Abstract
The symmetries of a crystal are notoriously uncorrelated to those of its constituent molecules. This symmetry breaking is typically thought to occur during crystallization. Here we demonstrate that one of the two symmetry elements of olanzapine crystals, an inversion centre, emerges in solute dimers extant in solution prior to crystallization. We combine time-resolved in situ scanning probe microscopy to monitor the crystal growth processes with all-atom molecular dynamics simulations. We show that crystals grow non-classically, predominantly by incorporation of centrosymmetric dimers. The growth rate of crystal layers exhibits a quadratic dependence on the solute concentration, characteristic of the second-order kinetics of the incorporation of dimers, which exist in equilibrium with a majority of monomers. We show that growth by dimers is preferred due to overwhelming accumulation of adsorbed dimers on the crystal surface, where it is complemented by dimerization and expedites dimer incorporation into growth sites.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The datasets generated during and/or analysed during the current study and the simulation and analysis codes used for these calculations are available with the manuscript files and/or from the corresponding authors upon reasonable request. Several hundred AFM files were used to determine step displacements and step velocities used in this work. Those needed to illustrate the presented results were output as images; they are included in the main and supplementary figures and are available in electronic format from the data repository of the University of Strathclyde at https://doi.org/10.15129/64d77d7f-d0e2-497e-84e5-3e201a677a75. Additional images are available from the corresponding authors upon reasonable request. Source data are provided with this paper.
References
Vainshtein, B. K. Fundamentals of Crystals. Symmetry, and Methods of Structural Crystallography 2nd edn (Springer, 1994).
Barker, T. V. Molecular and crystal symmetry. Nature 111, 632–633 (1923).
Gavezzotti, A. Are crystal strutures predictable? Acc. Chem. Res. 27, 309–314 (1994).
Landau, L. D. & Lifschitz, L. Statistical Physics, Part 1 3rd edn, Vol. 5 (Pergamon Press, 1977).
Braga, D., Grepioni, F. & Desiraju, G. R. Crystal engineering and organometallic architecture. Chem. Rev. 98, 1375–1406 (1998).
Bernstein, J. Polymorphism in Molecular Crystals (Oxford Univ. Press, 2007).
Stranski, I. N. Zur theorie des kristallwachstums. Z. Phys. Chem. 136, 259–278 (1928).
Blüh, O. Einige bei der Untersuchung von Kolloiden im Wechselfeld auftretende Erscheinungen. Kolloid-Zeitschrift 37, 267–270 (1925).
Burton, W. K., Cabrera, N. & Frank, F. C. The growth of crystals and equilibrium structure of their surfaces. Philos. Trans. R. Soc. Lond. A 243, 299–360 (1951).
Bennema, P. Analysis of crystal growth models for slightly supersaturated solutions. J. Cryst. Growth 1, 278–286 (1967).
Gliko, O. et al. A metastable prerequisite for the growth of lumazine synthase crystals. J. Am. Chem. Soc. 127, 3433–3438 (2005).
Li, D. et al. Direction-specific interactions control crystal growth by oriented attachment. Science 336, 1014–1018 (2012).
De Yoreo, J. J. et al. Crystallization by particle attachment in synthetic, biogenic and geologic environments. Science. 349, aaa6760 (2015).
Lupulescu, A. I. & Rimer, J. D. In situ imaging of silicalite-1 surface growth reveals the mechanism of crystallization. Science 344, 729–732 (2014).
Wawrzycka-Gorczyca, I., Borowski, P., Osypiuk-Tomasik, J., Mazur, L. & Koziol, A. Crystal structure of olanzapine and its solvates. Part 3. Two and three-component solvates with water, ethanol, butan-2-ol and dichloromethane. J. Mol. Struct. 830, 188–197 (2007).
Fulton, B. & Goa, K. L. Olanzapine. Drugs 53, 281–298 (1997).
Askin, S. et al. Olanzapine form IV: discovery of a new polymorphic form enabled by computed crystal energy landscapes. Cryst. Growth Des. 19, 2751–2757 (2019).
Bhardwaj, R. M. et al. Exploring the experimental and computed crystal energy landscape of olanzapine. Cryst. Growth Des. 13, 1602–1617 (2013).
Sun, Y. et al. Modeling olanzapine solution growth morphologies. Cryst. Growth Des. 18, 905–911 (2018).
Warzecha, M. et al. Direct observation of templated two-step nucleation mechanism during olanzapine hydrate formation. Cryst. Growth Des. 17, 6382–6393 (2017).
