H. Yamamoto & E. M. Carreira) 402–409 (Elsevier, 2012).
Rueping, M., Kuenkel, A. & Atodiresei, I. Chiral Brønsted acids in enantioselective carbonyl activations − activation modes and applications. Chem. Soc. Rev. 40, 4539–4549 (2011).
Min, C. & Seidel, D. Asymmetric Brønsted acid catalysis with chiral carboxylic acids. Chem. Soc. Rev. 46, 5889–5902 (2017).
Akiyama, T. & Mori, K. Stronger Brønsted acids: recent progress. Chem. Rev. 115, 9277–9306 (2015).
Maji, R., Mallojjala, S. C. & Wheeler, S. E. Chiral phosphoric acid catalysis: from numbers to insights. Chem. Soc. Rev. 47, 1142–1158 (2018).
Knowles, R. R. & Jacobsen, E. N. Attractive noncovalent interactions in asymmetric catalysis: Links between enzymes and small molecule catalysts. Proc. Natl Acad. Sci. USA 107, 20678–20685 (2010).
Neel, A. J., Hilton, M. J., Sigman, M. S. & Toste, F. D. Exploiting non-covalent π interactions for catalyst design. Nature 543, 637–646 (2017).
Huang, D., Xu, F., Lin, X. & Wang, Y. Highly enantioselective Pictet–Spengler reaction catalyzed by SPINOL phosphoric acids. Chem. Eur. J. 18, 3148–3152 (2012).
Okrasa, K. et al. Structure-guided directed evolution of alkenyl and arylmalonate decarboxylases. Angew. Chem. Int. Ed. 48, 7691–7694 (2009).
Brunner, H. & Baur, M. A. α-Amino acid derivatives by enantioselective decarboxylation. Eur. J. Org. Chem. 2854–2862 (2010).
Lin, X., Wang, L., Han, Z. & Chen, Z. Chiral spirocyclic phosphoric acids and their growing applications. Chin. J. Chem. 39, 802–824 (2021).
Xu, F. et al. SPINOL-derived phosphoric acids: synthesis and application in enantioselective Friedel–Crafts reaction of indoles with imines. J. Org. Chem. 75, 8677–8680 (2018).
Rahman, A. & Lin, X. Development and application of chiral spirocyclic phosphoric acids in asymmetric catalysis. Org. Biomol. Chem. 16, 4753–4777 (2018).
Messina, M. S. & Maynard, H. D. Modification of proteins using olefin metathesis. Mater. Chem. Front. 4, 1040–1051 (2020).
Parker, C. G. & Pratt, M. R. Click chemistry in proteomic investigations. Cell 180, 605–632 (2020).
Kim, C. H., Axup, J. Y. & Schultz, P. G. Protein conjugation with genetically encoded unnatural amino acids. Curr. Opin. Chem. Biol. 17, 412–419 (2013).
Arntson, K. E. & Pomerantz, W. C. K. Protein-observed fluorine NMR: a bioorthogonal approach for small molecule discovery. J. Med. Chem. 59, 5158–5171 (2016).
Trimble, J. S. et al. A designed photoenzyme for enantioselective [2+2] cycloadditions. Nature 611, 709–714 (2022).
Sun, N. et al. Enantioselective [2+2]-cycloadditions with triplet photoenzymes. Nature 611, 715–720 (2022).
Cheng, Z., Kuru, E., Sachdeva, A. & Vendrell, A. Fluorescent amino acids as versatile building blocks for chemical biology. Nat. Rev. Chem. 4, 275–290 (2020).
Fujita, T. et al. Synthesis of a complete set of l-difluorophenylalanines, l-(F2)Phe, as molecular explorers for the CH/π interaction between peptide ligand and receptor. Tetrahedron Lett. 41, 923–927 (2000).
Mann, M. Functional and quantitative proteomics using SILAC. Nat. Rev. Mol. Cell Biol. 7, 952–958 (2006).
Castellani, F. et al. Structure of a protein determined by solid-state magic-angle-spinning NMR spectroscopy. Nature 420, 99–102 (2002).
Shaginian, A. et al. Design, synthesis, and evaluation of an α-helix mimetic library targeting protein–protein interactions. J. Am. Chem. Soc. 131, 5564–5572 (2009).
Durcik, M. et al. New N-phenylpyrrolamide DNA gyrase B inhibitors: optimization of efficacy and antibacterial activity. Eur. J. Med. Chem. 154, 117–132 (2018).
