Abstract
A virus hijacks host cellular machineries and metabolites in order to reproduce. In response, the innate immune system activates different processes to fight back. Although many aspects of these processes have been well investigated, the key roles played by iron–sulfur [FeS] clusters, which are among the oldest classes of bio-inorganic cofactors, have barely been considered. Here we discuss how several [FeS] cluster-containing proteins activate, support and modulate the innate immune response to restrict viral infections, and how some of these proteins simultaneously support the replication of viruses. We also propose models of function of some proteins in the innate immune response and argue that [FeS] clusters in many of these proteins act as biological ‘fuses’ to control the response. We hope this overview helps to inspire future research in the emerging field of bio-inorganic virology/immunology and that such studies may reveal new molecular insight into the links between viral infections and diseases like cancer and neurodegeneration.

Similar content being viewed by others
Main
Iron–sulfur [FeS] clusters constitute one of the most versatile and ancient classes of bio-inorganic cofactors. In these clusters the Fe and S atoms exist in different stoichiometries and the coordinating ligands to the irons of the cluster can deviate in multiple ways from the archetypal all-cysteine (Cys) pattern (Fig. 1a)1. They have diverse functional roles (Fig. 1b–f), as described in the following. (1) They form a relay for long-range electron transfer2 in enzymes (the function of which is fundamental to life on Earth) such as hydrogenases (enabling many microorganisms to make or use H2 and conserve energy in the form of adenosine triphosphate (ATP))3, photosystem I (the integral membrane protein complex using light to generate electrons for the reduction of nicotinamide adenine dinucleotide phosphate (NADP+))4 or nicotinamide adenine dinucleotide (NADH):ubiquinone oxidoreductase (Fig. 1b; respiratory complex I in mitochondria, a giant, membrane-bound enzyme that plays an integral role in generating the proton-motive force driving ATP synthesis)5. In these enzymes, each cluster is characterized by a normally negative redox potential and a low activation barrier to electron transfer, and the clusters are usually not solvent-accessible. (2) They perform catalysis. A well-studied example is the [4Fe-4S](Cys)3 cluster of the radical S-adenosylmethionine (SAM) enzymes, which in most cases reductively cleaves SAM to generate the 5′-deoxyadenosyl radical (5′-dA• radical) intermediate (Fig. 1c)6,7. Members of the radical–SAM superfamily of enzymes play various biological functions from modification of RNA or DNA to biosynthesis of cofactors and peptides6,7. Another example is the [4Fe-4S](Cys)3 cluster in aconitase. This cluster acts as a Lewis acid (Fig. 1c) to facilitate isomerization of citrate to iso-citrate in the tricarboxylic acid cycle8. (3) The clusters provide structural stability to a protein or a multiprotein complex, for example, the [4Fe-4S](Cys)4 cluster in the catalytic subunit of DNA polymerase δ (DNA Pol-δ, a multi-subunit protein complex involved in the replication and repair of DNA in eukaryotes)9. The solvent-exposed cluster is located at the interface of the catalytic and regulatory subunits (Fig. 1d) and ensures the structural integrity and activity of the protein complex10. (4) The clusters act as sensors of cellular environments, responding to particular levels of O2, nitric oxide (NO) or hydrogen peroxide (H2O2). By altering the structure of a cluster, these agents induce conformational changes in the respective proteins, regulating their functions. Sensory clusters include the [4Fe-4S](Cys)4 cluster in the O2-sensing fumarate and nitrate reductase regulatory protein (FNR)11, the [4Fe-4S](Cys)3X (where X is unknown) rhizobial iron regulator (RirA)12 and the [4Fe-4S](Cys)3(Asp) cluster of the nitrite-responsive repressor (NsrR)13. In these proteins the reaction of solvent-exposed [FeS] clusters with O2 or NO induces or blocks the interaction of proteins with DNA, thereby regulating the expression of multiple genes. Reaction of NO with the [4Fe-4S](Cys)3(Asp) cluster of NsrR results in nitrosylation, degradation of the cluster and formation of a mixture of protein-bound iron nitrosyl species and apo-protein (Fig. 1e)14. The reaction of the cluster in FNR with O2 converts the solvent-exposed [4Fe-4S](Cys)4 cluster to a [2Fe-2S](Cys)4 cluster. This conversion induces conformational changes in the protein scaffold and switches the function (Fig. 1e)15. (5) The cluster acts as a source of sulfur. For example, the auxiliary [4Fe-4S](Cys)3(Ser) cluster in lipoyl synthase (LipA) supplies two S atoms for conversion of the protein-bound octanoyl chain to the lipoyl cofactor (Fig. 1f)16. In eukaryotes, synthesis and delivery of the [FeS] clusters to the target proteins is a complex process that has been reviewed in detail elsewhere17,18,19,20. Briefly, the mitochondrial iron–sulfur cluster (ISC) assembly machinery is involved in the maturation of all cellular iron–sulfur proteins present in mitochondria, the cytosol and the nucleus. Indeed, the ISC machinery presumably provides a [FeS] compound to the cytosolic iron–sulfur protein assembly (CIA) machinery21, which carries out the synthesis of [4Fe-4S] clusters in the cytosol17,18,19,20. In the cytosol, the assembled [4Fe-4S] clusters are then delivered to cytosolic and nuclear target proteins by iron-only hydrogenase-like protein 1 (IOP1) and accessory proteins including one or more components of the CIA targeting complex and the CIA2A protein17,18,19,20.
a, Six common forms of [FeS] clusters. These clusters have various functional roles in different proteins. b, Electron transfer. For example, a relay of [FeS] clusters in the respiratory complex I enables long-range electron transfer (PDB 6YJ4). c, Catalysis. For example, the [4Fe-4S]1+(Cys)3 cluster in most members of the radical–SAM enzymes reductively cleaves SAM to generate the 5′-dA• radical intermediate, or the [4Fe-4S](Cys)3 cluster in aconitase participates in Lewis-acid catalysis. d, Structural stability. For example, the solvent-exposed [4Fe-4S](Cys)4 cluster at the interface of catalytic and regulatory subunits of DNA Pol-δ gives structural integrity to the complex (PDB 6TNY). e, Sensor. For example, the solvent-exposed [4Fe-4S](Cys)3(Asp) cluster in NsrR (PDB 5N07 and 5N08) reacts with NO, leading to degradation of the cluster and formation of apo-NsrR and NsrR-bound iron-nitrosyl species. The solvent-exposed [4Fe-4S](Cys)4 cluster in FNR (PDB 5E44) reacts with O2, leading to formation of an FNR-bound [2Fe-2S](Cys)4 cluster, inducing a conformational change in the protein (PDB 5CVR). f, Sulfur donor. For example, the [4Fe-4S](Cys)3(Ser) auxiliary cluster in LipA (PDB 5EXJ) supplies two S atoms in the last step of the biosynthesis of the lipoyl cofactor. 5′-dAH, 5′-deoxyadenosine.
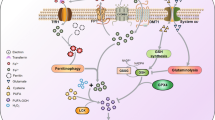
Viral infections are a global challenge and cause huge social and economic cost worldwide. The life cycle of all viruses can be divided into four stages (Fig. 2)22, the details of which vary depending on the virus: (1) host-cell entry, (2) replication, (3) assembly of viral particles and (4) egress. Some viral proteins contain [FeS] clusters23,24,25. In viral proteins, [FeS] clusters play a structural and/or catalytic role25. For example, the RNA-dependent RNA polymerase (RdRp) of SARS-CoV-2 has been shown to have two [4Fe-4S](Cys)3(His) clusters. The clusters are suggested to support the catalytic activity of the enzyme and supply structural stability to the RdRp complex25. Beside the viral factors (proteins, nucleic acids) involved in each step, cellular metabolites and proteins are hijacked by viruses to complete their life cycle. It is anticipated that the host [FeS] biosynthesis apparatus is also hijacked for the maturation of viral [FeS] proteins25.
Top: a summary of the viral life cycle. The steps leading to viral replication are indicated by orange arrows. After entry of a virus into a cell, viral factors such as nucleic acids and proteins are recognized by [FeS]-containing proteins (recognition step), beside other mechanisms. This signal is translated by [FeS] proteins to the overexpression of proinflammatory cytokines including IFNs and then expression of ISGs and activation of proinflammatory signalling (transcription and translation steps). At the same time, some viral [FeS] proteins (not shown) support replication of the viral genome and some host [FeS] proteins support translation of viral proteins (translation step). Some ISG proteins require [FeS] clusters to restrict replication of a virus. Expression of ISG proteins and activation of proinflammatory signalling pathways by cytokines cause inflammation, inducing metabolic changes and a burst in production (red arrow) of NO and H2O2 concomitant with harmful lowering levels (grey arrow) of O2 (hypoxia). The steps of the innate immune response leading to the formation of NO and H2O2 and to hypoxia are indicated by black arrows. The rising cellular levels of NO and H2O2 and hypoxia are sensed by [FeS] clusters modulating iron metabolism and mitophagy (feedforward mechanism, purple arrows). At the same time, NO and H2O2 can degrade the solvent-exposed [FeS] clusters of many proteins involved in all the steps of the innate immune response, halting an excessive production of NO and H2O2 (feedback mechanism, blue arrows).
In response to a viral infection, the innate immune system recognizes components of the virus, inducing expression of proinflammatory cytokines including type I and II interferons (IFNs)26,27. Next, type I and II IFNs induce expression of interferon stimulated genes (ISGs) to restrict replication of the virus (Fig. 2)28,29,30. Expression of ISG proteins and proinflammatory cytokines leads to inflammation, the hallmarks of which are a rise in the production of reactive oxygen and nitrogen species like H2O2 and NO (refs. 31,32) and insufficient levels of dioxygen (hypoxia; Fig. 2)33. In this Review, we will focus exclusively on the host cell [FeS] proteins involved in the immune response and viral replication process (Table 1). We discuss how [FeS] clusters in different proteins control and modulate the innate immune response to a virus (Fig. 2) from the early steps of viral entry to reprogramming metabolism and modulating mitophagy (the process by which damaged mitochondria are selectively degraded). We give examples of the cellular [FeS] proteins involved in inhibiting or catalysing the viral replication process and discuss their structural features, the function of [FeS] clusters, and the cellular requirement for [FeS] cluster insertion into these proteins, while highlighting common features. We propose that the [FeS] clusters present in some proteins act as a sensor of cellular levels of reactive oxygen species (ROS) and O2, enabling feedforward mechanisms to regulate processes such as iron homeostasis and mitophagy, while in many other proteins they act as biological ‘fuses’, enabling a feedback mechanism to prevent an uncontrolled immune response and excessive production of reactive oxygen and nitrogen species (Fig. 2). Our analysis highlights the importance of future studies to elucidate the role of [FeS] clusters in the viral replication process and the innate immune response.
