Abstract
Small cell lung cancer (SCLC) exists broadly in four molecular subtypes: ASCL1, NEUROD1, POU2F3 and Inflammatory. Initially, SCLC subtypes were thought to be mutually exclusive, but recent evidence shows intra-tumoural subtype heterogeneity and plasticity between subtypes. Here, using a CRISPR-based autochthonous SCLC genetically engineered mouse model to study the consequences of KDM6A/UTX inactivation, we show that KDM6A inactivation induced plasticity from ASCL1 to NEUROD1 resulting in SCLC tumours that express both ASCL1 and NEUROD1. Mechanistically, KDM6A normally maintains an active chromatin state that favours the ASCL1 subtype with its loss decreasing H3K4me1 and increasing H3K27me3 at enhancers of neuroendocrine genes leading to a cell state that is primed for ASCL1-to-NEUROD1 subtype switching. This work identifies KDM6A as an epigenetic regulator that controls ASCL1 to NEUROD1 subtype plasticity and provides an autochthonous SCLC genetically engineered mouse model to model ASCL1 and NEUROD1 subtype heterogeneity and plasticity, which is found in 35–40% of human SCLCs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Data from bulk RNA-seq experiments in Fig. 1 are included as Supplementary Table 2. Data from bulk ATAC-seq experiments from Fig. 2 or Fig. 3 are included as Supplementary Tables 3 and 4. Data from single-cell RNA-seq experiments in Fig. 3 are included as Supplementary Table 5. Data from Extended Data Fig. 5–7 are included as Supplementary Table 6. Data from ChIP–seq experiments in Fig. 5 are included as Supplementary Table 7. Data from Menin ChIP–seq and LISA analysis in Fig. 6 are included as Supplementary Table 8. Data from Extended Data Fig. 10 are included as Supplementary Tables 9 and 10. All source data are included within the supplementary information associated with this article. RNA-seq, ATAC-seq and ChIP–seq data that support the findings of this study have been deposited in the Gene Expression Omnibus under accession codes GSE228347 (RNA-seq Fig. 1, Menin-ChiP–seq Fig. 6), GSE215829 (ATAC-seq Figs. 2 and 3), GSE211242 (scRNA-seq Fig. 3) and GSE211165 (ChIP–seq Fig. 5). Any other data and materials can be requested from the corresponding author upon reasonable request. Source data are provided with this paper.
Code availability
Custom code for ATAC-seq analysis in Figs. 2 and 3 is here: https://bitbucket.org/cfce/viper/src/master/, https://bitbucket.org/cfce/cobra/src/master/ (ATAC-seq experiments). Custom R code for ChIP–seq analysis in Fig. 5 is here: https://doi.org/10.5281/zenodo.7786472.
References
Kalemkerian, G. P. et al. Small cell lung cancer. J. Natl Compr. Cancer Netw. 11, 78–98 (2013).
Rudin, C. M., Brambilla, E., Faivre-Finn, C. & Sage, J. Small-cell lung cancer. Nat. Rev. Dis. Prim. 7, 3 (2021).
Rudin, C. M. et al. Molecular subtypes of small cell lung cancer: a synthesis of human and mouse model data. Nat. Rev. Cancer 19, 289–297 (2019).
George, J. et al. Comprehensive genomic profiles of small cell lung cancer. Nature 524, 47–53 (2015).
Peifer, M. et al. Integrative genome analyses identify key somatic driver mutations of small-cell lung cancer. Nat. Genet. 44, 1104–1110 (2012).
Rudin, C. M. et al. Comprehensive genomic analysis identifies SOX2 as a frequently amplified gene in small-cell lung cancer. Nat. Genet. 44, 1111–1116 (2012).
Gay, C. M. et al. Patterns of transcription factor programs and immune pathway activation define four major subtypes of SCLC with distinct therapeutic vulnerabilities. Cancer Cell 39, 346–360 e347 (2021).
Baine, M. K. et al. POU2F3 in SCLC: clinicopathologic and genomic analysis with a focus on its diagnostic utility in neuroendocrine-low SCLC. J. Thorac. Oncol. 17, 1109–1121 (2022).
Baine, M. K. et al. SCLC subtypes defined by ASCL1, NEUROD1, POU2F3, and YAP1: a comprehensive immunohistochemical and histopathologic characterization. J. Thorac. Oncol. 15, 1823–1835 (2020).
Qu, S. et al. Molecular subtypes of primary SCLC tumors and their associations with neuroendocrine and therapeutic markers. J. Thorac. Oncol. 17, 141–153 (2022).
Huang, Y. H. et al. POU2F3 is a master regulator of a tuft cell-like variant of small cell lung cancer. Genes Dev. 32, 915–928 (2018).
Cejas, P. et al. Subtype heterogeneity and epigenetic convergence in neuroendocrine prostate cancer. Nat. Commun. 12, 5775 (2021).
Ireland, A. S. et al. MYC drives temporal evolution of small cell lung cancer subtypes by reprogramming neuroendocrine fate. Cancer Cell 38, 60–78 e12 (2020).
