Abstract
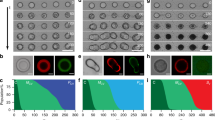
Bottom-up assembly of higher-order cytomimetic systems capable of coordinated physical behaviours, collective chemical signalling and spatially integrated processing is a key challenge in the study of artificial multicellularity. Here we develop an interactive binary population of coacervate microdroplets that spontaneously self-sort into chain-like protocell networks with an alternating sequence of structurally and compositionally dissimilar microdomains with hemispherical contact points. The protocell superstructures exhibit macromolecular self-sorting, spatially localized enzyme/ribozyme biocatalysis and interdroplet molecular translocation. They are capable of topographical reconfiguration using chemical or light-mediated stimuli and can be used as a micro-extraction system for macroscale biomolecular sorting. Our methodology opens a pathway towards the self-assembly of multicomponent protocell networks based on selective processes of coacervate droplet–droplet adhesion and fusion, and provides a step towards the spontaneous orchestration of protocell models into artificial tissues and colonies with ordered architectures and collective functions.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data supporting the findings of this study are available within the article and its Supplementary Information, and from the corresponding authors on reasonable request. Source Data are provided with this paper.
References
Buddingh, B. C., Elzinga, J. & van Hest, J. C. M. Intercellular communication between artificial cells by allosteric amplification of a molecular signal. Nat. Commun. 11, 1652 (2020).
Booth, M. J., Schild, V. R., Graham, A. D., Olof, S. N. & Bayley, H. Light-activated communication in synthetic tissues. Sci. Adv. 2, e1600056 (2016).
Tian, L., Li, M., Patil, A. J., Drinkwater, B. W. & Mann, S. Artificial morphogen-mediated differentiation in synthetic protocells. Nat. Commun. 10, 3321 (2019).
Gao, N. et al. Chemical-mediated translocation in protocell-based microactuators. Nat. Chem. 13, 868–879 (2021).
Adamala, K. P., Engelhart, A. E. & Szostak, J. W. Collaboration between primitive cell membranes and soluble catalysts. Nat. Commun. 7, 11041 (2016).
Bhattacharya, A., Cho, C. J., Brea, R. J. & Devaraj, N. K. Expression of fatty acyl-CoA ligase drives one-pot de novo synthesis of membrane-bound vesicles in a cell-free transcription–translation system. J. Am. Chem. Soc. 143, 11235–11242 (2021).
Saha, R., Verbanic, S. & Chen, I. A. Lipid vesicles chaperone an encapsulated RNA aptamer. Nat. Commun. 9, 2313 (2018).
Marguet, M., Bonduelle, C. & Lecommandoux, S. Multicompartmentalized polymeric systems: towards biomimetic cellular structure and function. Chem. Soc. Rev. 42, 512–529 (2013).
Peters, R. J., Louzao, I. & van Hest, J. C. M. From polymeric nanoreactors to artificial organelles. Chem. Sci. 3, 335–342 (2012).
Huang, X. et al. Interfacial assembly of protein–polymer nano-conjugates into stimulus-responsive biomimetic protocells. Nat. Commun. 4, 2239 (2013).
Li, M., Harbron, R. L., Weaver, J. V., Binks, B. P. & Mann, S. Electrostatically gated membrane permeability in inorganic protocells. Nat. Chem. 5, 529–536 (2013).
Torre, P., Keating, C. D. & Mansy, S. S. Multiphase water-in-oil emulsion droplets for cell-free transcription-translation. Langmuir 30, 5695–5699 (2014).
Wu, H., Du, X., Meng, X., Qiu, D. & Qiao, Y. A three-tiered colloidosomal microreactor for continuous flow catalysis. Nat. Commun. 12, 6113 (2021).
Mukwaya, V. et al. Lectin-glycan-mediated nanoparticle docking as a step toward programmable membrane catalysis and adhesion in synthetic protocells. ACS Nano. 14, 7899–7910 (2020).
Mukwaya, V. et al. Programmable membrane-mediated attachment of synthetic virus-like nanoparticles on artificial protocells for enhanced immunogenicity. Cell Rep. Phys. Sci. 2, 100291 (2021).