Chernov, A. A. Modern Crystallography III, Crystal Growth (Springer, 1984).
Gilmer, G. H., Ghez, R. & Cabrera, N. An analysis of combined volume and surface diffusion processes in crystal growth. J. Cryst. Growth 8, 79–93 (1971).
De Yoreo, J. J. & Vekilov, P. G. Principles of crystal nucleation and growth. Rev. Mineral. Geochem. 54, 57–93 (2003).
Warzecha, M., Safari, M. S., Florence, A. J. & Vekilov, P. G. Mesoscopic solute-rich clusters in olanzapine solutions. Cryst. Growth Des. 17, 6668–6676 (2017).
Jiang, Y. et al. Growth of organic crystals via attachment and transformation of nanoscopic precursors. Nat. Commun. 8, 15933 (2017).
Yau, S.-T., Thomas, B. R. & Vekilov, P. G. Molecular mechanisms of crystallization and defect formation. Phys. Rev. Lett. 85, 353–356 (2000).
Lovette, M. A. & Doherty, M. F. Multisite models to determine the distribution of kink sites adjacent to low-energy edges. Phys. Rev. E 85, 021604 (2012).
Joswiak, M. N., Doherty, M. F. & Peters, B. Ion dissolution mechanism and kinetics at kink sites on NaCl surfaces. Proc. Natl Acad. Sci. USA 115, 656–661 (2018).
Shtukenberg, A. G., Ward, M. D. & Kahr, B. Crystal growth with macromolecular additives. Chem. Rev. 117, 14042–14090 (2017).
Ma, W., Lutsko, J. F., Rimer, J. D. & Vekilov, P. G. Antagonistic cooperativity between crystal growth modifiers. Nature 577, 497–501 (2020).
Ehrlich, G. & Hudda, F. G. Asymetric capture at steps. J. Chem. Phys. 44, 1039–1052 (1966).
Olafson, K. N., Rimer, J. D. & Vekilov, P. G. Early onset of kinetic roughening due to a finite step width in hematin crystallization. Phys. Rev. Lett. 119, 198101 (2017).
Vekilov, P. G., Kuznetsov, Y. G. & Chernov, A. A. The effect of temperature on step motion; (101) ADP face. J. Cryst. Growth 121, 44–52 (1992).
Vekilov, P. G. What determines the rate of growth of crystals from solution? Cryst. Growth Des. 7, 2796–2810 (2007).
Dandekar, P., Kuvadia, Z. B. & Doherty, M. F. Engineering crystal morphology. Annu. Rev. Mater. Res. 43, 359–386 (2013).
Erdemir, D. et al. Relationship between self-association of glycine molecules in supersaturated solutions and solid state outcome. Phys. Rev. Lett. 99, 115702 (2007).
Parveen, S., Davey, R. J., Dent, G. & Pritchard, R. G. Linking solution chemistry to crystal nucleation: the case of tetrolic acid. Chem. Commun. 1531–1533 (2005).
Reutzel-Edens, S. M., Bush, J. K., Magee, P. A., Stephenson, G. A. & Byrn, S. R. Anhydrates and hydrates of olanzapine: crystallization, solid-state characterization and structural relationships. Cryst. Growth Des. 3, 897–907 (2003).
Thakuria, R. & Nangia, A. Olanzapinium salts, isostructural solvates and their physicochemical properties. Cryst. Growth Des. 13, 3672–3680 (2013).
Klamt, A. & Schüürmann, G. COSMO: a new approach to dielectric screening in solvents with explicit expressions for the screening energy and its gradient. J. Chem. Soc. Perkin Trans. 2, 799–805 (1993).
Qu, H. et al. Raman and ATR FTIR spectroscopy in reactive crystallization: simultaneous monitoring of solute concentration and polymorphic state of the crystals. J. Cryst. Growth 311, 3466–3475 (2009).
Hu, Y., Liang, J. K., Myerson, A. S. & Taylor, L. S. Crystallization monitoring by Raman spectroscopy: simultaneous measurement of desupersaturation profile and polymorphic form in flufenamic acid systems. Ind. Eng. Chem. Res. 44, 1233–1240 (2005).
Simone, E., Saleemi, A. N. & Nagy, Z. K. Application of quantitative Raman spectroscopy for the monitoring of polymorphic transformation in crystallization processes using a good calibration practice procedure. Chem. Eng. Res. Des. 92, 594–611 (2014).
Tanaka, N., Kitano, H. & Ise, N. Raman spectroscopic study of hydrogen bonding in aqueous carboxylic acid solutions. J. Phys. Chem. 94, 6290–6292 (1990).
