Abstract
Adherens junctions (AJs) create spatially, chemically and mechanically discrete microdomains at cellular interfaces. Here, using a mechanogenetic platform that generates artificial AJs with controlled protein localization, clustering and mechanical loading, we find that AJs also organize proteolytic hotspots for γ-secretase with a spatially regulated substrate selectivity that is critical in the processing of Notch and other transmembrane proteins. Membrane microdomains outside of AJs exclusively organize Notch ligand–receptor engagement (LRE microdomains) to initiate receptor activation. Conversely, membrane microdomains within AJs exclusively serve to coordinate regulated intramembrane proteolysis (RIP microdomains). They do so by concentrating γ-secretase and primed receptors while excluding full-length Notch. AJs induce these functionally distinct microdomains by means of lipid-dependent γ-secretase recruitment and size-dependent protein segregation. By excluding full-length Notch from RIP microdomains, AJs prevent inappropriate enzyme–substrate interactions and suppress spurious Notch activation. Ligand-induced ectodomain shedding eliminates size-dependent segregation, releasing Notch to translocate into AJs for processing by γ-secretase. This mechanism directs radial differentiation of ventricular zone-neural progenitor cells in vivo and more broadly regulates the proteolysis of other large cell-surface receptors such as amyloid precursor protein. These findings suggest an unprecedented role of AJs in creating size-selective spatial switches that choreograph γ-secretase processing of multiple transmembrane proteins regulating development, homeostasis and disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Previously published genomic sequence data that were re-analysed here are available from Ensembl for E-cadherin (CDH1) and N-cadherin (CDH2) protein from homo sapiens (gene IDs ENSG00000039068 and ENSG00000170558). Source data are provided with this paper. All raw images acquired using confocal, epifluorescence and time-lapse microscopy, and additional data that support the findings of this study are available from the corresponding authors upon reasonable request. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
Code availability
Custom Python code used for automatic segmentation and junction intensity ratio analysis for Notch activation and truncation studies is available at https://github.com/kmsouthard/JunctionAnalysis. Co-localization analysis was carried out in ImageJ and the JACOP plugin available at https://imagej.nih.gov/ij/plugins/track/jacop.html. Custom ImageJ codes for other analyses, including quantification of Manders’ overlap coefficients, Pearson’s coefficients and lipid polarization analyses, are available at https://github.com/sukgi333/yonsei-notch-activation. Selected sgRNAs for the candidate target sites were evaluated with DeepSpCas9 sgRNA prediction tool (http://deepcrispr.info/DeepSpCas9/)99.
References
Manz, B. N. & Groves, J. T. Spatial organization and signal transduction at intercellular junctions. Nat. Rev. Mol. Cell Biol. 11, 342–352 (2010).
Chen, C. S., Tan, J. & Tien, J. Mechanotransduction at cell–matrix and cell–cell contacts. Annu. Rev. Biomed. Eng. 6, 275–302 (2004).
Südhof, T. C. Synaptic neurexin complexes: a molecular code for the logic of neural circuits. Cell 171, 745–769 (2017).
Südhof, T. C. & Malenka, R. C. Understanding synapses: past, present, and future. Neuron 60, 469–476 (2008).
Zeng, M. et al. Reconstituted postsynaptic density as a molecular platform for understanding synapse formation and plasticity. Cell 174, 1172–1187.e16 (2018).
Sanes, J. R. & Zipursky, S. L. Synaptic specificity, recognition molecules, and assembly of neural circuits. Cell 181, 536–556 (2020).
Narui, Y. & Salaita, K. Membrane tethered Delta activates Notch and reveals a role for spatio-mechanical regulation of the signaling pathway. Biophys. J. 105, 2655–2665 (2013).
Schmid, E. M. et al. Size-dependent protein segregation at membrane interfaces. Nat. Phys. 12, 704–711 (2016).
Salaita, K. et al. Restriction of receptor movement alters cellular response: physical force sensing by EphA2. Science 327, 1380–1385 (2010).
Dustin, M. & Shaw, A. Costimulation: building an immunological synapse. Science 283, 649–650 (1999).
Groves, J. T. & Kuriyan, J. Molecular mechanisms in signal transduction at the membrane. Nat. Struct. Mol. Biol. 17, 659–665 (2010).
van der Merwe, P. A. & Dushek, O. Mechanisms for T cell receptor triggering. Nat. Rev. Immunol. 11, 47–55 (2010).
Belardi, B., Son, S., Felce, J. H., Dustin, M. L. & Fletcher, D. A. Cell–cell interfaces as specialized compartments directing cell function. Nat. Rev. Mol. Cell Biol. 21, 750–764 (2020).