Koga, S., Williams, D. S., Perriman, A. W. & Mann, S. Peptide–nucleotide microdroplets as a step towards a membrane-free protocell model. Nat. Chem. 3, 720–724 (2011).
Yewdall, N. A., André, A. A., Lu, T. & Spruijt, E. Coacervates as models of membraneless organelles. Curr. Opin. Colloid Interface Sci. 52, 101416 (2021).
Liu, S. et al. Enzyme-mediated nitric oxide production in vasoactive erythrocyte membrane-enclosed coacervate protocells. Nat. Chem. 12, 1165–1173 (2020).
Pir Cakmak, F., Marianelli, A. M. & Keating, C. D. Phospholipid membrane formation templated by coacervate droplets. Langmuir 37, 10366–10375 (2021).
Gao, N., Xu, C., Yin, Z., Li, M. & Mann, S. Triggerable protocell capture in nanoparticle-caged coacervate microdroplets. J. Am. Chem. Soc. 144, 3855–3862 (2022).
Zhang, Y. et al. Giant coacervate vesicles as an integrated approach to cytomimetic modeling. J. Am. Chem. Soc. 143, 2866–2874 (2021).
Weitz, M. et al. Diversity in the dynamical behaviour of a compartmentalized programmable biochemical oscillator. Nat. Chem. 6, 295–302 (2014).
Sokolova, E. et al. Enhanced transcription rates in membrane-free protocells formed by coacervation of cell lysate. Proc. Natl Acad. Sci. USA 110, 11692–11697 (2013).
Dubuc, E. et al. Cell-free microcompartmentalised transcription-translation for the prototyping of synthetic communication networks. Curr. Opin. Biotechnol. 58, 72–80 (2019).
Zhu, T. F. & Szostak, J. W. Coupled growth and division of model protocell membranes. J. Am. Chem. Soc. 131, 5705–5713 (2009).
Nakashima, K. K., van Haren, M. H., André, A. A., Robu, I. & Spruijt, E. Active coacervate droplets are protocells that grow and resist Ostwald ripening. Nat. Commun. 12, 3819 (2021).
Kurihara, K. et al. Self-reproduction of supramolecular giant vesicles combined with the amplification of encapsulated DNA. Nat. Chem. 3, 775–781 (2011).
Elani, Y., Law, R. V. & Ces, O. Vesicle-based artificial cells as chemical microreactors with spatially segregated reaction pathways. Nat. Commun. 5, 5305 (2014).
Drobot, B. et al. Compartmentalised RNA catalysis in membrane-free coacervate protocells. Nat. Commun. 9, 3643 (2018).
Iglesias-Artola, J. M. et al. Charge-density reduction promotes ribozyme activity in RNA–peptide coacervates via RNA fluidization and magnesium partitioning. Nat. Chem. 14, 407–416 (2022).
Donau, C. et al. Active coacervate droplets as a model for membraneless organelles and protocells. Nat. Commun. 11, 5167 (2020).
Wilson, D. A., Nolte, R. J. & Van Hest, J. C. M. Autonomous movement of platinum-loaded stomatocytes. Nat. Chem. 4, 268–274 (2012).
Kumar, B., Patil, A. J. & Mann, S. Enzyme-powered motility in buoyant organoclay/DNA protocells. Nat. Chem. 10, 1154–1163 (2018).
Rodríguez-Arco, L., Li, M. & Mann, S. Phagocytosis-inspired behaviour in synthetic protocell communities of compartmentalized colloidal objects. Nat. Mater. 16, 857–863 (2017).
Qiao, Y., Li, M., Booth, R. & Mann, S. Predatory behaviour in synthetic protocell communities. Nat. Chem. 9, 110–119 (2017).
Qiao, Y., Li, M., Qiu, D. & Mann, S. Response‐retaliation behavior in synthetic protocell communities. Angew. Chem. Int. Ed. 131, 17922–17927 (2019).
Merindol, R., Loescher, S., Samanta, A. & Walther, A. Pathway-controlled formation of mesostructured all-DNA colloids and superstructures. Nat. Nanotechnol. 13, 730–738 (2018).
