Abstract
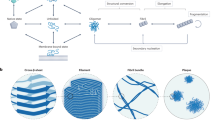
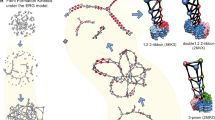
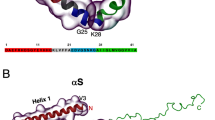
Oligomeric species populated during the aggregation of the Aβ42 peptide have been identified as potent cytotoxins linked to Alzheimer’s disease, but the fundamental molecular pathways that control their dynamics have yet to be elucidated. By developing a general approach that combines theory, experiment and simulation, we reveal, in molecular detail, the mechanisms of Aβ42 oligomer dynamics during amyloid fibril formation. Even though all mature amyloid fibrils must originate as oligomers, we found that most Aβ42 oligomers dissociate into their monomeric precursors without forming new fibrils. Only a minority of oligomers converts into fibrillar structures. Moreover, the heterogeneous ensemble of oligomeric species interconverts on timescales comparable to those of aggregation. Our results identify fundamentally new steps that could be targeted by therapeutic interventions designed to combat protein misfolding diseases.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The authors confirm that all data generated and analysed during this study are included in this published article and its Supplementary Information. Data are also available from the corresponding authors upon request.
Code availability
All the simulation and data analysis codes are included in this article and its Supplementary Information. Codes are available from the corresponding authors upon request.
Change history
17 April 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41557-020-0468-6
References
Alzheimer’s Association. 2014 Alzheimer’s disease facts and figures. Alzheimers Dement. 10, e47–e92 (2014).
Soto, C. Unfolding the role of protein misfolding in neurodegenerative diseases. Nat. Rev. Neurosci. 4, 49–60 (2003).
Dobson, C. M. Protein folding and misfolding. Nature 426, 884–890 (2003).
Chiti, F. & Dobson, C. M. Protein misfolding, functional amyloid, and human disease: a summary of progress over the last decade. Annu. Rev. Biochem. 86, 27 (2017).
Knowles, T. P. J., Vendruscolo, M. & Dobson, C. M. The amyloid state and its association with protein misfolding diseases. Nat. Rev. Mol. Cell Biol. 15, 384–396 (2014).
Selkoe, D. J. Folding proteins in fatal ways. Nature 426, 900–904 (2003).
Hardy, J. & Selkoe, D. J. The amyloid hypothesis of Alzheimer’s disease: progress and problems on the road to therapeutics. Science 297, 353–356 (2002).
Petkova, A. T. Self-propagating, molecular-level polymorphism in Alzheimer’s amyloid fibrils. Science 307, 262–265 (2005).
Campioni, S. et al. A causative link between the structure of aberrant protein oligomers and their toxicity. Nat. Chem. Biol. 6, 140–147 (2010).
Qiang, W., Yau, W.-M., Lu, J.-X., Collinge, J. & Tycko, R. Structural variation in amyloid-β fibrils from Alzheimer’s disease clinical subtypes. Nature 541, 217–221 (2017).
Cremades, N. et al. Direct observation of the interconversion of normal and toxic forms of α-synuclein. Cell 149, 1048–1059 (2012).
Benilova, I., Karran, E. & De Strooper, B. The toxic Aβ oligomer and Alzheimer’s disease: an emperor in need of clothes. Nat. Neurosci. 15, 349–357 (2012).
Catalano, S. M. et al. The role of amyloid-beta derived diffusible ligands (ADDLs) in Alzheimer’s disease. Curr. Top. Med. Chem. 6, 597–608 (2006).
Lashuel, H. A., Hartley, D., Petre, B. M., Walz, T. & Lansbury, P. T. Jr Neurodegenerative disease: amyloid pores from pathogenic mutations. Nature 418, 291 (2002).
Knowles, T. P. J. et al. An analytical solution to the kinetics of breakable filament assembly. Science 326, 1533–1537 (2009).
Cohen, S. I. A. et al. Proliferation of amyloid-β42 aggregates occurs through a secondary nucleation mechanism. Proc. Natl Acad. Sci. USA 110, 9758–9763 (2013).
Törnquist, M. et al. Secondary nucleation in amyloid formation. Chem. Commun. 54, 8667–8684 (2018).
Michaels, T. C. T. et al. Chemical kinetics for bridging molecular mechanisms and macroscopic measurements of amyloid fibril formation. Annu. Rev. Phys. Chem. 69, 273–298 (2018).
Walsh, D. M. et al. A facile method for expression and purification of the Alzheimer’s disease-associated amyloid β-peptide. FEBS J. 276, 1266–1281 (2009).
Szczepankiewicz, O. et al. N-terminal extensions retard Aβ42 fibril formation but allow cross-seeding and coaggregation with Aβ42. J. Am. Chem. Soc. 137, 14673–14685 (2015).
Nasir, I., Linse, S. & Cabaleiro-Lago, C. Fluorescent filter-trap assay for amyloid fibril formation kinetics in complex solutions. ACS Chem. Neurosci. 6, 1436–1444 (2015).
Colvin, M. T. et al. Atomic resolution structure of monomorphic Aβ-42 amyloid fibrils. J. Am. Chem. Soc. 138, 9663–9674 (2016).
Wälti, M. A. et al. Atomic-resolution structure of a disease-relevant Aβ(1–42) amyloid fibril. Proc. Natl Acad. Sci. USA 113, E4976–E4984 (2016).
Anwar, J., Khan, S. & Lindfors, L. Secondary crystal nucleation: nuclei breeding factory uncovered. Angew. Chem. Int. Ed. 54, 14681–14684 (2015).