Augert, A. et al. Small cell lung cancer exhibits frequent inactivating mutations in the histone methyltransferase KMT2D/MLL2: CALGB 151111 (Alliance). J. Thorac. Oncol. 12, 704–713 (2017).
Agger, K. et al. UTX and JMJD3 are histone H3K27 demethylases involved in HOX gene regulation and development. Nature 449, 731–734 (2007).
Lee, M. G. et al. Demethylation of H3K27 regulates polycomb recruitment and H2A ubiquitination. Science 318, 447–450 (2007).
Tran, N., Broun, A. & Ge, K. Lysine demethylase KDM6A in differentiation, development, and cancer. Mol. Cell Biol. https://doi.org/10.1128/MCB.00341-20 (2020).
Kim, J. H. et al. UTX and MLL4 coordinately regulate transcriptional programs for cell proliferation and invasiveness in breast cancer cells. Cancer Res. 74, 1705–1717 (2014).
Wang, L. & Shilatifard, A. UTX mutations in human cancer. Cancer Cell 35, 168–176 (2019).
Wang, S. P. et al. A UTX-MLL4-p300 transcriptional regulatory network coordinately shapes active enhancer landscapes for eliciting transcription. Mol. Cell 67, 308–321 e306 (2017).
Andricovich, J. et al. Loss of KDM6A activates super-enhancers to induce gender-specific squamous-like pancreatic cancer and confers sensitivity to BET inhibitors. Cancer Cell 33, 512–526 e518 (2018).
Faralli, H. et al. UTX demethylase activity is required for satellite cell-mediated muscle regeneration. J. Clin. Invest. 126, 1555–1565 (2016).
Gozdecka, M. et al. UTX-mediated enhancer and chromatin remodeling suppresses myeloid leukemogenesis through noncatalytic inverse regulation of ETS and GATA programs. Nat. Genet. 50, 883–894 (2018).
Morales Torres, C., Laugesen, A. & Helin, K. Utx is required for proper induction of ectoderm and mesoderm during differentiation of embryonic stem cells. PLoS ONE 8, e60020 (2013).
Tsherniak, A. et al. Defining a cancer dependency map. Cell 170, 564–576 e516 (2017).
Oser, M. G. et al. The KDM5A/RBP2 histone demethylase represses NOTCH signaling to sustain neuroendocrine differentiation and promote small cell lung cancer tumorigenesis. Genes Dev. 33, 1718–1738 (2019).
Chakraborty, A. A. et al. Histone demethylase KDM6A directly senses oxygen to control chromatin and cell fate. Science 363, 1217–1222 (2019).
Kalisz, M. et al. HNF1A recruits KDM6A to activate differentiated acinar cell programs that suppress pancreatic cancer. EMBO J. 39, e102808 (2020).
Borromeo, M. D. et al. ASCL1 and NEUROD1 reveal heterogeneity in pulmonary neuroendocrine tumors and regulate distinct genetic programs. Cell Rep. 16, 1259–1272 (2016).
Hong, D. et al. Plasticity in the absence of NOTCH uncovers a RUNX2-dependent pathway in small cell lung cancer. Cancer Res. https://doi.org/10.1158/0008-5472.can-21-1991 (2021).
Mollaoglu, G. et al. MYC drives progression of small cell lung cancer to a variant neuroendocrine subtype with vulnerability to aurora kinase inhibition. Cancer Cell 31, 270–285 (2017).
Schaffer, B. E. et al. Loss of p130 accelerates tumor development in a mouse model for human small-cell lung carcinoma. Cancer Res. 70, 3877–3883 (2010).
Marsich, E., Vetere, A., Di Piazza, M., Tell, G. & Paoletti, S. The PAX6 gene is activated by the basic helix-loop-helix transcription factor NeuroD/BETA2. Biochem. J. 376, 707–715 (2003).
Chan, J. M. et al. Signatures of plasticity, metastasis, and immunosuppression in an atlas of human small cell lung cancer. Cancer Cell 39, 1479–1496 e1418 (2021).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Mahadevan, N. R. et al. Intrinsic immunogenicity of small cell lung carcinoma revealed by its cellular plasticity. Cancer Discov. https://doi.org/10.1158/2159-8290.CD-20-0913 (2021).
Young, M. D. et al. ChIP–seq analysis reveals distinct H3K27me3 profiles that correlate with transcriptional activity. Nucleic Acids Res. 39, 7415–7427 (2011).
Qin, Q. et al. LISA: inferring transcriptional regulators through integrative modeling of public chromatin accessibility and ChIP–seq data. Genome Biol. 21, 32 (2020).
Krivtsov, A. V. & Armstrong, S. A. MLL translocations, histone modifications and leukaemia stem-cell development. Nat. Rev. Cancer 7, 823–833 (2007).
Krivtsov, A. V. et al. A Menin-MLL inhibitor induces specific chromatin changes and eradicates disease in models of MLL-rearranged leukemia. Cancer Cell 36, 660–673 e611 (2019).
Lissa, D. et al. Heterogeneity of neuroendocrine transcriptional states in metastatic small cell lung cancers and patient-derived models. Nat. Commun. 13, 2023 (2022).