Activating the innate immune response
The first step in activation of the innate immune response to viral infections is recognition of the invading virus to initiate a series of cellular processes and restrict viral replication. In this section we discuss how several [FeS] proteins play central roles in recognizing viral components to activate the innate immune response. We demonstrate that these [FeS] proteins recognize viral components such as nucleic acids and proteins and activate the expression of type I and II IFNs and potentially other proinflammatory cytokines, while at the same time some of them support expression of viral proteins.
Initiating transcription of IFNs and supporting translation
Many RNA and DNA processing proteins contain [FeS] clusters34, including the catalytic subunit of DNA polymerase α (DNA Pol-α)9,10,35, RNA Pol-III36,37 and the elongator complex (a multi-subunit protein complex that is highly conserved in eukaryotes, with roles in the activity of RNA Pol-II and the modification of transfer RNA (tRNA))38,39,40. The [FeS] clusters of POLA1, RNA Pol-III and the elongator complex support their functions in sensing viral components, initiating translation of IFNs and supporting translation (Fig. 3a). DNA Pol-α has four subunits: POLA1, a regulatory (POLA2) and two primase (PRIM1 and PRIM2) subunits. DNA Pol-α synthesizes the RNA:DNA primers required for DNA replication on both leading and lagging strands of DNA41. The C terminus of POLA1 has a [4Fe-4S](Cys)4 cluster that is essential for the formation of a DNA Pol-α protein complex and its enzymatic activity10. In the nucleus, RNA Pol-III transcribes genes encoding 5S ribosomal RNA, tRNA and other small RNAs42. Structural studies of RNA Pol-III in the presence and absence of DNA substrate revealed that the subunit 6 (RPC6) of RNA Pol-III has a [4Fe-4S](Cys)4 cluster36,37 that stabilizes the C-terminal domain of the protein (Fig. 3b). The [FeS] cluster domain of RPC6 forms a knot to tie the RPC7–RPC6–RPC3 heterotrimer to the core complex36,37. In the absence of DNA, the cluster is solvent-exposed via a shallow funnel and, upon binding of RNA Pol-III to DNA, the funnel is closed by the C-terminal tail of the RPC7 subunit (Fig. 3b).
a, A summary of the roles of [4Fe-4S] clusters of RNA Pol-III and Elp3 in activating and regulating the IFN response due to viral infection (see main text for details). Although the solvent-exposed clusters are essential to recognize viral components, activate the expression of IFNs and modify the cellular pool of tRNAs, they may also sense the increasing cellular levels of NO or H2O2 caused by the innate immune response. Consequently, the clusters are degraded, inducing conformational changes in the protein–protein complexes and halting their activity. Delivery of the clusters by the CIA targeting complex will reactivate the proteins. In the case of Elp3, an unknown reducing partner should provide electrons for catalysis. IRF, interferon regulatory factor; Ac, acetyl group. b, Structure of the RNA Pol-III and its [FeS] binding domain. Comparison of the structure of RNA Pol-III in the presence of DNA (PDB 7AEA) with that solved in the absence of DNA (PDB 7A6H) shows that binding of DNA allows the long tail of RPC7 to close the shallow funnel exposing the [4Fe-4S] cluster to solvent (dashed arrow). c, Structure of Elp3 in complex with Elp1 and Elp2 (PDB 6QK7). The KAT (light green) and rSAM (dark green) domains of Elp3 have a large interface forming a pocket, where the wobble position of tRNA can enter (blue arrow). Elp3 has a [4Fe-4S](Cys)3 cluster, which is exposed to solvent via two channels (dashed arrows). d, Mechanism of modification of tRNA by Elp3. In the cytoplasm, the cluster reductively cleaves SAM to initiate the radical–SAM catalysis and addition of carboxyl-methyl (cm) to the tRNA wobble uridine at C5 (cm5) (see main text for the details).
The multiprotein complex RNA Pol-II requires activity of the elongator complex43. The catalytic subunit of the elongator complex is Elp3, which harbours a [4Fe-4S](Cys)3 cluster (Fig. 3c)44. Elp3 has two catalytic domains: a C-terminal lysine acetyl–coenzyme A (acetyl-CoA) transferase (KAT) domain and an N-terminal radical–SAM (rSAM) domain, which binds a [4Fe-4S](Cys)3 cluster45. Unlike most radical–SAM enzymes, the three cysteine residues coordinating the [4Fe-4S] cluster are not part of a CX3CX2C motif but are coordinated by three cysteine residues in a CX9CX2C primary sequence (Fig. 3c). The cluster is accessible from solvent via two channels (Fig. 3c) and it provides structural integrity to the elongator complex46, analogous to the role of the [4Fe-4S](Cys)4 cluster in stabilizing RNA Pol-III and DNA Pols. When the elongator complex is in the nucleus, Elp3 catalyses acetylation of histones47,48. When in the cytoplasm, Elp3 catalyses the addition of carboxyl-methyl (cm) to the tRNA third anticodon nucleotide (wobble) uridine at C5 (ref. 49). This is referred to as ‘cm5’ modification. The mechanism of DNA acetylation by Elp3 is sparsely characterized, but the mechanism of tRNA modification has been studied in detail38,50,51. Amino-acid sequence analysis and biochemical studies using electron paramagnetic resonance (EPR) spectroscopy confirmed that SAM binds to the [4Fe-4S] cluster of Elp344. High-resolution cryogenic electron microscopy (cryo-EM) structures of Elp3 in the catalytic unit of the elongator complex38, consisting of Elp1, 2 and 3 (Elp123), show that Elp3 is sandwiched between Elp1 and Elp2 (Fig. 3c). The KAT and rSAM domains have a unique arrangement with a large interface, where tRNA can bind (Fig. 3c)38,50,51. Although the conformation of Elp3 and its catalytic pocket involving the [4Fe-4S] cluster and SAM appears not to change upon tRNA binding, the conformation of tRNA does change, allowing entry of the wobble uridine directly into the channel linked to the active site38. Like most radical–SAM enzymes, the reduced [4Fe-4S]1+ cluster reductively cleaves SAM to generate the 5′-dA• radical intermediate (Fig. 3d), which abstracts a H atom from the methyl group of acetyl-CoA49. The resulting radical attacks the C5 position of uridine, forming a C–C bond. This step is followed by rearrangements, loss of an electron and a proton, and formation of tRNA-acetyl-CoA, which is hydrolysed to generate CoA and cm5U-tRNA (Fig. 3d).
POLA1, RNA Pol-III and the elongator complex play vital roles in recognizing viral components and activating the innate immune response. POLA1 has similarity to the DNA polymerases of a number of DNA viruses52 and interacts with viral DNA in the cytosol53 or proteins like the non-structural protein (NSP1) of SARS-CoV-254,55. Although the functions of these interactions are not understood, they can act as a recognition mechanism to activate a type I IFN response. Specifically, interaction of viral DNAs or proteins with POLA1 may inhibit the activity of DNA Pol-α in the synthesis of RNA:DNA primers, affecting their cytosolic levels, which modulates IFN production56. Unlike POLA1, the role of RNA Pol-III in recognizing viral DNA is clearer. Mutations in the RNA Pol-III gene are associated with defects in viral DNA recognition and immune response57. In the cytoplasm, RNA Pol-III recognizes viral DNAs (Fig. 3a) via an unknown mechanism and catalyses the synthesis of double-stranded RNA (dsRNA) with a 5′-triphosphate moiety to stimulate IFN production (Fig. 3a)58,59,60. In addition, in the nucleus, RNA Pol-III modulates the IFN and antiviral immune response61,62, possibly by catalysing transcription of tRNAs and regulating their cellular pool (Fig. 3a)63. After recognizing viral components by cytosolic RNA Pol-III and other cellular mechanisms26,27, complex signalling cascades are activated to initiate transcription of the messenger RNAs (mRNAs) of IFNs and other proinflammatory cytokines in the nucleus26,27. The RNA Pol-II complex is recruited to IFN promoters to transcribe IFN mRNAs (Fig. 3a)64. Recruitment of RNA Pol-II is modulated by acetylation of histones, including those of H3 and H4, which are targets of Elp3 (Fig. 3a)47,48. Viral infection causes hyperacetylation of H3 and H4 at the IFN-β promoter65, and acetylation at H3Lys14 promotes hyperacetylation by inducing resistance to deacetylation and demethylation66, both of which could repress transcription of proinflammatory cytokines like type I IFNs67. Thus, acetylation of histones H3 and H4 by Elp3 of the elongator complex47,48 could prime the innate immune response recruiting RNA Pol-II. Consistently, the Elp3 subunit of the elongator complex plays a vital role in epigenetics and regulation of the immune response68, and in Drosophila melanogaster69, plants70 and humans71,72, Elp3 positively regulates the innate immune response. Additionally, in the cytoplasm the radical–SAM activity of Elp3 should play a role in priming innate and trained immunity. The radical–SAM activity of Elp3 induces cm5 modification of the tRNA wobble position49 to modulate the cellular pool of tRNA (Fig. 3a). Changes in the cellular pool of tRNA, as observed upon IFN stimulation and viral infection73, could regulate translation of type I and II IFNs and potentially other proinflammatory cytokines, ISGs and viral proteins.