Walsh, P. J. & Kozlowski, M. C. in Fundamentals of Asymmetric Catalysis 330–374 (University Science Books, 2009).
Lu, T. & Chen, Q. Independent gradient model based on Hirshfeld partition: A new method for visual study of interactions in chemical systems. J. Comput. Chem. 43, 539–555 (2022).
Horn, P. R., Mao, Y. & Head-Gordon, M. Defining the contributions of permanent electrostatics, Pauli repulsion, and dispersion in density functional theory calculations of intermolecular interaction energies. J. Chem. Phys. 144, 114107 (2016).
Horn, P. R. & Head-Gordon, M. Polarization contributions to intermolecular interactions revisited with fragment electric-field response functions. J. Chem. Phys. 143, 114111 (2015).
Horn, P. R., Mao, Y. & Head-Gordon, M. Probing non-covalent interactions with a second generation energy decomposition analysis using absolutely localized molecular orbitals. Phys. Chem. Chem. Phys. 18, 23067–23079 (2016).
Qi, X., Kohler, D. G., Hull, K. L. & Liu, P. Energy decomposition analyses reveal the origins of catalyst and nucleophile effects on regioselectivity in nucleopalladation of alkenes. J. Am. Chem. Soc. 141, 11892–11904 (2019).
Xi, Y. et al. Application of trimethylgermanyl-substituted bisphosphine ligands with enhanced dispersion interactions to copper-catalyzed hydroboration of disubstituted alkenes. J. Am. Chem. Soc. 142, 18213–18222 (2020).
Mao, Y. et al. Consistent inclusion of continuum solvation in energy decomposition analysis: theory and application to molecular CO2 reduction catalysts. Chem. Sci. 12, 1398–1414 (2021).
Sengupta, A., Li, B., Svatunek, D., Liu, F. & Houk, K. N. Cycloaddition reactivities analyzed by energy decomposition analyses and the frontier molecular orbital model. Acc. Chem. Res. 55, 2467–2479 (2022).
Emamian, S., Lu, T., Kruse, H. & Emamian, H. Exploring nature and predicting strength of hydrogen bonds: a correlation analysis between atoms‐in‐molecules descriptors, binding energies, and energy components of symmetry‐adapted perturbation theory. J. Comput. Chem. 40, 2868–2881 (2019).
Tsuzuki, S., Honda, K., Uchimaru, T., Mikami, M. & Tanabe, K. Origin of attraction and directionality of the π/π interaction: model chemistry calculations of benzene dimer interaction. J. Am. Chem. Soc. 124, 104–112 (2002).
Carter-Fenk, K. & Herbert, J. M. Electrostatics does not dictate the slip-stacked arrangement of aromatic π–π interactions. Chem. Sci. 11, 6758–6765 (2020).
Acknowledgements
We thank the University of Hong Kong for a start-up fund, the National Natural Science Foundation of China (22201222, X.Q.), the National Key R&D Program of China (2022YFA1505100, X.Q.), the Fundamental Research Funds for the Central Universities (2042022kf1038, X.Q.) and the Research Grants Council of Hong Kong (17304523, Z.H.). We acknowledge funding support from the Laboratory for Synthetic Chemistry and Chemical Biology under the Health@InnoHK Program launched by the Innovation and Technology Commission, the Government of HKSAR. J. Yip and B. Yan are acknowledged for mass spectrometry and NMR spectroscopy, respectively. X.Q. acknowledges the supercomputing system in the Supercomputing Center of Wuhan University.
Author information
Authors and Affiliations
Contributions
Z.H. conceived and designed the project. W.-F.Z. carried out the experiments. W.-F.Z. and Z.H. analysed the experimental data. J.C. and X.Q. carried out the theoretical studies and analysed the data. All authors wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemistry thanks Tian Lu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–19, experimental procedures, product characterization data, and mechanistic studies.
Supplementary Data 1
Crystallographic data for compound (+)-6; CCDC reference 2206851.
Source data
Source Data Fig. 5a and d
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zheng, WF., Chen, J., Qi, X. et al. Modular and diverse synthesis of amino acids via asymmetric decarboxylative protonation of aminomalonic acids. Nat. Chem. 15, 1672–1682 (2023). https://doi.org/10.1038/s41557-023-01362-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-023-01362-3
This article is cited by
-
An acid for an acid
Nature Chemistry (2023)