Villar, G., Graham, A. D. & Bayley, H. A tissue-like printed material. Science 340, 48–52 (2013).
Alcinesio, A. et al. Controlled packing and single-droplet resolution of 3D-printed functional synthetic tissues. Nat. Commun. 11, 2105 (2020).
Booth, M. J., Restrepo Schild, V., Box, S. J. & Bayley, H. Light-patterning of synthetic tissues with single droplet resolution. Sci. Rep. 7, 9315 (2017).
Tian, L. et al. Spontaneous assembly of chemically encoded two-dimensional coacervate droplet arrays by acoustic wave patterning. Nat. Commun. 7, 13068 (2016).
Yang, Z., Wei, J., Sobolev, Y. I. & Grzybowski, B. A. Systems of mechanized and reactive droplets powered by multi-responsive surfactants. Nature 553, 313–318 (2018).
Dupin, A. & Simmel, F. C. Signalling and differentiation in emulsion-based multi-compartmentalized in vitro gene circuits. Nat. Chem. 11, 32–39 (2019).
Ramsay, K., Levy, J., Gobbo, P. & Elvira, K. S. Programmed assembly of bespoke prototissues on a microfluidic platform. Lab. Chip. 21, 4574–4585 (2021).
Deshpande, S. et al. Spatiotemporal control of coacervate formation within liposomes. Nat. Commun. 10, 1800 (2019).
Gobbo, P. et al. Programmed assembly of synthetic protocells into thermoresponsive prototissues. Nat. Mater. 17, 1145–1153 (2018).
Galanti, A. et al. A floating mold technique for the programmed assembly of protocells into protocellular materials capable of non‐equilibrium biochemical sensing. Adv. Mater. 33, 2100340 (2021).
Liu, J. et al. Hydrogel‐immobilized coacervate droplets as modular microreactor assemblies. Angew. Chem. Int. Ed. 59, 6853–6859 (2020).
McMullen, A. et al. Self-assembly of emulsion droplets through programmable folding. Nature 610, 502–506 (2022).
Gong, J., Tsumura, N., Sato, Y. & Takinoue, M. Computational DNA droplets recognizing miRNA sequence inputs based on liquid–liquid phase separation. Adv. Funct. Mater. 32, 2202322 (2022).
Funasaki, N. & Neya, S. Multiple complexation of didecyldimethylammonium bromide and cyclodextrins deduced from electromotive force measurements. Langmuir 16, 5343–5346 (2000).
Mu, W. et al. Membrane-confined liquid-liquid phase separation toward artificial organelles. Sci. Adv. 7, eabf9000 (2021).
Fraccia, T. P. & Jia, T. Z. Liquid crystal coacervates composed of short double-stranded DNA and cationic peptides. ACS Nano. 14, 15071–15082 (2020).
Jing, H. et al. Fission and internal fusion of protocell with membraneless ‘organelles’ formed by liquid–liquid phase separation. Langmuir 36, 8017–8026 (2020).
Fisher, R. S. & Elbaum-Garfinkle, S. Tunable multiphase dynamics of arginine and lysine liquid condensates. Nat. Commun. 11, 4628 (2020).
Lu, T. & Spruijt, E. Multiphase complex coacervate droplets. J. Am. Chem. Soc. 142, 2905–2914 (2020).
Kaur, T. et al. Sequence-encoded and composition-dependent protein–RNA interactions control multiphasic condensate morphologies. Nat. Commun. 12, 872 (2021).
Katzir, I., Haimov, E. & Lampel, A. Tuning the dynamics of viral-factories-inspired compartments formed by peptide–RNA liquid–liquid phase separation. Adv. Mater. 34, 2206371 (2022).
Valiev, M. et al. NWChem: a comprehensive and scalable open-source solution for large scale molecular simulations. Comput. Phys. Commun. 181, 1477–1489 (2010).
Rego, L. G. C. & Bortolini, G. Modulating the photoisomerization mechanism of semiconductor-bound azobenzene-functionalized compounds. J. Phys. Chem. C 123, 5692–5698 (2019).