Ayala, A. P. et al. Solid state characterization of olanzapine polymorphs using vibrational spectroscopy. Int. J. Pharm. 326, 69–79 (2006).
Hunter, C. A., McCabe, J. F. & Spitaleri, A. Solvent effects of the structures of prenucleation aggregates of carbamazepine. CrystEngComm 14, 7115–7117 (2012).
Spitaleri, A., Hunter, C. A., McCabe, J. F., Packer, M. J. & Cockroft, S. L. A 1H NMR study of crystal nucleation in solution. CrystEngComm 6, 490–493 (2004).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2, 19–25 (2015).
Horn, H. W. et al. Development of an improved four-site water model for biomolecular simulations: TIP4P-Ew. J. Chem. Phys. 120, 24503 (2004).
Dodda, L. S., Cabeza de Vaca, I., Tirado-Rives, J. & Jorgensen, W. L. LigParGen web server: an automatic OPLS-AA parameter generator for organic ligands. Nucleic Acids Res. 45, W331–W336 (2017).
Dodda, L. S., Vilseck, J. Z., Tirado-Rives, J. & Jorgensen, W. L. 1.14*CM1A-LBCC: localized bond-charge corrected CM1A charges for condensed-phase simulations. J. Phys. Chem. B 121, 3864–3870 (2017).
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577–8593 (1995).
Miyamoto, S. & Kollman, P. A. SETTLE: an analytical version of the SHAKE and RATTLE algorithm for rigid water models. J. Comput. Chem. 13, 952–962 (1992).
Hess, B., Berendsen, H. J. C., Fraaije, J. G. E. M. & Bekker, H. LINCS: a linear constraint solver for molecular simulations. J. Comput. Chem. 18, 1463–1472 (1997).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101 (2007).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: a new molecular dynamics method. Strain fluctuations and elastic constants. J. Chem. Phys. 52, 234505 (1981).
Guevara-Carrion, G., Vrabec, J. & Hasse, H. Prediction of self-diffusion coefficient and shear viscosity of water and its binary mixtures with methanol and ethanol by molecular simulation. J. Chem. Phys. 134, 74508 (2011).
Hess, B. & Van Der Vegt, N. F A. Hydration thermodynamic properties of amino acid analogues: a systematic comparison of biomolecular force fields and water models. J. Phys. Chem. B 110, 17616–17626 (2006).
Caleman, C. et al. Force field benchmark of organic liquids: density, enthalpy of vaporization, heat capacities, surface tension, isothermal com-pressibility, volumetric expansion coefficient and dielectric constant. J. Chem. Theory Comput. 8, 61–74 (2012).
Kästner, J. Umbrella sampling. WIREs Comput. Mol. Sci. 1, 932–942 (2011).
Torrie, G. M. & Valleau, J. P. Monte Carlo free energy estimates using non-Boltzmann sampling: application to the sub-critical Lennard–Jones fluid. Chem. Phys. Lett. 28, 578–581 (1974).
Torrie, G. M. & Valleau, J. P. Nonphysical sampling distributions in Monte Carlo free-energy estimation: umbrella sampling. J. Comput. Phys. 23, 187–199 (1977).
Bonomi, M. et al. PLUMED: a portable plugin for free-energy calculations with molecular dynamics. Comput. Phys. Commun. 180, 1961–1972 (2009).
Ferguson, A. L. BayesWHAM: a Bayesian approach for free energy estimation, reweighting and uncertainty quantification in the weighted histogram analysis method. J. Comput. Chem. 38, 1583–1605 (2017).
Kumar, S. et al. The weighted histogram analysis method for free-energy calculations on biomolecules. I. The method. J. Comput. Chem. 13, 1011–1021 (1992).
Fiorin, G., Klein, M. L. & Hénin, J. Using collective variables to drive molecular dynamics simulations. Mol. Phys. 111, 3345–3362 (2013).
De Jong, D. H. et al. Determining equilibrium constants for dimerization reactions from molecular dynamics simulations. J. Comput. Chem. 32, 1919–1928 (2011).
Roy, A., Hua, D. P., Ward, J. M. & Post, C. B. Relative binding enthalpies from molecular dynamics simulations using a direct method terms of use. J. Chem. Theory Comput. 10, 2759–2768 (2014).
Chernov, A. A. The spiral growth of crystals. Sov. Phys. Uspekhi 4, 116–148 (1961).