Although the [4Fe-4S] clusters of RNA Pol-III and Elp3 have both structural and/or catalytic roles to support the function of proteins in activating the innate immune response, the solvent-exposed clusters may also act as sensors of the rising cellular levels of NO or H2O2 during inflammation (Fig. 3a), which are caused by the innate immune response (Fig. 2). The mechanism by which the solvent-exposed [4Fe-4S] clusters in RNA Pol-III and Elp3 may sense NO or H2O2 is not clear, but it may be similar to that observed for the solvent-exposed [FeS] clusters of NO and O2 sensory proteins (Fig. 1e). Specifically, NO or H2O2 reacts with the clusters and degrades them, thereby altering and destabilizing the structure of RNA Pol-III and the elongator complex. These structural changes block the ability of these proteins to maintain continuous expression of IFNs and other proinflammatory cytokines, serving as a feedback mechanism to prevent excessive expression of ISG proteins and formation of cytotoxic levels of NO or H2O2 (Fig. 3a). Reactivation of the proteins requires the CIA targeting complex to restore the [FeS] clusters and, in the case of Elp3, an unknown reducing partner to supply electrons for catalysis (Fig. 3a).
Enabling translation of cytokines, ISGs and viral proteins
After transcription of proinflammatory cytokines, ISGs and viral mRNAs, ribosomes translate the mRNAs to proteins (Fig. 2). The eukaryotic translation process consists of four steps: initiation, elongation, termination and recycling. Two proteins for which clusters regulate activity of the translation machinery (Fig. 4a) are the Dph2 homodimer in archaea (the Dph1–Dph2 heterodimer in eukaryotes)74 and ABCE1, originally known as an inhibitor of RNase L (RLI1)75. Dph1–Dph2 catalyses the first step in the biosynthesis of diphthamide, a post-translational modification of the histidine 715 residue (mammals) in the translation elongation factor 2 (EF2)74. Although the exact function of diphthamide modification has remained elusive, it is shown to be essential for the function of EF2 in translocation of peptidyl-tRNA from the A site of the ribosome to the P site (peptidyl-tRNA site) and the fidelity of the translation process76. While a specific role of diphthamide in the innate immune response is unknown, its loss pre-activates nuclear factor kappa B (NF-κB)77, a transcription factor and a regulator of the innate immunity. The heterodimeric Dph1–Dph2 has two [4Fe-4S](Cys)3 clusters similar to that present in members of the radical–SAM enzymes. The structure of Dph2 shows that the clusters are solvent-exposed (Fig. 4b)78; however, unlike most radical–SAM enzymes, each cluster is coordinated to three distant cysteines in the primary sequence, CX105CX125C (Fig. 4b). Thus, the cluster should confer structural stability and integrity besides its role in catalysis. Biochemical and spectroscopic studies using the eukaryotic Dph1–Dph2 heterodimer revealed that the cluster in Dph1 is essential for catalysis and the cluster in Dph2 is involved in electron transfer to the cluster in Dph179. Unlike most radical–SAM enzymes, the [4Fe-4S]1+ cluster of Dph1–Dph2 reductively cleaves SAM at the γ carbon of the methionine to generate a 3-amino-3-carboxypropyl (ACP)-[4Fe-4S]3+ organometallic intermediate and 5′-methylthioadenosine (MTA; Fig. 4c)78,80. Subsequently, the organometallic intermediate attacks the imidazole ring of the histidine residue of EF2, generating an organic radical intermediate, which, on losing an electron and proton, generates ACP-modified histidine (Fig. 4c)78,80.
a, The [FeS] cluster-containing proteins Dph1–Dph2 and ABCE1 orchestrate the translation machinery to express proinflammatory cytokines, ISG proteins and viral proteins. The solvent-exposed clusters sense the increasing cellular level of reactive oxygen or nitrogen species to modulate translation and prevent excessive expression of proinflammatory cytokines and ISG proteins. b, Structure of Dph2 homodimer (PDB 3LZD) showing the solvent-exposed [4Fe-4S](Cys)3 cluster (dashed arrow). c, The proposed mechanism by which Dph1–Dph2 is involved in the first step of biosynthesis of diphthamide (see main text for detail). MTA, 5′-methylthioadenosine. d, Structure of ABCE1 showing two [4Fe-4S] clusters in the [FeS] cluster-binding domain and two ADP in the twin ATP-binding domain (PDB 3BK7). e, The proposed molecular mechanism of the function of ABCE1 and its [FeS] cluster-binding domain in ribosome recycling and translation (see main text for details). After splitting of ribosomal subunits, other factors (not shown, to retain figure clarity)94 are needed for formation of the initiation complex.
After the elongation step, the highly conserved ABCE1 protein orchestrates ribosome recycling. This protein is present in all archaea and eukaryotes, and its function is vital both for cells81 and for translation of viral mRNAs82. The protein interacts with several translation initiation factors and with the 40S ribosomal subunit83,84,85. Biochemical and biophysical experiments using EPR and Mössbauer spectroscopy confirmed that the protein has two diamagnetic [4Fe-4S]2+ clusters, which cannot be reduced using sodium dithionite86. Structural studies of the ABCE1 from Pyrococcus abyssi revealed that the protein has an N-terminal [FeS] cluster-containing domain with two [4Fe-4S](Cys)4 clusters (Fig. 4d)87. Each cluster is coordinated by three nearby cysteines and one that is distant in the primary sequence (Fig. 4d), [4Fe-4S]A:CX4CX4CX39C and [4Fe-4S]B:CX25CX2CX2C, implying the importance of the [4Fe-4S] clusters for the structural stability of the protein. One cysteine residue of each cluster is solvent-exposed (Fig. 4d) and the clusters are prone to degradation by ROS88. Extensive structural and biochemical studies have yielded molecular insight into the role of the [FeS] cluster domain in ribosome recycling89,90,91,92,93,94,95,96. Briefly, upon ATP binding to both sites in ABCE1, the protein undergoes a conformational switch from an open to a closed ATP-occluded state, which drives ribosome dissociation (Fig. 4e)95,96. In the closed state, the [FeS] cluster domain of ABCE1 rotates through 150° and repositions itself into a binding pocket on the 40S subunit. This movement suggests a collision with A-site factors splitting the ribosomal subunits (Fig. 4e)93. ATP hydrolysis is required for ABCE1 release from the 40S subunit to re-enter a new cycle.
The activities of [FeS] proteins Dph1–2Dph2 and ABCE1 together orchestrate translation of the mRNAs (Fig. 4a). Their [4Fe-4S] clusters give structural stability to the proteins and enable their function (Fig. 4). The clusters are solvent-exposed and can sense the cellular levels of NO or H2O2, which degrade the clusters and inhibit their activity in translation. Hence, like the [FeS] clusters of RNA Pol-III and Elp3, the [FeS] clusters of Dph1–Dph2 and ABCE1 enable a feedback mechanism to respond to the increasing cellular levels of NO and H2O2 caused by the innate immune response. This feedback mechanism can serve as a safeguard to prevent excessive production of proinflammatory cytokines and ISG proteins and consequently the formation of cytotoxic levels of NO and H2O2. After degradation of the clusters, formation of active holo-ABCE1 requires the function of the CIA targeting complex and two accessory proteins, namely YAE1D1 (Yae1 domain containing 1) and OROAV1 (the oral cancer overexpressed 1) proteins, to deliver two [4Fe-4S] clusters (Fig. 4a)97,98. The specific cellular requirements for formation of holo-Dph1–Dph2 are unknown and it is proposed that the iron-binding protein Dph3 can provide electrons to Dph1–Dph2 (Fig. 4a)99.
Reprogramming metabolism
A consequence of induction of the expression of proinflammatory cytokines like type I and II IFNs is the induction of expression of ISG proteins28,29,30. One of the ISG proteins is the [FeS] protein RSAD2100,101. This protein and other ISG proteins collectively restrict replication of a wide range of viruses and induce a metabolic shift in the cell. Many metabolic proteins also contain [FeS] clusters including respiratory complexes I–III, mitochondrial LipA, and PPAT, which is the first enzyme in the de novo biosynthesis of purine nucleotides. In this section, we discuss how the [FeS] clusters in RSAD2 and PPAT regulate immunometabolism (Fig. 5a).
a, A summary of the role of the [FeS] clusters of RSAD2 and PPAT in immunometabolism. The nucleotide analogue ddhCTP is proposed to act as a chain terminator of viral RdRps or to inhibit the glycolytic enzyme GAPDH and regulate metabolism (see main text for details). Inhibition of GAPDH by ddhCTP increases the levels of intermediates in the pentose phosphate pathway (PPP), which support de novo synthesis of purines. The first enzyme in purine biosynthesis is the [FeS] protein PPAT. This enzyme catalyses the first step in the biosynthesis of inosine monophosphate (IMP) to supply ATP and guanosine triphosphate (GTP) for the activity of [FeS] proteins RNA Pol-II and RNA Pol-III supporting transcription of proinflammatory cytokines and ISGs. At the same time, it supplies ATP and GTP for viral genome synthesis. F6P, fructose 6-phosphate. b, Structure of vertebrate RSAD2 (viperin) showing the solvent-exposed [4Fe-4S](Cys)3 cluster (PDB 5VSL). Structural analysis of vertebrate RSAD2 in the presence of SAH and CTP (PDB 6Q2P) shows that H4′ faces toward the C5′ of the ribose in SAH, suggesting that this H atom is abstracted by the 5′-dA• radical. The highly conserved Tyr302, which could play a role in catalysis, is shown. c, The proposed mechanism of the catalytic conversion of CTP to ddhCTP by RSAD2. d, Structure of Bacillus subtilis PPAT (PDB 1AO0). The enzyme is a homotetramer, and each subunit has a solvent-exposed [4Fe-4S](Cys)4 cluster.