Easwaran, H., Tsai, H. C. & Baylin, S. B. Cancer epigenetics: tumor heterogeneity, plasticity of stem-like states, and drug resistance. Mol. Cell 54, 716–727 (2014).
Flavahan, W. A., Gaskell, E. & Bernstein, B. E. Epigenetic plasticity and the hallmarks of cancer. Science https://doi.org/10.1126/science.aal2380 (2017).
Stewart, C. A. et al. Single-cell analyses reveal increased intratumoral heterogeneity after the onset of therapy resistance in small-cell lung cancer. Nat. Cancer 1, 423–436 (2020).
Simpson, K. L. et al. A biobank of small cell lung cancer CDX models elucidates inter- and intratumoral phenotypic heterogeneity. Nat. Cancer 1, 437–451 (2020).
Adam, M. P. et al. Kabuki syndrome: international consensus diagnostic criteria. J. Med. Genet. 56, 89–95 (2019).
Cheon, C. K. et al. Identification of KMT2D and KDM6A mutations by exome sequencing in Korean patients with Kabuki syndrome. J. Hum. Genet. 59, 321–325 (2014).
DuPage, M., Dooley, A. L. & Jacks, T. Conditional mouse lung cancer models using adenoviral or lentiviral delivery of Cre recombinase. Nat. Protoc. 4, 1064–1072 (2009).
GPP sgRNA Designer (Broad Institute, 2023); https://portals.broadinstitute.org/gpp/public/analysis-tools/sgrnadesign
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Yang, L. et al. Validation of a hypoxia related gene signature in multiple soft tissue sarcoma cohorts. Oncotarget 9, 3946–3955 (2018).
Benita, Y. et al. An integrative genomics approach identifies Hypoxia Inducible Factor-1 (HIF-1)-target genes that form the core response to hypoxia. Nucleic Acids Res. 37, 4587–4602 (2009).
Bankhead, P. et al. QuPath: open source software for digital pathology image analysis. Sci. Rep. 7, 16878 (2017).
Buenrostro, J. D. et al. Single-cell chromatin accessibility reveals principles of regulatory variation. Nature 523, 486–490 (2015).
Taing, L. et al. CHIPS: a Snakemake pipeline for quality control and reproducible processing of chromatin profiling data [version 1; peer review: 1 not approved]. F1000Research https://doi.org/10.12688/f1000research.52878.1 (2021).
Qiu, X. et al. CoBRA: containerized bioinformatics workflow for reproducible ChIP/ATAC-seq analysis. Genomics Proteom. Bioinform. 19, 652–661 (2021).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-–seq (MACS). Genome Biol. 9, R137–R137 (2008).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Neph, S. et al. BEDOPS: high-performance genomic feature operations. Bioinformatics 28, 1919–1920 (2012).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Ramirez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
H. Wickham. ggplot2: Elegant Graphics for Data Analysis (Springer, 2016).
Cornwell, M. et al. VIPER: visualization pipeline for RNA-seq, a Snakemake workflow for efficient and complete RNA-seq analysis. BMC Bioinform. 19, 135–135 (2018).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Trapnell, C. et al. Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Wang, L., Wang, S. & Li, W. RSeQC: quality control of RNA-seq experiments. Bioinformatics 28, 2184–2185 (2012).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550–550 (2014).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Zerbino, D. R., Johnson, N., Juettemann, T., Wilder, S. P. & Flicek, P. WiggleTools: parallel processing of large collections of genome-wide datasets for visualization and statistical analysis. Bioinformatics 30, 1008–1009 (2014).
Satpathy, A. T. et al. Massively parallel single-cell chromatin landscapes of human immune cell development and intratumoral T cell exhaustion. Nat. Biotechnol. 37, 925–936 (2019).
Satpathy, A. T. et al. Massively parallel single-cell chromatin landscapes of human immune cell development and intratumoral T cell exhaustion. Nat. Biotechnol. 37, 925–936 (2019).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 e1821 (2019).
Stuart, T., Srivastava, A., Madad, S., Lareau, C. A. & Satija, R. Single-cell chromatin state analysis with Signac. Nat. Methods 18, 1333–1341 (2021).
Platt, R. J. et al. CRISPR–Cas9 knockin mice for genome editing and cancer modeling. Cell 159, 440–455 (2014).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–587 e3529 (2021).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2019).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172 (2019).
Heng, T. S. & Painter, M. W., Immunological Genome Project, C. The Immunological Genome Project: networks of gene expression in immune cells. Nat. Immunol. 9, 1091–1094 (2008).
Benayoun, B. A. et al. Remodeling of epigenome and transcriptome landscapes with aging in mice reveals widespread induction of inflammatory responses. Genome Res 29, 697–709 (2019).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Kharchenko, P. V., Tolstorukov, M. Y. & Park, P. J. Design and analysis of ChIP–seq experiments for DNA-binding proteins. Nat. Biotechnol. 26, 1351–1359 (2008).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Thorvaldsdóttir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
Casper, J. et al. The UCSC Genome Browser database: 2018 update. Nucleic Acids Res. 46, D762–D769 (2018).
Moore, J. E. et al. Expanded encyclopaedias of DNA elements in the human and mouse genomes. Nature 583, 699–710 (2020).