Benmensour, M. A. et al. Azobased iminopyridine ligands and their rhenium metal complexes: syntheses, spectroscopic, trans–cis photoisomerization and theoretical studies. J. Photoch. Photobio. A 368, 78–84 (2019).
Klamt, A. & Schüürmann, G. COSMO: a new approach to dielectric screening in solvents with explicit expressions for the screening energy and its gradient. J. Chem. Soc. Perkin Trans. 2, 799–805 (1993).
Stefanovich, E. V. & Truong, T. N. Optimized atomic radii for quantum dielectric continuum solvation models. Chem. Phys. Lett. 244, 65–74 (1995).
Acknowledgements
This work was supported by the Strategic Priority Research Program of the Chinese Academy of Sciences (grant no. XDB0480000) to Y.Q., the National Natural Science Foundation of China (grant nos. 22272183 and 22072159 to Y.Q., and 22172007 to Y.L.), the Science Fund for Creative Research Groups of the National Natural Science Foundation of China (grant no. 52221006) to Y.L., and the Fundamental Research Funds for the Central Universities (grant nos. buctrc202015 and PT2208 to Y.L). S.M. was funded by the ERC Advanced Grant Scheme (grant no. EC-2016-674 ADG 740235).
Author information
Authors and Affiliations
Contributions
W.M. and L.J. performed the experiments and analysed the data. M.Z. and J.W. performed computational experiments. Y.Q. led the project. Y.L., S.M. and Y.Q. conceived, designed and supervised the study, analysed the data and wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemistry thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Video captions 1–4, materials and methods, Figs. 1–33, Tables 1–3 and references.
Supplementary Video 1
CLSM video showing the assembly of chain-like protocell networks comprising an alternating interconnected sequence of Nile-red-loaded DDAB/trans-AzoAsp2 (red fluorescence) and HPTS-loaded PDDA/trans-AzoAsp2 (green fluorescence) coacervate droplets are observed. Video is shown at ×540 of real-time speed at 3 fps. Total duration of recording in real time is 60 min; scale bar, 13 μm.
Supplementary Video 2
CLSM video showing HRP/H2O2-mediated peroxidation in coacervate-based protocell networks. Resorufin (red fluorescence) is produced and retained specifically within the DDAB/trans-AzoAsp2 domains. Video is shown at ×100 of real-time speed at 5 fps. Total duration of recording in real time is 600 s; scale bar, 5 μm.
Supplementary Video 3
CLSM video showing transfer of TAMRA-ssDNA from a DDAB/trans-AzoAsp2 droplet (red fluorescence at t = 0) to an adjacent PDDA/trans-AzoAsp2 domain (no fluorescence at t = 0) located in the same protocell chain. Video is shown at ×5 of real-time speed at 10 fps. Total duration of recording in real time is 38.5 s; scale bar, 10 μm.
Supplementary Video 4
Optical microscopy video showing UV-induced reconfiguration of protocell networks due to light-mediated trans-to-cis isomerization of azobenzene in ordered chains of DDAB/trans-AzoAsp2 and PDDA/trans-AzoAsp2 droplets. UV source; 325 nm < λ < 375 nm, 120 W short-arc Hg light. Video is shown at ×75 of real-time speed at 5 fps. Total duration of recording in real time is 10 min; scale bar, 5 μm.
Source data
Source Data Fig. 2
Statistical Source Data.
Source Data Fig. 3
Statistical Source Data.
Source Data Fig. 4
Statistical Source Data.
Source Data Fig. 5
Statistical Source Data.
Source Data Fig. 6
Statistical Source Data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mu, W., Jia, L., Zhou, M. et al. Superstructural ordering in self-sorting coacervate-based protocell networks. Nat. Chem. 16, 158–167 (2024). https://doi.org/10.1038/s41557-023-01356-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-023-01356-1
This article is cited by
-
Multicompartmental coacervate-based protocell by spontaneous droplet evaporation
Nature Communications (2024)
-
Self-assembly of stabilized droplets from liquid–liquid phase separation for higher-order structures and functions
Communications Chemistry (2024)