Modulating central carbon metabolism
The ISG protein RSAD2 was originally identified as an antiviral enzyme100,101, which localizes at the cytosolic face of the endoplasmic reticulum102 and lipid droplets103. Thus, the enzyme was termed viperin (virus inhibitory protein, endoplasmic reticulum-associated, interferon-inducible)104. This name, however, does not precisely describe the function of the enzyme because its expression can be induced by lipopolysaccharides (LPSs)105, its activity can enhance the infectivity of human cytomegalovirus (HCMV)106, and growing evidence supports a role of the protein in maturation of dendritic cells107, promoting adaptive behaviour of natural killer (NK) cells108, optimal T helper 2 (Th2) cell cytokine production109 and thermogenesis in adipose tissue110. Spectroscopic studies of human and fungal RSAD2111,112 and structural analysis of vertebrate RSAD2113 have shown that the protein has a [4Fe-4S](Cys)3 cluster (Fig. 5b). The cluster is solvent-exposed via a shallow funnel (Fig. 5b), like the [FeS] clusters of many proteins involved in activating the innate immune response. The radical–SAM activity of human RSAD2 catalyses the transformation of cytidine triphosphate (CTP) to the nucleotide analogue 3′-deoxy-3′,4′-didehydro CTP (ddhCTP)114. Structural analysis in the presence of CTP substrate revealed that the H4′ of ribose in CTP is positioned towards the C5′ of ribose in S-adenosylhomocysteine (SAH)115 (Fig. 5b), suggesting that H4′ is abstracted by the 5′-dA• radical, consistent with isotope labelling experiments114. The resulting substrate radical intermediate undergoes rearrangement reactions and one-electron reduction to generate the ddhCTP product (Fig. 5c). The highly conserved tyrosine (Tyr302), which is ~5.5 Å away from the ribose of CTP, could act as an electron donor for one-electron reduction of the substrate radical intermediate via a proton-coupled electron transfer step (Fig. 5c)116. The nucleotide analogue ddhCTP is proposed to be a chain terminator of RNA-dependent RNA polymerases (RdRps) of a number of flaviviruses with IC50 values (the inhibitor concentration at which the activity of an enzyme drops to 50% of its maximum value) in the range of 20–3,000 mM (Fig. 5a)114. These IC50 values appear to be not physiologically relevant given that the cellular concentration of ddhCTP is ~0.1–0.3 mM (ref. 114). Furthermore, the role of ddhCTP as a chain terminator of viral RdRps does not explain many reports describing the effect of RSAD2 enzymatic activity on cellular metabolism and behaviour, for example, glucose homeostasis117, inhibition of mitochondrial trifunctional enzyme complex118, thermogenesis in adipose tissue110, interference with the metabolism of amino acids119, maturation of dendritic cells107 or promotion of the adaptive behaviour of NK cells108. These activities together support an immunometabolic function of the product of the radical–SAM activity of RSAD2101. A role of ddhCTP in the regulation of immunometabolism was proposed through metabolomics studies of authentic macrophages derived from human-induced pluripotent stem cells (iPSCs)120. Metabolomics studies suggested that the radical–SAM activity of RSAD2 inhibits the NAD+-dependent activity of different enzymes, including the glycolytic enzyme glyceraldehyde 3-phosphate dehydrogenase (GAPDH; Fig. 5a)120. Biochemical studies confirmed that ddhCTP can inhibit the NAD+-dependent activity of different enzymes, including that of GAPDH, with an approximate IC50 value of 0.05 mM (ref. 120). Inhibition of the NAD+-dependent activity of GAPDH increases the cellular level of glyceraldehyde 3-phosphate (G3P or GAP)120 and potentially NAD+. The concomitant increase in the levels of G3P and NAD+ can support the synthesis of triacylglycerols (TAGs) and phosphatidylcholine (PC), and stimulate the synthesis of cyclic ADP-ribose (cADPr) and protein ADP-ribosylation (Fig. 5a)121.
Together, TAGs, PC, cADPr and protein ADP-ribosylation can stimulate the immune response in various ways (Fig. 5a)121, including (1) supporting the synthesis of eicosanoids, which are signalling lipids, (2) supporting antigen cross-presentation via major histocompatibility complex class-I (MHC-I), (3) stimulating the activity of transcription factors NFAT (nuclear factor of activated T cells) and NF-κB and (4) inducing synthesis of NO. Besides the function of the [4Fe-4S](Cys)3 cluster in the radical–SAM-mediated synthesis of ddhCTP, the cluster provides structural stability to the enzyme122, analogous to the [4Fe-4S] cluster in proteins activating the innate immune response (Figs. 3 and 4). The solvent-exposed [4Fe-4S] cluster of RSAD2 can act as a sensor of NO and H2O2, which degrade the cluster, induce conformational change in the protein structure and abolish activity of the enzyme. The degradation of the cluster can modulate the interactions of RSAD2 with other proposed partner proteins100. Thus, the [FeS] cluster of RSAD2 could enable a feedback mechanism to regulate the innate immune response and production of toxic levels of ddhCTP, which may cause cell death119,123. Although the cellular requirement for the reduction of the cluster is not known, insertion of the [4Fe-4S] cluster to form holo-enzyme is proposed to require the CIAO1 protein of the CIA targeting complex (Fig. 5a)124.
Modulating purine metabolism
Nucleotide metabolism is another metabolic process important to the innate immune response and viral infection. Although the innate immune response requires nucleotides for the transcription and synthesis of proinflammatory cytokines and ISGs mRNAs, and tRNAs, viruses require access to the cellular pool of nucleotides to replicate their genomic material (Fig. 5a). De novo biosynthesis of purines requires activation of the pentose phosphate pathway (PPP) to supply 5-phosphoribosyl-1-pyrophosphate (PRPP; Fig. 5d). Consistently, activation of PPP is a signature of proinflammatory macrophages (M1 macrophages)125, and cells infected with viruses like influenza activate the de novo pathway of purine metabolism126,127. The radical–SAM activity of the ISG protein RSAD2 increases the cellular levels of metabolites of PPP including ribose 5-phosphate (R5P)120, which is the substrate for the synthesis of PRPP. Subsequently, the [FeS] enzyme PPAT catalyses the first step in de novo biosynthesis of purines by converting PRPP into 5-phosphoribosyl-1-amine (PRA) using the side-chain amine group of a glutamine (Fig. 5a). Crystallographic studies of PPAT from Bacillus subtilis128, Escherichia coli129 and Arabidopsis thaliana130, in combination with sequence alignments, show that the enzyme from most archaeal, bacterial and eukaryotic species, including human, is a homotetramer with one [4Fe-4S](Cys)4 cluster per subunit (Fig. 5d). The cluster in the B. subtilis enzyme does not differ notably in geometric128 or electronic and magnetic structure131,132 from the [4Fe-4S]2+ clusters in electron-transfer proteins; however, it is not reduced under physiological conditions (reduction requires a potential more negative than −600 mV)133. The cluster is thought not to be directly involved in catalysis because it is located too far away from both the site at which glutamine is hydrolysed and a phosphoribosyl transferase domain where the amine group is added to PRPP128,134. Thus, the cluster is a structural element comparable to that of the Zn(II)(Cys)4 site in, for example, zinc fingers, as it connects distant cysteine residues (Fig. 5d), for example, in B. subtilis PPAT CX145CX76CX2C. Oxidation of the cluster leads to its disintegration and loss of enzyme activity133. One of the inorganic sulfides of the cluster is solvent-exposed via a shallow funnel of 7-Å length and 7-Å entry diameter (Fig. 5d)128. Therefore, similar to the [4Fe-4S] cluster of RSAD2, the [4Fe-4S] cluster in PPAT can act as lockable switch to turn activity off as a function of the concentrations of NO and H2O2 (Fig. 5a). Formation of holo-PPAT and regeneration of the activity of oxidatively damaged enzyme requires the CIA targeting complex135.
Redox sensing
A universal outcome of the innate immune response to viral infections is inflammation, a hallmark of which is excessive production of reactive oxygen or nitrogen species like H2O2 and NO (refs. 31,32) and hypoxia (Fig. 2)33. These changes are sensed by [FeS] proteins modulating iron homeostasis and mitophagy (Fig. 6a). Iron homeostasis is at the crossroads of viral infection and the immune response136,137,138, and mitophagy is a positive regulator of the innate immune response139. The [FeS] proteins involved in these processes are IRP1140,141, FBXL5142 and mitoNEET (Fig. 6a)143. First, IRP1 exists in two forms: holo-protein containing a solvent-exposed [4Fe-4S](Cys)3 cluster and apo-protein lacking the cluster (Fig. 6b). Whereas holo-IRP1 is in a closed state, apo-IRP1 adopts an open state (Fig. 6b) and interacts with specific nucleotide sequences, commonly known as iron-responsive elements (IREs), in the untranslated regions of the mRNAs of ferritin and transferrin receptor (TfR)140,141. Second, FBXL5 harbours a [2Fe-2S](Cys)4 cluster with a solvent-exposed cysteine residue (Fig. 6c)142. Under normal O2 levels, the cluster is in the oxidized state, [2Fe-2S]2+, and is able to organize the C-terminal loop of FBXL5 that is responsible for recruiting IRP2 (iron-responsive element-binding protein 2; Fig. 6c)142. This protein is not an iron–sulfur binding protein, but, like IRP1, can bind to IREs. Finally, mitoNEET is a homodimer and each monomer contains a redox-active [2Fe-2S](Cys)3(His) cluster (Fig. 6d)143. The histidine and a cysteine residue coordinating the [2Fe-2S] cluster are exposed to the solvent (Fig. 6d)144,145. The [2Fe-2S] cluster of mitoNEET has a midpoint redox potential of ~0 mV (pH 7.0)146, which is well above the cytosolic redox potential, that is ~−325 mV (pH 7.0)147. Thus, under normal physiological conditions mitoNEET is reduced. The cluster is stable in the presence of NO (ref. 148) and is reversibly oxidized by H2O2 (ref. 149).
a, A summary of the roles of the [FeS] clusters of IRP1, FBXL5 and mitoNEET in the regulation of iron homeostasis and mitophagy in response to the cellular levels of NO, H2O2 and O2 (feedforward). b, The [4Fe-4S](Cys)3 cluster of IRP1 acts as a conformational switch: holo-IRP1 (PDB 2B3X) has a closed conformation (top) and cannot bind to IRE, whereas apo-IRP1 has an open conformation and binds to IRE (PDB 3SN2). c, FBXL5 has a [2Fe-2S](Cys)4 cluster, and one of the cysteine residues is solvent-exposed (dashed arrow; PDB 6VCD). When the [2Fe-2S](Cys)4 cluster of FBXL5 is oxidized, the protein binds to IRP2. d, Each monomer of the homodimeric mitoNEET has a redox-active [2Fe-2S](Cys)3(His) cluster (PDB 3EW0). The histidine and a cysteine residue coordinating to the cluster are solvent-exposed (dashed arrow). Ub, ubiquitin.