Caeser, R. et al. Genomic and transcriptomic analysis of a library of small cell lung cancer patient-derived xenografts. Nat. Commun. 13, 2144 (2022).
Acknowledgements
We thank A. Chakraborty for insightful discussions and members of the Oser, Barbie and Janne laboratories for thoughtful comments during our lab meetings. M.G.O. is a William Raveis Charitable Fund Damon Runyon Clinical Investigator. This work was supported by the Damon Runyon Cancer Research Foundation (CI-101-19, M.G.O.), an NCI/NIH K08 grant (no. K08CA222657, M.G.O.), NCI R37 grant (no. R37CA269990, M.G.O.) and the Kaplan Family Fund (M.G.O.). S.A.A. was supported by NIH grants CA176745, CA259273 and CA066996. S.L. was supported by the NCI Research Specialist Award (R50CA251956).
Author information
Authors and Affiliations
Contributions
Conceptualization: L.D., Y.L. and M.G.O.; methodology: L.D., Y.L., M.A.B., Y.X., S.N.O., P.C., M.Y.T., H.W.L. and M.G.O.; investigation: L.D., Y.L., M.A.B., Y.X., S.N.O., R.A.P., D.H., T.D., E.W., Y.N.L., J.S., S.L., P.C., H.W.L., M.Y.T., M.C.H. and M.G.O.; formal analysis: L.D., Y.L., M.B., Y.X., C.H., R.L., Y.J., S.S., X.Q., R.T.B. and M.G.O; writing: L.D., Y.L., M.A.B.,Y.X., S.N.O., S.A.A., H.W.L., M.Y.T., M.C.H. and M.G.O.; review and editing: all authors; funding acquisition: M.G.O.
Corresponding author
Ethics declarations
Competing interests
M.G.O. has (currently or previously) sponsored research agreements with Takeda, Eli Lilly, Novartis, Circle Pharma and Bristol Myers Squibb, and is a consultant for Daiichi Sankyo and Ananke Therapeutics. None of these sponsored research agreements was used to fund this work. S.A.A. has been a consultant and/or shareholder for Neomorph Inc, Imago Biosciences, Cyteir Therapeutics, C4 Therapeutics, Nimbus Therapeutics and Accent Therapeutics. S.A.A. has received research support from Janssen and Syndax. S.A.A. is named as an inventor on a patent application related to MENIN inhibition WO/2017/132398A1. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks Charles Rudin, Paul Wade and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 KDM6A Inactivation in an Autochthonous SCLC Mouse Model Promotes NEUROD1 Expression Leading to SCLC Tumors that Express both ASCL1 and NEUROD1.
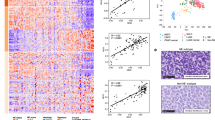
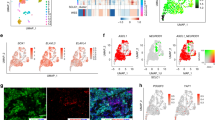
(a) Immunoblot analysis of 293T cells infected with adenoviruses that encoded Cre recombinase and the indicated sgRNAs. (b) Kaplan-Meier survival analysis of LSL-Cas9 mice IT injected with the indicated adenoviruses. p = 0.3447 for sgKdm6a RPP vs. sgControl RPP (non-targeting), p = 0.2781 for sgKdm6a RPP vs. sgControl RPP (intron-targeting), p = 0.3481 for sgControl RPP (non-targeting) vs. sgControl RPP (intron-targeting). n = 11 mice sgControl RPP (non-targeting), n = 5 mice sgControl RPP (intron-targeting), and n = 8 mice (sgKdm6a#4 RPP). (c) Principal component analysis (PCA) of gene expression from RNA-seq data of 6 Kdm6a-WT SCLC mice tumors (18,221, 18,222, 18,227, 535, 542, 645) and 7 Kdm6a-Mutant SCLC mice tumors (236L, 236R, 656, 651L, 651R, 672-1, 672-2) in Fig. 1d. (d, e) mRNA expression of ASCL1 vs. PAX6 (D) or NEUROD1 vs. PAX6 (e) from publicly available RNA-seq data set of 81 human SCLC samples from George et al. Nature 20154. p-values are generated from cbioportal.org. (f) Gene set enrichment analysis (GSEA) of RNA-seq data (from Fig. 1d, Extended Data Fig. 1c) of NEUROD1 correlated genes (401 genes; see Borromeo et al. Cell Reports 201629, see Supplementary Table 2)29. FDR q-value calculated using GSEA is indicated. (g–k) mRNA expression from the RNA-seq data (from Fig. 1d) for Ascl1 (g), Neurod1 (h), Chromogranin A (i), Synaptophysin (j) and Insm1 (k) of individual Kdm6a-WT and Kdm6a-Mutant mouse SCLC tumors. For i, j, and k, lower graphs show average mRNA expression in Kdm6a-WT vs. Kdm6a-Mutant lung tumors (see Supplementary Table 2). For i,j,k, data are presented as mean values ± SEM. Statistical significance was calculated using unpaired, two-tailed students t-test and p-values are indicated on each figure panel. n = 6 Kdm6a-WT tumors from independent mice and n = 7 Kdm6a-Mutant tumors.