Case, L. B., Zhang, X., Ditlev, J. A. & Rosen, M. K. Stoichiometry controls activity of phase-separated clusters of actin signaling proteins. Science 363, 1093–1097 (2019).
James, J. R. & Vale, R. D. Biophysical mechanism of T-cell receptor triggering in a reconstituted system. Nature 487, 64–69 (2012).
Case, L. B., Ditlev, J. A. & Rosen, M. K. Regulation of transmembrane signaling by phase separation. Annu. Rev. Biophys. 48, 465–494 (2019).
Kovall, R. A., Gebelein, B., Sprinzak, D. & Kopan, R. The canonical Notch signaling pathway: structural and biochemical insights into shape, sugar, and force. Dev. Cell 41, 228–241 (2017).
Kopan, R. & Ilagan, M. X. G. The canonical Notch signaling pathway: unfolding the activation mechanism. Cell 137, 216–233 (2009).
Bray, S. J. Notch signalling in context. Nat. Rev. Mol. Cell Biol. 17, 722–735 (2016).
van Es, J. H. et al. Notch/γ-secretase inhibition turns proliferative cells in intestinal crypts and adenomas into goblet cells. Nature 435, 959–963 (2005).
Artavanis-Tsakonas, S. & Notch, M. R. R. L. Signaling: cell fate control and signal integration in development. Science 284, 770–776 (1999).
Boukhatmi, H., Martins, T., Pillidge, Z., Kamenova, T. & Bray, S. Notch mediates inter-tissue communication to promote tumorigenesis. Curr. Biol. 30, 1809–1820.e4 (2020).
South, A. P., Cho, R. J. & Aster, J. C. The double-edged sword of Notch signaling in cancer. Semin. Cell Dev. Biol. 23, 458–464 (2012).
Nicolas, M. et al. Notch1 functions as a tumor suppressor in mouse skin. Nat. Genet. 33, 416–421 (2003).
Lai, E. C. Notch signaling: control of cell communication and cell fate. Development 131, 965–973 (2004).
Haines, N. & Irvine, K. D. Glycosylation regulates Notch signalling. Nat. Rev. Mol. Cell Biol. 4, 786–797 (2003).
Stanley, P. & Okajima, T. Roles of glycosylation in Notch signaling. Curr. Top. Dev. Biol. 92, 131–164 (2010).
Wang, X. & Ha, T. Defining single molecular forces required to activate integrin and Notch signaling. Science 340, 991–994 (2013).
Gordon, W. R. et al. Mechanical allostery: evidence for a force requirement in the proteolytic activation of Notch. Dev. Cell 33, 729–736 (2015).
Seo, D. et al. A mechanogenetic toolkit for interrogating cell signaling in space and time. Cell 165, 1507–1518 (2016).
Marambaud, P. et al. A presenilin-1/γ-secretase cleavage releases the E-cadherin intracellular domain and regulates disassembly of adherens junctions. EMBO J. 21, 1948–1956 (2002).
Georgakopoulos, A. et al. Presenilin-1 forms complexes with the cadherin/catenin cell-cell adhesion system and is recruited to intercellular and synaptic contacts. Mol. Cell 4, 893–902 (1999).
Guillaume, E. et al. Flotillin microdomains stabilize cadherins at cell–cell junctions. J. Cell Sci. 126, 5293–5304 (2013).
Brasch, J., Harrison, O. J., Honig, B. & Shapiro, L. Thinking outside the cell: how cadherins drive adhesion. Trends Cell Biol. 22, 299–310 (2012).
Honig, B. & Shapiro, L. Adhesion protein structure, molecular affinities, and principles of cell–cell recognition. Cell 181, 520–535 (2020).
Yap, A. S., Gomez, G. A. & Parton, R. G. Adherens junctions revisualized: organizing cadherins as nanoassemblies. Dev. Cell 35, 12–20 (2015).
Falo-Sanjuan, J. & Bray, S. J. Membrane architecture and adherens junctions contribute to strong Notch pathway activation. Development 148, 19 (2021).
Taulet, N. et al. N-cadherin/p120 catenin association at cell–cell contacts occurs in cholesterol-rich membrane domains and is required for RhoA activation and myogenesis. J. Biol. Chem. 284, 23137–23145 (2009).
Causeret, M., Taulet, N., Comunale, F., Favard, C. & Gauthier-Rouvière, C. N-Cadherin association with lipid rafts regulates its dynamic assembly at cell–cell junctions in C2C12 myoblasts. Mol. Biol. Cell 16, 2168–2180 (2005).