The [FeS] clusters of IRP1 and FBXL5 modulate iron homeostasis during inflammation (Fig. 6a). The cellular level of IRP1 is affected by inflammation and the synthesis of NO (ref. 150), which degrades the [4Fe-4S] cluster, presumably via a mechanism similar to that proposed for NO sensory protein NsrR (Fig. 1e)13. At the same time, the hypoxic conditions caused by inflammation result in a reduced [2Fe-2S]1+ cluster in FBXL5, blocking its ability to recruit and mark IRP2 for degradation (Fig. 6a)142. Consequently, apo-IRP1 and IRP2 bind to the IRE at the 5′-untranslated region of ferritin mRNA, blocking its translation (Fig. 6a), and bind to the 3′-untranslated regions of TfR mRNA, stabilizing the mRNA and inducing translation. Overexpression of TfR and a reduction in the cellular level of ferritin modulates the immune response to viral infections in two general ways (Fig. 6a). First, it causes an increase in the intracellular level of available iron (that is, not stored in ferritin). Iron is a critical component of many cofactors, for example haem, mononuclear/dinuclear iron (di-iron) sites and [FeS] clusters. The proteins carrying these bio-inorganic cofactors perform many cellular functions essential for the immune response and viral replication, for example, [FeS] proteins such as RNA Pol-III, Elp3, ABCE1 and RSAD2 and di-iron ribonucleotide reductase of humans and some viruses, such as human cytomegalovirus (HCMV)151. Second, an increase in the expression of TfR can support entry of some viruses to host cells (Fig. 6a)138,152.
On the other hand, available data suggest that the [FeS] cluster of mitoNEET senses the cellular level of H2O2 to regulate mitophagy and the innate immune response (Fig. 6a). An elevated cellular level of H2O2 during inflammation, caused by the activity of enzymes such as NADPH oxidases (NOXs) and/or superoxide dismutase (SOD)153, leads to oxidation of the [2Fe-2S] cluster of mitoNEET. Oxidized mitoNEET interacts with voltage-dependent anion channels (VDACs) and abolishes their redox-gating function in crosstalk between the mitochondria and cytosol154. Consequently, the mitochondrial respiration rate is suppressed (Fig. 6a). These observations are consistent with previous reports showing that loss of mitoNEET may increase the mitochondrial respiration rate155. A decrease in mitochondrial respiration rate induced by the interaction of oxidized mitoNEET with VDACs will lead to a lowering of the mitochondrial membrane potential (ΔΨm). A reduction in the mitochondrial respiration and membrane potential activates the tensin homologue (PTEN)-induced putative kinase 1 (PINK1; Fig. 6a), which phosphorylates and recruits the E3 ubiquitin protein ligase (PARKIN). The latter induces ubiquitination of target proteins and activates mitophagy156,157. Recent studies also support a function of mitoNEET in linking the innate immune response and cellular levels of H2O2 to the function of the CIA machinery. Specifically, the complex of anamorsin and NADPH-dependent diflavin oxidoreductase 1 (Ndor1) protein, which acts as an electron donor in the CIA machinery158, interacts with mitoNEET. This complex reduces the [2Fe-2S] clusters of mitoNEET159, and the clusters are transferred to anamorsin160. These activities will prevent mitoNEET-induced mitophagy.
Concluding remarks and perspective
We have discussed the essential roles of [FeS] clusters in all steps of the innate immune response (Fig. 2), namely recognizing viral components, activating the expression of proinflammatory cytokines and ISG proteins, the functioning of the ISG protein RSAD2 and metabolic enzymes like PPAT, and sensing cellular levels of NO, H2O2 and O2. We have shown that some of the proteins involved, such as ABCE1 and PPAT, also support viral reproduction. We have highlighted that there are two common features associated with the [FeS] clusters in the proteins involved: (1) the clusters confer structural stability on the proteins or protein complexes and (2) they are solvent-exposed and thus accessible by NO, H2O2 or O2. We discussed that these features in [FeS] proteins like mitoNEET, IRP1 and FBXL5 enable a feedforward mechanism to modulate the innate immune response, while in other proteins like RNA Pol-III, Elp3, ABCE1, Dph1–Dph2, RSAD2 and PPAT they provide a feedback loop to block an excessive immune response. In the feedback mechanism, the increasing levels of NO and H2O2, which are caused by the innate immune response, are sensed by the solvent-exposed [FeS] clusters. Consequently, the clusters are degraded, inactivating the proteins and preventing an excessive immune response and the production of toxic levels of NO and H2O2 or metabolites like ddhCTP, which may have cytotoxicity119,123. This period of suppression continues until the cellular levels of NO and H2O2 are decreased to a low level and the [FeS] biogenesis machinery restores the activity of the proteins. Recent studies support the proposed feedback mechanism and the action of reactive nitrogen or oxygen species like H2O2 in preventing an excessive innate immune response161,162. The function of [FeS] clusters as biological ‘fuses’ in enzymes like ABCE1 and PPAT, the activities of which not only enable the innate immune system to restrict viral replication but also support the reproduction of viral particles, will also limit the viral replication process.
We have discussed examples of some host cell [FeS] proteins as inhibitors and catalysts of the viral replication process. The roles of many [FeS] proteins (such as Dph1–Dph2, Elp3, ABCE1, mitoNEET, RSAD2 and PPAT) in the innate immune response are still not well understood, and [FeS] clusters are emerging as essential components of some viral proteins, including RdRp of SARS-CoV-225. Many questions need to be answered. How is the [FeS] biogenesis machinery regulated in response to viral infection and during activation of the immune system? What are the critical levels of reactive oxygen and nitrogen species at which [FeS] clusters in different proteins become sensitive? What are the kinetics laws governing the reaction of [FeS] clusters with NO and H2O2 and those of [FeS] insertion into target proteins, and how do the different steps combine to regulate the innate immune response? What is the cellular protein supplying electrons for activating enzymes like Elp3 and RSAD2? How exactly do viral proteins gain their [FeS] clusters? Is the ability of SARS-CoV-2 RdRps to catalyse replication of viral RNA either with [FeS] clusters or Zn(II)25 an adaptation mechanism to cope with the increasing cellular levels of NO and H2O2 caused by the innate immune response? Future studies should help to address these questions, revealing fundamental insights into the function of [FeS] clusters in the innate immune system and viral replication processes. These studies will identify new drug targets to fight a wide range of RNA and DNA viruses. Additionally, as the links between the antiviral innate immune response and cancer or neurodegeneration become clearer163,164, and because mutations in proteins involved in the [FeS] biogenesis machinery such as frataxin165 can lead to neurodegenerative diseases like Friedreich’s ataxia and oxidative stress166, we expect that a better understanding of the roles of [FeS] clusters in the innate immune response will help in understanding the molecular details connecting viral infections and inflammation to cancer and neurodegenerative diseases.
Change history
05 April 2022
In the version of this article initially published online, various labels dropped out of the HTML version of Fig. 3. The errors have been corrected in the online version of the article.
References
Beinert, H., Holm, R. H. & Munck, E. Iron-sulfur clusters: nature’s modular, multipurpose structures. Science 277, 653–659 (1997).
Cracknell, J. A., Vincent, K. A. & Armstrong, F. A. Enzymes as working or inspirational electrocatalysts for fuel cells and electrolysis. Chem. Rev. 108, 2439–2461 (2008).
Lubitz, W., Ogata, H., Rudiger, O. & Reijerse, E. Hydrogenases. Chem. Rev. 114, 4081–4148 (2014).
Nelson, N. & Ben-Shem, A. The complex architecture of oxygenic photosynthesis. Nat. Rev. Mol. Cell Biol. 5, 971–982 (2004).
Verkhovskaya, M. & Bloch, D. A. Energy-converting respiratory complex I: on the way to the molecular mechanism of the proton pump. Int. J. Biochem. Cell Biol. 45, 491–511 (2013).
Broderick, J. B., Duffus, B. R., Duschene, K. S. & Shepard, E. M. Radical S-adenosylmethionine enzymes. Chem. Rev. 114, 4229–4317 (2014).
Nicolet, Y. Structure–function relationships of radical SAM enzymes. Nat. Catal. 3, 337–350 (2020).
Castro, L., Tortora, V., Mansilla, S. & Radi, R. Aconitases: non-redox iron-sulfur proteins sensitive to reactive species. Acc. Chem. Res. 52, 2609–2619 (2019).
Lancey, C. et al. Structure of the processive human Pol δ holoenzyme. Nat. Commun. 11, 1109 (2020).
Netz, D. J. A. et al. Eukaryotic DNA polymerases require an iron–sulfur cluster for the formation of active complexes. Nat. Chem. Biol. 8, 125–132 (2012).
Crack, J. C., Green, J., Thomson, A. J. & Le Brun, N. E. Iron-sulfur clusters as biological sensors: the chemistry of reactions with molecular oxygen and nitric oxide. Acc. Chem. Res. 47, 3196–3205 (2014).
Martinez, M. T. P. et al. Mechanisms of iron- and O2-sensing by the [4Fe-4S] cluster of the global iron regulator RirA. eLife 8, e47804 (2019).
Volbeda, A. et al. Crystal structures of the NO sensor NsrR reveal how its iron–sulfur cluster modulates DNA binding. Nat. Commun. 8, 15052 (2017).
Crack, J. C., Hamilton, C. J. & Le Brun, N. Mass spectrometric detection of iron nitrosyls, sulfide oxidation and mycothiolation during nitrosylation of the NO sensor [4Fe–4S] NsrR. Chem. Commun. 54, 5992–5995 (2018).
Volbeda, A., Darnault, C., Renoux, O., Nicolet, Y. & Fontecilla-Camps, C. The crystal structure of the global anaerobic transcriptional regulator FNR explains its extremely fine-tuned monomer-dimer equilibrium. Sci. Adv. 1, e1501086 (2015).
McCarthy, E. L. & Booker, S. J. Destruction and reformation of an iron-sulfur cluster during catalysis by lipoyl synthase. Science 358, 373–377 (2017).
Braymer, J. J. & Lill, R. Iron-sulfur cluster biogenesis and trafficking in mitochondria. J. Biol. Chem. 292, 12754–12763 (2017).