Extended Data Fig. 2 IHC and ATAC-seq Data from Kdm6a-Mutant vs. Kdm6a-WT Autochthonous SCLC tumors from Figs. 1 and 2.
(a) RT-qPCR for Neurod1 at the times indicated after transduction of mouse SCLC cell lines derived from Kdm6a-WT mice (1014) with 2 independent Kdm6a sgRNAs or a non-targeting sgRNA (sgControl). Data are relative to Actb and then normalized to Neurod1 expression in sgControl sample. n = 2 biological independent experiments. Data are presented as mean values ± SEM. Statistical significance was calculated using unpaired, two-tailed students t-test and p-values are indicated. (b) Immunoblot analysis of the cells from a 24 days after transduction. NCI-H82 cells, a human NEUROD1-high SCLC cell line, is used as a positive control for NEUROD1 expression. (c) Immunohistochemistry (IHC) for ASCL1 and NEUROD1 from 3 Kdm6a-WT and 3 Kdm6a-Mutant mouse SCLC lung tumors. Scale Bar=50 µm. (d) Quantification of ASCL1-positive, NEUROD1-positive and ASCL1/NEUROD1-negative (double-negative) cells in Kdm6a-WT tumors (n = 6 tumors) and Kdm6a-Mutant tumors (n = 7 tumors) from the multiplex-IF data in Fig. 1j. (e) Representative H&E and IHC staining for NEUROD1 and ASCL1 of a Kdm6a-Mutant tumor (236L). (f, g) Representative H&E of a Kdm6a-Mutant tumor (656) (f) and a Kdm6a-WT tumor (1015) (g). For e and f, black arrows show enlarged nuclei of NEUROD1-positive cells with occasional multi-nucleated giant cells. Scale Bar=20 µm. (h) Bar graph of upregulated (red) and downregulated (blue) differential accessible peaks with LFC > 2, pAdj<0.05 from ATAC-seq data in Fig. 2a in Kdm6a-Mutant vs Kdm6a-WT tumors. p-values are calculated using Wald test in DEseq2 and adjusted for multiple hypothesis testing. (i) Pie charts of genomic location of differential accessible peaks from a. (j) Heat maps of ATAC-seq read densities from pseudo-bulk analysis of scATAC-seq data from Fig. 2f showing gain (n = 11,760 peaks) or loss (n = 10,587) of accessibility of Kdm6aMutantNEUROD1Only vs Kdm6aWT SCLC tumors (see Extended Data Fig. 3d).
Extended Data Fig. 3 Analysis of Bulk ATAC-seq from Kdm6a-Mutant vs. Kdm6a-WT Tumors from Fig. 3.
(a) RT-qPCR for Ascl1 mRNA expression relative to Actb and then normalized to Ascl1 expression in the 236L cell line of mouse SCLC lung tumors from Fig. 3a. (b) IHC for ASCL1 and NEUROD1 from the 1119 Kdm6a-Mutant mouse SCLC tumor showing rare NEUROD1-positive cells. Scale Bar= 50 µm. (c) Immunoblot analysis of SCLC lung tumors formed in an additional cohort of LSL-Cas9 mice injected with sgControl RPP (Kdm6a-WT) or sgKdm6a RPP (Kdm6a-Mutant) adenoviruses. For the immunoblot on the left, a separate piece of each tumor was used for bulk ATAC-seq and scRNA-seq experiments and hence immunoblots were run for each sample. Note that the piece of tumor 1270 used for ATAC-seq did show faint NEUROD1 expression while the 1270 sample used for scRNA-seq did not show any detectable NEUROD1 expression. This was likely related to heterogeneity within the tumor sample. (d) Bar graph of upregulated (red) and downregulated (blue) differential accessible peaks with LFC > 2, pAdj<0.05 from ATAC-seq data used to identify accessibility changes in Kdm6aMutantNEUROD1Only tumors for analyses in Fig. 3e, f and hence differential accessibility analysis was performed comparing Kdm6aMutantNEUROD1Only tumors (236, 656) vs. Kdm6aWT SCLC tumors (158, 18227, 165, 1198, 168, 645, 535, sc535). p-values are calculated using Wald test in DEseq2 and adjusted for multiple hypothesis testing. (e) Pie charts of genomic location of differential accessible peaks from d. (f, g) Chromatin accessibility tracks for the average of each phenotype indicated (see Fig. 3c–g) at Neurod1 (f) or Ascl1 (g) from all ATAC-seq data from Fig. 3d. (h, i) PCA of all ATAC-seq data from Fig. 3c (h) and Fig. 3d (i) now classified by genotype (Kdm6a-Mutant vs. Kdm6a-WT) rather than phenotype. (j) Heat maps of ATAC-seq read densities from all bulk ATAC-seq data from Fig. 3d–f showing gain (n = 11,760 peaks) or loss (n = 10,587) of accessibility in Kdm6aMutantNEUROD1Only vs Kdm6aWT SCLC tumors (see Extended Data Fig. 3d) for all phenotypes indicated.