Orgel, L. E. Prion replication and secondary nucleation. Chem. Biol. 3, 413–414 (1996).
Vekilov, P. G. The two-step mechanism of nucleation of crystals in solution. Nanoscale 2, 2346–2357 (2010).
Sear, R. P. The non-classical nucleation of crystals: microscopic mechanisms and applications to molecular crystals, ice and calcium carbonate. Int. Mater. Rev. 57, 328–356 (2012).
Koffie, R. et al. Oligomeric amyloid associates with postsynaptic densities and correlates with excitatory synapse loss near senile plaques. Proc. Natl Acad. Sci. USA 106, 4012–4017 (2009).
Cohen, S. I. A. et al. A molecular chaperone breaks the catalytic cycle that generates toxic Aβ oligomers. Nat. Struct. Mol. Biol. 22, 207–213 (2015).
Cukalevki, R. et al. The Aβ40 and Aβ42 peptides self-assemble into separate homomolecular fibrils in binary mixtures but cross-react during primary nucleation. Chem. Sci. 6, 4215–4233 (2015).
Šarić, A. et al. Physical determinants of the self-replication of protein fibrils. Nat. Phys. 12, 874–880 (2016).
Fändrich, M., Fletcher, M. A. & Dobson, C. M. Amyloid fibrils from muscle myoglobin. Nature 410, 165–166 (2001).
Allison, J. R., Varnai, P., C. M. Dobson, C. M. & Vendruscolo, M. Determination of the free energy landscape of alpha-synuclein using spin label nuclear magnetic resonance measurements. J. Am. Chem. Soc. 131, 18314–18326 (2009).
Šarić, A., Chebaro, Y. C., Knowles, T. P. J. & Frenkel, D. Crucial role of nonspecific interactions in amyloid nucleation. Proc. Natl Acad. Sci. USA 111, 17869–17874 (2014).
Šarić, A., Michaels, T. C. T., Zaccone, A., Knowles, T. P. J. & Frenkel, D. Kinetics of spontaneous filament nucleation via oligomers: insights from theory and simulation. J. Chem. Phys. 145, 211926 (2016).
Michaels, T. C. T. et al. Reaction rate theory for supramolecular kinetics: application to protein aggregation. Mol. Phys. 116, 3055–3065 (2018).
Galkin, O. et al. Two-step mechanism of homogeneous nucleation of sickle cell hemoglobin polymers. Biophys. J. 93, 902–913 (2007).
Auer, S., Dobson, C. M. & Vendruscolo, M. Characterization of the nucleation barriers for protein aggregation and amyloid formation. HFSP J. 1, 137–146 (2007).
Lee, C.-T. & Terentjev, E. M. Mechanisms and rates of nucleation of amyloid fibrils. J. Chem. Phys. 147, 105103 (2017).
Chen, S. W. et al. Structural characterization of toxic oligomers that are kinetically trapped during alpha-synuclein fibril formation. Proc. Natl Acad. Sci. USA 112, E1994–E2003 (2015).
Garai, K., Posey, A. E., Li, X., Buxbaum, J. N. & Pappu, R. V. Inhibition of amyloid beta fibril formation by monomeric human transthyretin. Protein Sci. 27, 1252–1261 (2018).
Acknowledgements
We acknowledge support from Peterhouse (T.C.T.M.), the Swiss National Science foundation (T.C.T.M.), the Royal Society (A.Š.), the Academy of Medical Sciences (A.Š.), the UCL Institute for the Physics of Living Systems (S.C.), Sidney Sussex College (G.M.), the Wellcome Trust (A.Š., M.V., C.M.D. and T.P.J.K.), the Schiff Foundation (A.J.D.), the Cambridge Centre for Misfolding Diseases (M.V., C.M.D. and T.P.J.K.), the BBSRC (C.M.D. and T.P.J.K.), the Frances and Augustus Newman Foundation (T.P.J.K.), the Swedish Research Council (S.L.) and the ERC grant MAMBA (S.L., agreement no. 340890). The research that led to these results received funding from the European Research Council under the European Union’s Seventh Framework Programme (FP7/2007-2013) through the ERC grant PhysProt (agreement no. 337969).
Author information
Authors and Affiliations
Contributions
All the authors were involved in the design of the study; T.C.T.M. developed the theoretical model and performed the kinetic analysis; S.L. and K.B. performed the experiments; A.Š. and S.C. performed computer simulations; all the authors participated in interpreting the results and writing the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Experimental methods, details on computer simulations, definition and solution of mathematical models of oligomer dynamics, details on data analysis, Supplementary Figs. 1–19, Tables 1 and 2, and Video 1.
Rights and permissions
About this article
Cite this article
Michaels, T.C.T., Šarić, A., Curk, S. et al. Dynamics of oligomer populations formed during the aggregation of Alzheimer’s Aβ42 peptide. Nat. Chem. 12, 445–451 (2020). https://doi.org/10.1038/s41557-020-0452-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-020-0452-1
This article is cited by
-
Misfolded protein oligomers: mechanisms of formation, cytotoxic effects, and pharmacological approaches against protein misfolding diseases
Molecular Neurodegeneration (2024)
-
High-throughput and proteome-wide discovery of endogenous biomolecular condensates
Nature Chemistry (2024)
-
Identification of potential aggregation hotspots on Aβ42 fibrils blocked by the anti-amyloid chaperone-like BRICHOS domain
Nature Communications (2024)
-
Data-mining unveils structure–property–activity correlation of viral infectivity enhancing self-assembling peptides
Nature Communications (2023)
-
Engineering synthetic biomolecular condensates
Nature Reviews Bioengineering (2023)