Cardenas-Rodriguez, M., Chatzi, A. & Tokatlidis, K. Iron-sulfur clusters: from metals through mitochondria biogenesis to disease. J. Biol. Inorg. Chem. 23, 509–520 (2018).
Lill, R. & Freibert, S.-A. Mechanisms of mitochondrial iron-sulfur protein biogenesis. Annu. Rev. Biochem. 89, 471–499 (2020).
Ciofi-Baffoni, S., Nasta, V. & Banci, L. Protein networks in the maturation of human iron-sulfur proteins. Metallomics 10, 49–72 (2018).
Pandey, A. K., Pain, J., Dancis, A. & Pain, D. Mitochondria export iron-sulfur and sulfur intermediates to the cytoplasm for iron-sulfur cluster assembly and tRNA thiolation in yeast. J. Biol. Chem. 294, 9489–9502 (2019).
Jones, J. E., Le Sage, V. & Lakdawala, S. S. Viral and host heterogeneity and their effects on the viral life cycle. Nat. Rev. Microbiol. 19, 272–282 (2021).
Martin, D. et al. The rotavirus nonstructural protein NSP5 coordinates a [2Fe-2S] iron-sulfur cluster that modulates interaction to RNA. FASEB J. 27, 1074–1083 (2013).
Tsang, S. H. et al. The oncogenic small tumor antigen of Merkel cell polyomavirus is an iron-sulfur cluster protein that enhances viral DNA replication. J. Virol. 90, 1544–1556 (2016).
Maio, N. et al. Fe-S cofactors in the SARS-CoV-2 RNA-dependent RNA polymerase are potential antiviral targets. Science 373, 236–241 (2021).
Abe, T., Marutani, Y. & Shoji, I. Cytosolic DNA‐sensing immune response and viral infection. Microbiol. Immunol. 63, 51–64 (2019).
Kawai, T. & Akira, S. Innate immune recognition of viral infection. Nat. Immunol. 7, 131–137 (2006).
Schneider, W. J., Chevillotte, M. D. & Rice, C. M. Interferon-stimulated genes: a complex web of host defenses. Annu. Rev. Immunol. 32, 513–545 (2014).
Schoggins, J. W. Interferon-stimulated genes: what do they all do? Annu. Rev. Virol. 6, 567–584 (2019).
Sadler, A. J. & Williams, B. R. G. Interferon-inducible antiviral effectors. Nat. Rev. Immunol. 8, 559–568 (2008).
Schwarz, K. B. Oxidative stress during viral infection: a review. Free Radic. Biol. Med. 21, 641–649 (1996).
Rouse, B. T. & Sehrawat, S. Immunity and immunopathology to viruses: what decides the outcome? Nat. Rev. Immunol. 10, 514–526 (2010).
Taylor, C. T. & Colgan, S. P. Regulation of immunity and inflammation by hypoxia in immunological niches. Nat. Rev. Immunol. 17, 774–785 (2017).
Puig, S., Ramos-Alonso, L., Maria Romero, A. & Martinez-Pastor, M. T. The elemental role of iron in DNA synthesis and repair. Metallomics 9, 1483–1500 (2017).
Ter Beek, J. et al. Structural evidence for an essential Fe-S cluster in the catalytic core domain of DNA polymerase ϵ.Nucleic Acids Res 47, 5712–5722 (2019).
Girbig, M. et al. Cryo-EM structures of human RNA polymerase III in its unbound and transcribing states. Nat. Struct. Mol. Biol. 28, 210–219 (2021).
Wang, Q. et al. Structural insights into transcriptional regulation of human RNA polymerase III. Nat. Struct. Mol. Biol. 28, 220–227 (2021).
Dauden, M. I. et al. Molecular basis of tRNA recognition by the elongator complex. Sci. Adv. 5, eaaw2326 (2019).
Dauden, M. I., Jaciuk, M., Muller, C. W. & Glatt, S. Structural asymmetry in the eukaryotic elongator complex. FEBS Lett. 592, 502–515 (2018).
Svejstrup, J. Q. Elongator complex: how many roles does it play? Curr. Opin. Cell Biol. 19, 331–336 (2007).
Lehman, I. R. & Kaguni, L. S. DNA polymerase alpha. J. Biol. Chem. 264, 4265–4268 (1989).
Vannini, A. et al. Molecular basis of RNA polymerase III transcription repression by Maf1. Cell 143, 59–70 (2010).
Shilatifard, A., Conaway, R. C. & Conaway, J. W. The RNA polymerase II elongation complex. Annu. Rev. Biochem. 72, 693–715 (2003).
Paraskevopoulou, C., Fairhurst, S. A., Lowe, D. J., Brick, P. & Onesti, S. The elongator subunit Elp3 contains a Fe4S4 cluster and binds S‐adenosylmethionine. Mol. Microbiol. 59, 795–806 (2006).
Wittschieben, B. et al. A novel histone acetyltransferase is an integral subunit of elongating RNA polymerase II holoenzyme. Mol. Cell 4, 123–128 (1999).
Greenwood, C., Selth, L. A., Dirac-Svejstrup, A. B. & Svejstrup, J. Q. An iron-sulfur cluster domain in Elp3 important for the structural integrity of elongator. J. Biol. Chem. 284, 141–149 (2009).
Winkler, G. S., Kristjuhan, A., Erdjument-Bromage, H., Tempst, P. & Svejstrup, J. Q. Elongator is a histone H3 and H4 acetyltransferase important for normal histone acetylation levels in vivo. Proc. Natl Acad. Sci. USA 99, 3517–3522 (2002).
Keating, S. T., van Diepen, J. A., Riksen, N. P. & El-Osta, A. Epigenetics in diabetic nephropathy, immunity and metabolism. Diabetologia 61, 6–20 (2018).
Selvadurai, K., Wang, P., Seimetz, J. & Huang, R. H. Archaeal Elp3 catalyzes tRNA wobble uridine modification at C5 via a radical mechanism. Nat. Chem. Biol. 10, 810–812 (2014).
Glatt, S. et al. Structural basis for tRNA modification by Elp3 from Dehalococcoides mccartyi. Nat. Struct. Mol. Biol. 23, 794–802 (2016).
Lin, T.-Y. et al. The elongator subunit Elp3 is a non-canonical tRNA acetyltransferase. Nat. Commun. 10, 625 (2019).
Wang, T. S.-E., Wong, S. W. & Korn, D. Human DNA polymerase α: predicted functional domains and relationships with viral DNA polymerases. FASEB J. 3, 14–21 (1989).
Tang, L., Sheraz, M., McGrane, M., Chang, J. & Guo, J.-T. DNA polymerase alpha is essential for intracellular amplification of hepatitis B virus covalently closed circular DNA. PLoS Pathog. 15, e1007742 (2019).
Gordon, D. E. et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 583, 459–468 (2020).
Chen, Z. et al. Interactomes of SARS-CoV-2 and human coronaviruses reveal host factors potentially affecting pathogenesis.EMBO J. 40, e107776 (2021).
Starokadomskyy, P. et al. DNA polymerase-α regulates the activation of type I interferons through cytosolic RNA:DNA synthesis. Nat. Immunol. 17, 495–504 (2016).
Carter-Timofte, M. E., Hansen, A. F., Christiansen, M., Paludan, S. R. & Mogensen, T. H. Mutations in RNA polymerase III genes and defective DNA sensing in adults with varicella-zoster virus CNS infection. Genes Immun. 20, 214–223 (2019).
Jarrous, N. & Rouvinski, A. RNA polymerase III and antiviral innate immune response. Transcription 12, 1–11 (2021).
Chiu, Y.-H., MacMilan, J. B. & Chen, Z. J. RNA polymerase III detects cytosolic DNA and induces type I interferons through the RIG-I pathway. Cell 138, 576–591 (2009).
Ablasser, A. et al. RIG-I-dependent sensing of poly(dA:dT) through the induction of an RNA polymerase III-transcribed RNA intermediate. Nat. Immunol. 10, 1065–1072 (2009).
Graczyk, D., White, R. J. & Ryan, K. M. Involvement of RNA polymerase III in immune responses. Mol. Cell. Biol. 35, 1848–1859 (2015).
Ramanathan, A. et al. A mutation in POLR3E impairs antiviral immune response and RNA polymerase III. Proc. Natl Acad. Sci. USA 117, 22113–22121 (2020).
Reverendo, M. et al. Polymerase III transcription is necessary for T cell priming by dendritic cells. Proc. Natl Acad. Sci. USA 116, 22721–22729 (2019).
Tailor, P. et al. The feedback phase of type I interferon induction in dendritic cells requires interferon regulatory factor 8. Immunity 27, 228–239 (2007).
Parekh, B. & Maniatis, T. Virus infection leads to localized hyperacetylation of histones H3 and H4 at the IFN-β promoter. Mol. Cell 3, 125–129 (1999).
Wu, M. et al. Lysine-14 acetylation of histone H3 in chromatin confers resistance to the deacetylase and demethylase activities of an epigenetic silencing complex. eLife 7, e37231 (2018).
Zhang, Q. & Cao, X. Epigenetic regulation of the innate immune response to infection. Nat. Rev. Immunol. 19, 417–432 (2019).
Otero, G. et al. Elongator, a multisubunit component of a novel RNA polymerase II holoenzyme for transcriptional elongation. Mol. Cell 3, 109–118 (1999).
Walker, J. et al. Role of elongator subunit Elp3 in Drosophila melanogaster larval development and immunity. Genetics 187, 1067–1075 (2011).
DeFraia, C. T., Wang, Y., Yao, J. & Mou, Z. Elongator subunit 3 positively regulates plant immunity through its histone acetyltransferase and radical S-adenosylmethionine domains. BMC Plant Biol. 13, 102 (2013).
Lemaitre, P. et al. Loss of the transfer RNA wobble uridine-modifying enzyme Elp3 delays T cell cycle entry and impairs T follicular helper cell responses through deregulation of Atf4. J. Immunol. 206, 1077–1087 (2021).
Rosu, A. et al. Loss of tRNA-modifying enzyme Elp3 activates a p53-dependent antitumor checkpoint in hematopoiesis. J. Exp. Med. 218, e20200662 (2021).