Extended Data Fig. 4 Additional scRNA-seq Data from Kdm6a-Mutant vs. Kdm6a-WT Tumors from Fig. 3 and Additional RT-qPCR Data from Tumor Derived Cell Lines from Fig. 4.
(a) Dot plot showing the average expression of marker genes to identify immune cell subpopulations of all cells in Fig. 3h. The size of the dot represents the percentage of cells expressing a particular gene while color represents the mean gene expression levels (blue is low and red is high). (b) Elbow plot showing the standard deviation associated with the top 40 PCs in the tumor population. (c) Feature plot of Ascl1 of all tumor cells from scRNA-seq data in Kdm6a-Mutant vs. Kdm6a-WT (see Fig. 3i). (d) Neurod1 violin plot using scRNA-seq data from Fig. 3j of tumor cells from Kdm6a-Mutant and Kdm6a-WT tumors. For d, non-parametric Wilcoxon rank sum test was used to generate a two-tailed p-value adjusted for multiple comparisons by Bonferroni correction. n = 35,446 Kdm6a-Mutant and n = 6,612 Kdm6a-WT tumor cells. Each individual dot represents a cell. (e–j) RT-qPCR for Chromogranin A (e), Synaptophysin (f), Ascl1 (g), Neurod1 (h) Pou2f3 (i) and Yap1 (j) mRNA expression in the early vs. late passage tumor-derived cell lines of individual Kdm6a-WT and Kdm6a-Mutant (upper graphs) or average mRNA expression in Kdm6a-WT vs. Kdm6a-Mutant cell lines (lower graphs). n = 6 biological independent experiments for each genotype (2 biological independent experiments for each cell line). Data are presented as mean values ± SEM. Statistical significance was calculated using unpaired, two-tailed students t-test and p-values are indicated on each figure panel. For all t-tests comparing Kdm6a-Mutant vs. Kdm6a-WT cell lines, all early passage Kdm6a-Mutant were compared to all early passage Kdm6a-WT cell lines.
Extended Data Fig. 5 Comparison of Kdm6a-Mutant SCLC GEMM with the Myc-driven RPM SCLC GEMM using RNA-seq and IHC from tumors.
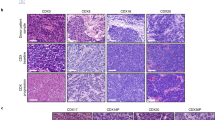
(a–c) mRNA expression from the RNA-seq data (from Fig. 1d) for Myc (a), Mycl (b) and Mycn (c) of individual Kdm6a-WT and Kdm6a-Mutant mouse SCLC tumors (top) or combined tumors by genotype (bottom). See Supplementary Table 2. For a,b,c, data are presented as mean values ± SEM. Statistical significance was calculated using unpaired, two-tailed students t-test and p-values are indicated on each figure panel. n = 6 Kdm6a-WT tumors from independent mice and n = 7 Kdm6a-Mutant tumors. (d) H&E and IHC staining for ASCL1, NEUROD1, c-Myc and Synaptophysin from 3 Kdm6a-WT (1014, 1015, 222) and 3 Kdm6a-Mutant (656, 236R, 670) mouse SCLC lung tumors. Scale Bar=50 µm. (e) IHC staining for c-Myc from tumor adjacent lymphocytes and high c-Myc prostate cancer GEMM as positive control of c-Myc staining. Scale Bar=50 µm. (f–h) GSEA of genes upregulated in the RPM vs. RPP GEMM13 with publicly available scRNA-seq data from human SCLCs of the ASCL1 and NEUROD1 subtypes34 (see Methods) (f), or with RNA-seq data from Kdm6a-Mutant vs. Kdm6a-WT tumors from Fig. 1d (g), or using GSEA of genes upregulated in Kdm6a-Mutant tumors (see Fig. 1e) with late RPM vs. early RPM subtypes13. FDR q-values from GSEA adjusted for multiple hypothesis testing are indicated.
Extended Data Fig. 6 Myc and Mycl mRNA Expression and Chromatin Accessibility in Kdm6a-Mutant vs. Kdm6a-WT Tumors.
(a–d) Feature plots with violin plot of single cells from tumors for Myc (a,b) and Mycl (c,d) expression from scRNA-seq data from Fig. 3i. For b,d, non-parametric Wilcoxon rank sum test was used to generate two-tailed p-values adjusted for multiple comparisons by Bonferroni correction. n = 35,446 Kdm6a-Mutant and n = 6,612 Kdm6a-WT tumor cells. Each individual dot represents a cell. Minimum and maximum values defined the range of violin plot. (e–g) Chromatin accessibility tracks for the average of each phenotype indicated (see Fig. 3c–g) at Myc (e), Mycl (f) and Mycn (g) from all ATAC-seq data from Fig. 3d.
Extended Data Fig. 7 Analysis of c-Myc and L-Myc in Kdm6a-Mutant vs. Kdm6a-WT Tumor Derived Cell Lines.