Chernov, A. A. & Komatsu, H. in Science and Technology of Crystal Growth (eds van der Eerden, J. P. & Bruinsma, O. S. L.) 67–80 (Kluwer Academic, 1995).
Olafson, K. N., Ketchum, M. A., Rimer, J. D. & Vekilov, P. G. Mechanisms of hematin crystallization and inhibition by the antimalarial drug chloroquine. Proc. Natl Acad. Sci. USA 112, 4946–4951 (2015).
Vekilov, P. G., Kuznetsov, Y. G. & Chernov, A. A. Interstep interaction in solution growth; (101) ADP face. J. Cryst. Growth 121, 643–655 (1992).
Maruyama, M., Tsukamoto, K., Sazaki, G., Nishimura, Y. & Vekilov, P. G. Chiral and achiral mechanisms of regulation of calcite crystallization. Cryst. Growth Des. 9, 127–135 (2009).
Olafson, K. N., Ketchum, M. A., Rimer, J. D. & Vekilov, P. G. Molecular mechanisms of hematin crystallization from organic solvent. Cryst. Growth Des. 15, 5535–5542 (2015).
Petsev, D. N., Chen, K., Gliko, O. & Vekilov, P. G. Diffusion-limited kinetics of the solution-solid phase transition of molecular substances. Proc. Natl Acad. Sci. USA 100, 792–796 (2003).
Vekilov, P. G. Incorporation at kinks: kink density and activation barriers. In AIP Conference Proceedings Vol. 916 (eds Skowronski, M., DeYoreo, J. J. & Wang, C. A.) 235–267 (2007).
Elhadj, S., De Yoreo, J. J., Hoyer, J. R. & Dove, P. M. Role of molecular charge and hydrophilicity in regulating the kinetics of crystal growth. Proc. Natl Acad. Sci. USA 103, 19237–19242 (2006).
Elhadj, S. et al. Peptide controls on calcite mineralization: polyaspartate chain length affects growth kinetics and acts as a stereochemical switch on morphology. Cryst. Growth Des. 6, 197–201 (2006).
Teng, H. H. How ions and molecules organize to form crystals. Elements 9, 189–194 (2013).
Acknowledgements
We thank M. Doherty, S. Price, E. Vlieg, D. Maes and J. Rimer for invaluable discussions and suggestions. This work was supported by the National Science Foundation (award no. DMR-1710354), NASA (award nos. NNX14AD68G and NNX14AE79G) and The Welch Foundation (Grant E-1882). We acknowledge the CMAC National Facility, housed within the University of Strathclyde’s Technology and Innovation Centre, and funded with a UK Research Partnership Investment Fund (UKRPIF) capital award, SFC ref H13054, from the Higher Education Funding Council for England (HEFCE). Computational resources were provided by the Hewlett Packard Enterprise Data Science Institute and the University of Houston and the Texas Advanced Computing Center at the University of Texas at Austin.
Author information
Authors and Affiliations
Contributions
P.G.V., M.W. and A.J.F. conceived this work. P.G.V. and M.W. designed the experiments and M.W. performed all experiments and analysed data. B.F.J. and M.W. modelled the Raman spectra and adsorption on the OZPN crystal surface. P.G.V. developed kinetic and thermodynamic models. L.V. and J.C.P. carried out molecular dynamics simulations. P.G.V., M.W., L.V., J.C.P. and A.J.F. wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
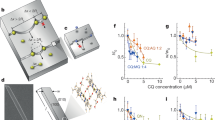
Extended Data Fig. 1 NMR study of self-association of OZPN molecules in solution.
a, concentration dependent 1H NMR spectra of OZPN in 1/1 (vol/vol) EtOD/D2O with assigned protons at listed concentrations. b, 1H NMR spectra of OZPN in CDCl3 with assigned protons at listed concentrations. Vertical dotted lines highlight shift evolution with increased concentration. c, Normalized changes in chemical shift observed for OZPN protons as a function of concentration in CDCl3 solution. Lines are guides for the eye. d, The C−H···π interactions stabilizing SC0 dimer illustrated using the Mercury software package.
Extended Data Fig. 2 Raman spectra of OZPN solutions.
a, b, Measured at listed concentrations in CHCl3 and EtOH/H2O. c, d, Simulated spectra of OZPN monomer in c and dimers, in d, in H2O, EtOH, and CHCl3.