Zhu, C., Sun, B., Nie, A. & Zhou, Z. The tRNA-associated dysregulation in immune responses and immune diseases. Acta Physiol. 228, e13391 (2020).
Su, X., Lin, Z. & Lin, H. The biosynthesis and biological function of diphthamide. Crit. Rev. Biochem. Mol. Biol. 48, 515–521 (2013).
Bisbal, C., Marinand, C., Silhol, M., Lebleu, B. & Salehzada, T. Cloning and characterization of a RNAse L inhibitor. A new component of the interferon-regulated 2-5A pathway. J. Biol. Chem. 270, 13308–13317 (1995).
Hawer, H. et al. Importance of diphthamide modified EF2 for translational accuracy and competitive cell growth in yeast. PLoS ONE 13, e0205870 (2018).
Stahl, S. et al. Loss of diphthamide pre-activates NF-κB and death receptor pathways and renders MCF7 cells hypersensitive to tumor necrosis factor. Proc. Natl Acad. Sci. USA 112, 10732–10737 (2015).
Zhang, Y. et al. Diphthamide biosynthesis requires an organic radical generated by an iron–sulphur enzyme. Nature 465, 891–896 (2010).
Dong, M. et al. The asymmetric function of Dph1-Dph2 heterodimer in diphthamide biosynthesis. J. Biol. Inorg. Chem. 24, 777–782 (2019).
Dong, M. et al. Organometallic and radical intermediates reveal mechanism of diphthamide biosynthesis. Science 359, 1247–1250 (2018).
Winzeler, E. A. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999).
Anderson, D. E. et al. Comparative loss-of-function screens reveal ABCE1 as an essential cellular host factor for efficient translation of Paramyxoviridae and Pneumoviridae. mBio 10, e00826-19 (2019).
Gavin, A.-C. et al. Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature 415, 141–147 (2002).
Dong, J. et al. The essential ATP-binding cassette protein RLI1 functions in translation by promoting preinitiation complex assembly. J. Biol. Chem. 279, 42157–42168 (2004).
Chen, Z.-Q. et al. The essential vertebrate ABCE1 protein interacts with eukaryotic initiation factors. J. Biol. Chem. 281, 7452–7457 (2006).
Barthelme, D. et al. Structural organization of essential iron-sulfur clusters in the evolutionarily highly conserved ATP-binding cassette protein ABCE1. J. Biol. Chem. 282, 14598–14607 (2007).
Karcher, A., Schele, A. & Hopfner, K.-P. X-ray structure of the complete ABC enzyme ABCE1 from Pyrococcus abyssi. J. Biol. Chem. 283, 7962–7971 (2008).
Alhebshi, A., Sideri, T. C., Holland, S. L. & Avery, S. V. The essential iron-sulfur protein Rli1 is an important target accounting for inhibition of cell growth by reactive oxygen species. Mol. Biol. Cell 23, 3582–3590 (2012).
Savelsbergh, A., Rodnina, M. V. & Wintermeyer, W. Distinct functions of elongation factor G in ribosome recycling and translocation. RNA 15, 772–780 (2009).
Pisarev, A. V. et al. The role of ABCE1 in eukaryotic posttermination ribosomal recycling. Mol. Cell 37, 196–210 (2010).
Preis, A. et al. Cryoelectron microscopic structures of eukaryotic translation termination complexes containing eRF1-eRF3 or eRF1-ABCE1. Cell Rep. 8, 59–65 (2014).
Young, D. J., Guydosh, N. R., Zhang, F., Hinnebusch, A. G. & Green, R. Rli1/ABCE1 recycles terminating ribosomes and controls translation reinitiation in 3′UTRs in vivo. Cell 162, 872–884 (2015).
Heuer, A. et al. Structure of the 40S–ABCE1 post-splitting complex in ribosome recycling and translation initiation. Nat. Struct. Mol. Biol. 24, 453–460 (2017).
Kratzat, H. et al. A structural inventory of native ribosomal ABCE1‐43S pre‐initiation complexes. EMBO J. 40, e105179 (2021).
Barthelme, D. et al. Ribosome recycling depends on a mechanistic link between the FeS cluster domain and a conformational switch of the twin-ATPase ABCE1. Proc. Natl Acad. Sci. USA 108, 3228–3233 (2011).
Nurenberg-Goloub, E. et al. Molecular analysis of the ribosome recycling factor ABCE1 bound to the 30S post-splitting complex. EMBO J. 39, e103788 (2020).
Paul, V. D. et al. The deca-GX3 proteins Yae1-Lto1 function as adaptors recruiting the ABC protein Rli1 for iron-sulfur cluster insertion. eLife 4, e08231 (2015).
Prusty, N. R., Camponeschi, F., Ciofi-Baffoni, S. & Banci, L. The human YAE1-ORAOV1 complex of the cytosolic iron-sulfur protein assembly machinery binds a [4Fe-4S] cluster. Inorg. Chim. Acta 518, 120252 (2021).
Dong, M. et al. Dph3 is an electron donor for Dph1-Dph2 in the first step of eukaryotic diphthamide biosynthesis. J. Am. Chem. Soc. 136, 1754–1757 (2014).
Ghosh, S. & Marsh, E. N. G. Viperin: an ancient radical-SAM enzyme finds its place in modern cellular metabolism and innate immunity. J. Biol. Chem. 295, 11513–11528 (2020).
Ebrahimi, K. H. A unifying view of the broad-spectrum antiviral activity of RSAD2 (viperin) based on its radical-SAM chemistry. Metallomics 10, 539–552 (2018).
Hinson, E. R. & Cresswell, P. The N-terminal amphipathic α-helix of viperin mediates localization to the cytosolic face of the endoplasmic reticulum and inhibits protein secretion. J. Biol. Chem. 284, 4705–4712 (2009).
Hinson, E. R. & Cresswell, P. The antiviral protein, viperin, localizes to lipid droplets via its N-terminal amphipathic α-helix. Proc. Natl Acad. Sci. USA 106, 20452–20457 (2009).
Fitzgerald, K. A. The interferon inducible gene: viperin. J. Interf. Cytokine Res. 31, 131–135 (2011).
Severa, M., Coccia, E. M. & Fitzgerald, K. A. Toll-like receptor-dependent and -independent viperin gene expression and counter-regulation by PRDI-binding factor-1/BLIMP1. J. Biol. Chem. 281, 26188–26195 (2006).
Seo, J.-Y., Yaneva, R., Hinson, E. R. & Cresswell, P. Human cytomegalovirus directly induces the antiviral protein viperin to enhance infectivity. Science 332, 1093–1097 (2011).
Jang, J.-S. et al. Rsad2 is necessary for mouse dendritic cell maturation via the IRF7-mediated signaling pathway. Cell Death Dis. 9, 823 (2018).
Wiedemann, G. M., Geary, C. D., Lau, C. M. & Sun, J. C. Cutting edge: STAT1-mediated epigenetic control of Rsad2 promotes clonal expansion of antiviral NK cells. J. Immunol. 205, 21–25 (2020).
Qiu, L.-Q., Cresswell, P. & Chin, K.-C. Viperin is required for optimal Th2 responses and T-cell receptor-mediated activation of NF-kB and AP-1. Blood 113, 3520–3529 (2009).
Eom, J. et al. Intrinsic expression of viperin regulates thermogenesis in adipose tissues. Proc. Natl Acad. Sci. USA 116, 17419–17428 (2019).
Duschene, K. S. & Broderick, J. B. The antiviral protein viperin is a radical SAM enzyme. FEBS Lett. 584, 1263–1267 (2010).
Ebrahimi, K. H. et al. The radical‐SAM enzyme viperin catalyzes reductive addition of a 5′‐deoxyadenosyl radical to UDP‐glucose in vitro. FEBS Lett. 591, 2394–2405 (2017).
Fenwick, M. K., Li, Y., Cresswell, P., Modis, Y. & Ealick, S. E. Structural studies of viperin, an antiviral radical SAM enzyme. Proc. Natl Acad. Sci. USA 114, 6806–6811 (2017).
Gizzi, A. S. et al. A naturally occurring antiviral ribonucleotide encoded by the human genome. Nature 558, 610–614 (2018).
Fenwick, M. K., Su, D., Dong, M., Lin, H. & Ealick, S. E. Structural basis of the substrate selectivity of viperin. Biochemistry 59, 652–662 (2020).
Ebrahimi, K. H., Rowbotham, J., McCullagh, J. & James, W. S. Mechanism of diol dehydration by a promiscuous radical‐SAM enzyme homologue of the antiviral enzyme viperin (RSAD2). ChemBioChem 21, 1605–1612 (2020).
Qi, Z. et al. Targeting viperin improves diet-induced glucose intolerance but not adipose tissue inflammation. Oncotarget 8, 101418–101436 (2017).
Dumbrepatil, A. B., Zegalia, K. A., Sajja, K., Kennedy, R. T. & Marsh, E. N. G. Targeting viperin to the mitochondrion inhibits the thiolase activity of the trifunctional enzyme complex. J. Biol. Chem. 295, 2839–2849 (2020).
Ebrahimi, K. H. et al. Viperin, through its radical-SAM activity, depletes cellular nucleotide pools and interferes with mitochondrial metabolism to inhibit viral replication. FEBS Lett. 594, 1624–1630 (2020).
Ebrahimi, K. H., Vowles, J., Browne, C., McCullagh, J. & James, W. S. ddhCTP produced by the radical‐SAM activity of RSAD2 (viperin) inhibits the NAD+‐dependent activity of enzymes to modulate metabolism. FEBS Lett. 594, 1631–1644 (2020).
Ebrahimi, K. H., Gilbert-Jaramillo, J., James, W. S. & McCullagh, J. S. O. Interferon-stimulated gene products as regulators of central carbon metabolism. FEBS J. 288, 3715–3726 (2021).
Haldar, S., Paul, S., Joshi, N., Dasgupta, A. & Chattopadhyay, K. The presence of the iron-sulfur motif is important for the conformational stability of the antiviral protein, viperin. PLoS ONE 7, e31797 (2012).
Torrance, C. J., Agrawal, V., Vogelstein, B. & Kinzler, K. W. Use of isogenic human cancer cells for high-throughput screening and drug discovery. Nat. Biotechnol. 19, 940–945 (2001).