(a) Immunoblot analysis for c-Myc with a c-Myc antibody (Y69) in Kdm6a-WT and Kdm6a-Mutant SCLC cell lines cultured in ultra-low attachment flasks at late times (>2 months) after cell line generation. NCI-H82 and CORL279 human SCLC cell lines are included as benchmark controls for SCLCs with high c-Myc expression. (b, c) RT-qPCR for Myc mRNA expression in the early vs. late passage Kdm6a-WT and Kdm6a-Mutant tumor-derived cell lines shown as individual lines (b) or grouped by genotype (c). (d) Immunoblot analysis for L-Myc with a specific L-Myc antibody (Cell Signaling E3M5P) in Kdm6a-WT and Kdm6a-Mutant SCLC cell lines cultured in ultra-low attachment flasks at late times (>2 months) after cell line generation. CORL47 and NCI-H1092 human SCLC cell lines are included as benchmark controls for SCLC cell lines with high L-Myc expression. CORL47 and NCI-H1092 express the RLF-MYCL fusion and hence MYCL migrates at higher molecular weights compared to WT L-Myc. (e–h) RT-qPCR for Mycl (e,f) or Mycn (g,h) mRNA expression in the early vs. late passage Kdm6a-WT and Kdm6a-Mutant lines shown as individual lines (e,g) or grouped by genotype (f,h). For all RT-qPCR experiments, n = 6 biological independent experiments for each genotype (2 biological independent experiments for each cell line). Data are presented as mean values ± SEM. Statistical significance was calculated using unpaired, two-tailed students t-test and p-values are indicated. For all t-tests comparing Kdm6a-Mutant vs. Kdm6a-WT cell lines, early passage Kdm6a-Mutant vs. Kdm6a-WT were compared. (i, j) Tracks of H3K4me1, and KDM6A ChIP-seq for Mycl (i), and Myc (j) from ChIP-seq data from Fig. 5. Each track is the sum of 2 Kdm6a-WT cell lines (1014, 159-1) and 2 Kdm6a-Mutant cell lines (236L,236R) peaks with their respective input. (k) Immunoblot analysis for c-Myc and L-Myc in a Kdm6a-Mutant SCLC cell line (672-2) using highly specific antibodies with benchmark controls above. (l) Immunoblot analysis of 672-2 cells transduced with 4 independent Myc sgRNAs or a non-targeting control (sgControl) acutely after transduction and puromycin selection.
Extended Data Fig. 8 Correlation and GSEA Analyses of KDM6A, H3K27me3, H3K4me1, and H3K4me2 ChIP-seq Data in Kdm6a-WT and Kdm6a-Mutant SCLC Primary Cell Lines.
(a) Genome-wide correlation of log fold change in H3K27me3 ChIP-seq (Kdm6a-Mutant/Kdm6a-WT) vs. log fold change in KDM6A ChIP-seq at enhancers. (b) Genome-wide correlation of log fold change in H3K4me1 ChIP-seq (Kdm6a-Mutant/Kdm6a-WT) vs. log fold change in KDM6A ChIP-seq at enhancers. (c) Genome-wide correlation of log fold change in H3K4me2 ChIP-seq (Kdm6a-Mutant/Kdm6a-WT) vs. log fold change in KDM6A ChIP-seq at enhancers. For a-c, r Pearson correlation coefficient and p-values were calculated using a two-sided Pearson’s correlation test and are indicated. (d, e) Normalized Enrichment Score from GSEA of H3K27me3 ChIP-seq at transcription start sites of conserved ASCL1 target genes or conserved NEUROD1 target genes (d) or top 5 enriched Hallmarks (e) from the ChIP-seq data in Fig. 5. (f, g) Normalized Enrichment Score from GSEA of H3K4me2 ChIP-seq at enhancers of conserved ASCL1 target genes or conserved NEUROD1 target genes (f) or top 5 enriched Hallmarks (g) from the ChIP-seq data in Fig. 5. For d-g, p-values were generated by GSEA using a permutation test adjusted for multiple hypothesis testing using the Benjamini-Hochberg correction. Adjusted p-values are indicated.
Extended Data Fig. 9 NEUROD1 Expression after KDM6A Inactivation is Partially Mediated by KMT2A.
(a, b) Tracks of H3K27me3 and KDM6A ChIP-seq of Neurod1 (a) and Ascl1 (b) from ChIP-seq data from Fig. 5. Each track is the sum of the peaks of: 2 Kdm6a-WT tumor-derived cell lines for Kdm6a-WT (1014, 159-1) and 2 Kdm6a-Mutant tumor-derived cell lines for Kdm6a-Mutant (236L, 236R) with their respective input. (c) Cistrome analysis of transcription factors and chromatin regulators of Neurod1 gene in human hg38 (http://dbtoolkit.cistrome.org/) within 10 kilobases of the Neurod1 gene. (d) Pseudo-bulk differentially expressed gene (DEG) analysis from the scRNA-seq data in Fig. 3i in Kdm6a-Mutant vs Kdm6a-WT tumors cells of the genes in c. Non-parametric Wilcoxon rank sum test was used and p-values adjusted for multiple hypothesis testing are shown. (e) List of the top 20 candidate regulators of accessible peaks determined by LISA analysis of the 100 top differentially accessible peaks at TSSs with LFC > 1 sorted by ascending p-value in Kdm6a-Mutant vs. Kdm6a-WT using ATAC-seq data from Fig. 2a (see Supplementary Table 8, tab 1 for complete list).