Extended Data Fig. 3 Evaluation of the OZPN dimerization equilibrium constant.
a, In 1/1 (vol/vol) EtOH/H2O. b, In CHCl3. Root mean squared deviation (RMSD) of the intensity of the characteristic dimer Raman peak at 1517 cm−1 ID computed using Equation. 8 from the experimentally measured values at five OZPN concentrations in each solvent. The minima of these correlations indicate the values of KD that best fit the Raman spectra at different concentrations.
Extended Data Fig. 4 Predicted correlations between the step velocity and solute concentration.
Predicted correlations assuming four scenarios summarized in Extended Data Table 2. i. Monomers dominate in the solution and growth occurs by incorporation of monomers. ii. Monomers dominate in solution, but the growth occurs by incorporation of dimers. iii. Dimers dominate in solution but the growth occurs by monomers. iv. Dimers dominate in solution and growth occurs by the incorporation dimers.
Extended Data Fig. 5 The surface diffusion mechanism operates during growth of OZPN HE crystals.
a, The correlation between the step velocity v and the step separation ℓ at C = 2.38 mM. b, The correlation between v and ℓ in coordinates [σ/ν](1/ ℓ), where σ = (C − Ce)/Ce. In both plots, regions of depressed v enforced by step supply field overlap at ℓ < 250 nm are highlighted in gray. Error bars represent the standard deviation from the average of five independent measurements and are smaller than the symbol size for some data points.
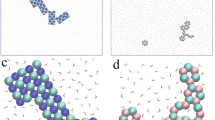
Extended Data Fig. 6 Adsorption of OZPN monomers and dimers on the (002) face of 2OZPN·EtOH·2H2O crystals in contact with a 1/1 (vol/vol) EtOH H2O solution.
a, Schematic of dimerization in the solution bulk and on the crystal surface. \({\mathrm{\Delta }}H_D^0\) are the respective equilibrium enthalpies of dimerization in the bulk and at the surface and \({\mathrm{\Delta }}H_{ads}^0\), of adsorption of monomers and dimers. b, c, Structuring of the water, in b, and ethanol, in c, molecules at the crystal interface. Only the lattice water molecules are represented with their van der Waals surfaces, where red and white encode for O and H, respectively. In the solution, C atoms are shown in cyan, O and H in red. Hydrogen bonds between water molecules are depicted with dashed green lines in b. d–g, OZPN monomers, in d and f, and dimers, in e and g, displace distinct numbers of solvent EtOH and H2O molecules in their various surface conformations. Only two confirmations for each species are shown, a dormant in d and e, and a rampant, in f and g.
Extended Data Fig. 7 X-ray identification of the crystal form of 2OZPN·EtOH·2H2O.
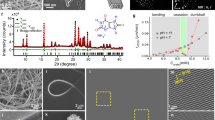
a, Power x-ray diffraction spectrum. b, Identification of the crystal faces by single-crystal X-ray diffraction.
Extended Data Fig. 8 Lack of interaction between the steps on the surface of OZPN HE during data collection for Fig. 1d and g.
a, The separation between steps ℓ at several OZPN concentrations during the measurements of v. Error bars correspond to standard deviations of about 30 measurements. b, The correlation between the step velocity v and ℓ for ℓ > 300 nm at C = 3.65 mM. Observed step separations during measurements of v were binned in 75 nm wide groups. Error bars correspond to standard deviations of 10 to 15 v data points in each group and are smaller than the symbol size for some data points.
Supplementary information
Supplementary Information
Supplementary text, references, tables and figures.
Source data
Source Data Fig. 1
CDX file for OZPN structure.
Source Data Fig. 1
Raw data for Fig. 1e–g.
Source Data Fig. 2
Raw data for Fig. 2a.
Source Data Fig. 4
Raw data for Fig. 4a–c.
Source Data Fig. 5
Raw data for Fig. 5d.
Source Data Extended Data Fig. 1
Source Data Extended Data Fig. 1
Source Data Extended Data Fig. 2
Source Data Extended Data Fig. 2
Source Data Extended Data Fig. 4
Source Data Extended Data Fig. 4
Source Data Extended Data Fig. 6
Source Data Extended Data Fig. 6
Source Data Extended Data Fig. 7
Source Data Extended Data Fig. 7
Rights and permissions
About this article
Cite this article
Warzecha, M., Verma, L., Johnston, B.F. et al. Olanzapine crystal symmetry originates in preformed centrosymmetric solute dimers. Nat. Chem. 12, 914–920 (2020). https://doi.org/10.1038/s41557-020-0542-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-020-0542-0
This article is cited by
-
Non-classical crystallization in soft and organic materials
Nature Reviews Materials (2024)
-
Symmetry in the making
Nature Chemistry (2020)