Upadhyay, A. S. et al. Cellular requirements for iron-sulfur cluster insertion into the antiviral radical SAM protein viperin. J. Biol. Chem. 292, 13879–13889 (2017).
Viola, A., Munari, F., Sanchez-Rodriguez, R., Scolaro, T. & Categna, A. The metabolic signature of macrophage responses. Front. Immunol. 10, 1462 (2019).
Tian, X. et al. Metabolomic analysis of influenza A virus A/WSN 1933 (H1N1) infected A549 cells during first cycle of viral replication. Viruses 11, 1007 (2019).
Chandler, J. D. et al. Metabolic pathways of lung inflammation revealed by high-resolution metabolomics (HRM) of H1N1 influenza virus infection in mice. Am. J. Physiol. Regul. Integr. Comp. Physiol. 311, R906–R916 (2016).
Smith, J. L. et al. Structure of the allosteric regulatory enzyme of purine biosynthesis. Science 264, 1427–1433 (1994).
Muchmore, C. R. A., Krahn, J. M., Smith, J. L., Kim, J. H. & Zalkin, H. Crystal structure of glutamine phosphoribosylpyrophosphate amidotransferase from Escherichia coli. Protein Sci. 7, 39–51 (1998).
Cao, X. et al. Crystal structure of the chloroplastic glutamine phosphoribosylpyrophosphate amidotransferase GPRAT2 from Arabidopsis thaliana. Front. Plant Sci. 11, 157 (2020).
Averill, B. A. et al. Evidence for a tetranuclear iron-sulfur center in glutamine phosphoribosylpyrophosphate amidotransferase from Bacillus subtilis. J. Biol. Chem. 255, 6007–6010 (1980).
Onate, Y. A., Vollmer, S. J., Switzer, R. L. & Johnson, M. K. Spectroscopic characterization of the iron-sulfur cluster in Bacillus subtilis glutamine phosphoribosylpyrophosphate amidotransferase. J. Biol. Chem. 264, 18386–18391 (1989).
Vollmer, S. J., Switzer, R. L. & Debrunner, P. G. Oxidation-reduction properties of the iron-sulfur cluster in Bacillus subtilis glutamine phosphoribosylpyrophosphate amidotransferase. J. Biol. Chem. 258, 14284–14293 (1983).
Chen, S. et al. Mechanism of the synergistic end-product regulation of Bacillus subtilis glutamine phosphoribosylpyrophosphate amidotransferase by nucleotide. Biochemistry 36, 10718–10726 (1997).
Stehling, O. et al. Human CIA2A-FAM96A and CIA2B-FAM96B integrate iron homeostasis and maturation of different subsets of cytosolic-nuclear iron-sulfur proteins. Cell Metab. 18, 187–198 (2013).
Weinberg, E. D. Iron withholding: a defense against infection and neoplasia. Physiol. Rev. 64, 65–102 (1984).
Ganz, T. & Nemeth, E. Iron homeostasis in host defence and inflammation. Nat. Rev. Immunol. 15, 500–510 (2015).
Drakesmith, H. & Prentice, A. Viral infection and iron metabolism. Nat. Rev. Microbiol. 6, 541–552 (2008).
Cho, D.-H., Kim, J. K. & Jo, E.-K. Mitophagy and innate immunity in infection. Mol. Cells 43, 10–22 (2020).
Anderson, C. P., Shen, M., Eisenstein, R. S. & Leibold, E. A. Mammalian iron metabolism and its control by iron regulatory proteins. Biochim. Biophys. Acta 1823, 1468–1483 (2012).
Wallander, M. L., Leibold, E. A. & Eisenstein, R. S. Molecular control of vertebrate iron homeostasis by iron regulatory proteins. Biochim. Biophys. Acta 1763, 668–689 (2006).
Wang, H. et al. FBXL5 regulates IRP2 stability in iron homeostasis via an oxygen-responsive [2Fe2S] cluster. Mol. Cell 78, 31–41 (2020).
Wiley, S. E., Murphy, A. N., Ross, S. A., van der Geer, P. & Dixon, J. E. MitoNEET is an iron-containing outer mitochondrial membrane protein that regulates oxidative capacity. Proc. Natl Acad. Sci. USA 104, 5318–5323 (2007).
Hou, X. et al. Crystallographic studies of human MitoNEET. J. Biol. Chem. 282, 33242–33246 (2007).
Lin, J., Zhou, T., Ye, K. & Wang, J. Crystal structure of human MitoNEET reveals distinct groups of iron-sulfur proteins. Proc. Natl Acad. Sci. USA 104, 14640–14645 (2007).
Bak, D. W., Zuris, J. A., Paddock, M. L., Jennings, P. A. & Elliott, S. J. Redox characterization of the FeS protein MitoNEET and impact of thiazolidinedione drug binding. Biochemistry 48, 10193–10195 (2009).
Dooley, C. T. et al. Imaging dynamic redox changes in mammalian cells with green fluorescent protein indicators. J. Biol. Chem. 279, 22284–22293 (2004).
Cheng, Z., Landry, A. P., Wang, Y. & Ding, H. Binding of nitric oxide in CDGSH-type [2Fe-2S] clusters of the human mitochondrial protein Miner2. J. Biol. Chem. 292, 3146–3153 (2017).
Landry, A. P. & Ding, H. Redox control of human mitochondrial outer membrane protein MitoNEET [2Fe-2S] clusters by biological thiols and hydrogen peroxide. J. Biol. Chem. 289, 4307–4315 (2014).
Phillips, J. D., Kinikini, D. V., Yu, Y., Guo, B. & Leibold, E. A. Differential regulation of IRP1 and IRP2 by nitric oxide in rat hepatoma cells. Blood 87, 2983–2992 (1996).
Jordan, A. & Reichard, P. Ribonucleotide reductases. Annu. Rev. Biochem. 67, 71–98 (1998).
Radoshitzky, S. R. et al. Transferrin receptor 1 is a cellular receptor for New World haemorrhagic fever arenaviruses. Nature 446, 92–96 (2007).
Mittal, M., Rizwan, M., Tran, K., Reddy, S. P. & Malik, A. B. Reactive oxygen species in inflammation and tissue injury. Antioxid. Redox Signal. 20, 1126–1167 (2014).
Lipper, C. H. et al. Redox-dependent gating of VDAC by MitoNEET. Proc. Natl Acad. Sci. USA 116, 19924–19929 (2019).
Kusminski, C. M. et al. MitoNEET-driven alterations in adipocyte mitochondrial activity reveal a crucial adaptive process that preserves insulin sensitivity in obesity. Nat. Med. 18, 1539–1549 (2012).
Sun, Y., Vashisht, A. A., Tchieu, J., Wohlschlegel, J. A. & Dreier, L. Voltage-dependent anion channels (VDACs) recruit Parkin to defective mitochondria to promote mitochondrial autophagy. J. Biol. Chem. 287, 40652–40660 (2012).
Eiyama, A. & Okamoto, K. PINK1/Parkin-mediated mitophagy in mammalian cells. Curr. Opin. Cell Biol. 33, 95–101 (2015).
Banci, L. et al. Molecular view of an electron transfer process essential for iron-sulfur protein biogenesis. Proc. Natl Acad. Sci. USA 110, 7136–7141 (2013).
Camponeschi, F., Ciofi-Baffoni, S. & Banci, L. Anamorsin/Ndor1 complex reduces [2Fe–2S]-MitoNEET via a transient protein-protein interaction. J. Am. Chem. Soc. 139, 9479–9482 (2017).
Lipper, C. H. et al. Cancer-related NEET proteins transfer 2Fe-2S clusters to anamorsin, a protein required for cytosolic iron-sulfur cluster biogenesis. PLoS ONE 10, e0139699 (2015).
Timblin, G. A. et al. Mitohormesis reprogrammes macrophage metabolism to enforce tolerance. Nat. Metab. 3, 618–635 (2021).
Zarse, K. & Ristow, M. Mitochondrial ROS signals prevent excessive immune response. Nat. Metab. 3, 588–589 (2021).
Fuertes, M. B., Woo, S.-R., Burnett, B., Fu, Y.-X. & Gajewski, T. F. Type I interferon response and innate immune sensing of cancer. Trends Immunol. 34, 67–73 (2013).
Heneka, M. T., Kummer, M. P. & Latz, E. Innate immune activation in neurodegenerative disease. Nat. Rev. Immunol. 14, 463–477 (2014).
Prischi, F. et al. Structural bases for the interaction of frataxin with the central components of iron–sulphur cluster assembly. Nat. Commun. 1, 95 (2010).
Lupoli, F., Vannocci, T., Longo, G., Niccolai, N. & Pastore, A. The role of oxidative stress in Friedreich’s ataxia. FEBS Lett. 592, 718–727 (2017).
Acknowledgements
We dedicate this Review to the memory of Andrew Thomson, a true pioneer of iron–sulfur cluster biochemistry who inspired us all in different ways. We acknowledge support from the European COST action FeSBioNET. K.H.E. is grateful for support from the European Molecular Biology Organization (EMBO; ALTF 157-2015), the Edward Penley Abraham Research Fund (RF259) and Cancer Research UK (CRUK) Oxford (CRUKDF-0221-KHE). Y.N. thanks the Commissariat à l'Energie Atomique (CEA) for institutional support. N.E.L.B. thanks the Biotechnology and Biological Sciences Research Council (BBSRC; grant no. BB/S001018/1). S.C.-B. acknowledges support from the Fondazione Cassa di Risparmio di Firenze (grant no. CRF2018.0920).
Author information
Authors and Affiliations
Contributions
K.H.E. conceived the idea for this Review. K.H.E. wrote the manuscript with contributions from S.C.-B., P.-L.H., Y.N., N.E.L.B., W.R.H. and F.A.A.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemistry thanks Maxie Roessler and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Honarmand Ebrahimi, K., Ciofi-Baffoni, S., Hagedoorn, PL. et al. Iron–sulfur clusters as inhibitors and catalysts of viral replication. Nat. Chem. 14, 253–266 (2022). https://doi.org/10.1038/s41557-021-00882-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-021-00882-0
This article is cited by
-
Can iron chelators ameliorate viral infections?
BioMetals (2024)