Extended Data Fig. 10 Analyses of KMT2D Loss or Hypoxia with NEUROD1 Expression in SCLC.
(a) Correlation of KDM6A dependency vs. KMT2D depedency across hundreds of cancer cell lines25. KMT2D is the #1 co-dependency with KDM6A (p-value = 9.09×10–67 calculated using dependency map25). (b) GSEA using publicly available datasets of human SCLCs with KMT2D LOF mutations or KMT2D WT4 using genes upregulated in Kdm6a-Mutant vs. Kdm6a-WT SCLC GEMMs (see Fig. 1d). FDR q-value generated using GSEA, adjusted for multiple-hypothesis testing, is indicated. (c) Proportion of mixed ASCL1 and NEUROD1 human SCLCs from RNA-seq data from 3 independent data sets4,41,87 (see Methods) in SCLCs with KMT2D LOF mutations vs. KMT2D WT. Fisher’s exact test was used to generate p-value. For b,c, n = 14 KMT2D LOF, n = 142 KMT2D WT. See also Supplementary Table 9. (d, e) CRISPR amplicon sequencing of 1014 Kdm6a-WT cells transduced with 2 independent Kmt2d sgRNAs. sgRNA sequences are in blue and gene editing is in red. (f) RT-qPCR for Neurod1 in 1014 Kmt2d knockout cells in d,e 3 weeks after transduction. Data are relative to Actb and then normalized to Neurod1 expression in the sgControl (non-targeting) cell line. n = 4 biological replicates and error bars represent mean ± SEM. Statistical significance was calculated using unpaired, two-tailed students t-test and p-values are indicated. (g, h) Immunoblot (g) and histone blot (h) analyses in 1014 cells transduced with 4 independent Kmt2c and Kmt2d sgRNAs or a non-targeting control (sgControl) 30 days post-transduction. (i, j) Immunoblot analysis of NEUROD1 and ASCL1 expression of (i) NCI-H69 and (j) DMS79 ASCL1-positive human SCLC cells cultured under 5% or 21% O2 for 7 days. (k) Immunoblot analysis for HIF1α protein of cells lines in i,j. (l, m) GSEA analysis of scRNA-seq from human ASCL1 or NEUROD1 SCLC tumors34 using (l) HIF1 target gene list52 or (m) 24 genes induced by hypoxia51 (see Methods) (see Supplementary Table 10). FDR q-value adjusted for multiple hypothesis testing is indicated. Model-based Analysis of Single Cell Transcriptomics (MAST) was used to generate the GSEA expression profile.
Supplementary information
Supplementary Information
Supplementary Figs. 1–6 with accompanying figure legends.
Supplementary Table 1
Supplementary Table 1. Histology of autochthonous SCLC sgKdm6a-RPP (Kdm6a-Mutant) and sgControl RPP (Kdm6a-WT) tumours. Supplementary Table 2. RNA-seq data from autochthonous Kdm6a-Mutant versus Kdm6a-WT SCLC tumours. Supplementary Table 3. Bulk ATAC-seq data from autochthonous Kdm6a-Mutant versus Kdm6a-WT SCLC tumours. Supplementary Table 4. Bulk ATAC-seq data from autochthonous Kdm6aMutantAscl1Only and Kdm6aMutantAscl1HighNeurod1Low versus Kdm6aWT SCLC tumours. Supplementary Table 5. scRNA-seq differential expression analysis, Neurod47_score gene list, cell cycle phase gene lists and subpopulation cell counts. Supplementary Table 6. Differential expression analysis of RPM versus RPP and RPM early versus late. Supplementary Table 7. KDM6A, H3K4me1, H3K4me2 and H3K27me3 ChIP–seq data from Kdm6a-Mutant versus Kdm6a-WT SCLC primary cell lines. Supplementary Table 8. LISA analysis and Menin-ChIP–seq data. Supplementary Table 9. Human SCLCs KMT2D mutation status and bulk RNA-seq analysis. Supplementary Table 10. Hypoxia-related gene lists and GSEA expression profile.
Source data
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig./Table 1
Statistical source data.
Source Data Extended Data Fig. 2
Unprocessed western blots.
Source Data Extended Data Fig./Table 2
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Source Data Extended Data Fig./Table 3
Statistical source data.
Source Data Extended Data Fig./Table 4
Statistical source data.
Source Data Extended Data Fig. 7
Unprocessed western blots.
Source Data Extended Data Fig./Table 7
Statistical source data.
Source Data Extended Data Fig. 10
Unprocessed western blots.
Source Data Extended Data Fig./Table 10
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Duplaquet, L., Li, Y., Booker, M.A. et al. KDM6A epigenetically regulates subtype plasticity in small cell lung cancer. Nat Cell Biol 25, 1346–1358 (2023). https://doi.org/10.1038/s41556-023-01210-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-023-01210-z
This article is cited by
-
Small cells – big issues: biological implications and preclinical advancements in small cell lung cancer
Molecular Cancer (2024)
-
Genetically-engineered mouse models of small cell lung cancer: the next generation
Oncogene (2024)
-
Gut microbial metabolite facilitates colorectal cancer development via ferroptosis inhibition
Nature Cell Biology (2024)