Szaruga, M. et al. Alzheimer’s-causing mutations shift Aβ length by destabilizing γ-secretase-Aβn interactions. Cell 170, 443–456.e14 (2017).
Lee, S.-J. et al. A detergent-insoluble membrance compartment contains Aβ in vivo. Nat. Med. 4, 730–734 (1998).
Vetrivel, K. S. et al. Spatial segregation of γ-secretase and substrates in distinct membrane domains. J. Biol. Chem. 280, 25892–25900 (2005).
Chow, V. W., Mattson, M. P., Wong, P. C. & Gleichmann, M. An overview of APP processing enzymes and products. Neuromolecular Med. 12, 1–12 (2009).
Ehehalt, R., Keller, P., Haass, C., Thiele, C. & Simons, K. Amyloidogenic processing of the Alzheimer β-amyloid precursor protein depends on lipid rafts. J. Cell Biol. 160, 113–123 (2003).
Wahrle, S. et al. Cholesterol-dependent γ-secretase activity in buoyant cholesterol-rich membrane microdomains. Neurobiol. Dis. 9, 11–23 (2002).
Vetrivel, K. S. et al. Association of γ-secretase with lipid rafts in post-Golgi and endosome membranes. J. Biol. Chem. 279, 44945–44954 (2004).
Urano, Y. et al. Association of active γ-secretase complex with lipid rafts. J. Lipid Res. 46, 904–912 (2005).
Galbiati, F., Razani, B. & Lisanti, M. P. Emerging themes in lipid rafts and caveolae. Cell 106, 403–411 (2001).
Owen, D. M., Rentero, C., Magenau, A., Abu-Siniyeh, A. & Gaus, K. Quantitative imaging of membrane lipid order in cells and organisms. Nat. Protoc. 7, 24–35 (2012).
Komura, N. et al. Raft-based interactions of gangliosides with a GPI-anchored receptor. Nat. Chem. Biol. 12, 402–410 (2016).
Raghupathy, R. et al. Transbilayer lipid interactions mediate nanoclustering of lipid-anchored proteins. Cell 161, 581–594 (2015).
Kusumi, A. et al. Paradigm shift of the plasma membrane concept from the two-dimensional continuum fluid to the partitioned fluid: high-speed single-molecule tracking of membrane molecules. Annu. Rev. Biophys. Biomol. Struct. 34, 351–378 (2005).
Struhl, G. & Adachi, A. Requirements for presenilin-dependent cleavage of Notch and other transmembrane proteins. Mol. Cell 6, 625–636 (2000).
Palmer, W. H. & Deng, W.-M. Ligand-independent mechanisms of Notch activity. Trends Cell Biol. 25, 697–707 (2015).
Kouchi, Z. et al. p120 catenin recruits cadherins to γ-secretase and inhibits production of Aβ peptide. J. Biol. Chem. 284, 1954–1961 (2009).
Kim, J., Jeong, H., Southard, K. M., Jun, Y. & Cheon, J. Magnetic nanotweezers for interrogating biological processes in space and time. Acc. Chem. Res. 51, 839–849 (2018).
Kwak, M. et al. Small, clickable, and monovalent magnetofluorescent nanoparticles enable mechanogenetic regulation of receptors in a crowded live-cell microenvironment. Nano Lett. 19, 3761–3769 (2019).
Harrison, O. J. et al. The extracellular architecture of adherens junctions revealed by crystal structures of type I cadherins. Structure 19, 244–256 (2011).
Polacheck, W. J. et al. A non-canonical Notch complex regulates adherens junctions and vascular barrier function. Nature 552, 258–262 (2017).
Yang, Z., Yu, Z., Cai, Y., Du, R. & Cai, L. Engineering of an enhanced synthetic Notch receptor by reducing ligand-independent activation. Commun. Biol. 3, 1–7 (2020).
Mumm, J. S. et al. A ligand-induced extracellular cleavage regulates γ-secretase-like proteolytic activation of Notch1. Mol. Cell 5, 197–206 (2000).
Bolduc, D. M., Montagna, D. R., Gu, Y., Selkoe, D. J. & Wolfe, M. S. Nicastrin functions to sterically hinder γ-secretase–substrate interactions driven by substrate transmembrane domain. Proc. Natl Acad. Sci. USA 113, E509–E518 (2016).
Liang, S. I. et al. Phosphorylated EGFR dimers are not sufficient to activate Ras. Cell Rep. 22, 2593–2600 (2018).
Shah, S. et al. Nicastrin functions as a γ-secretase-substrate receptor. Cell 122, 435–447 (2005).
Sprinzak, D. et al. Cis-interactions between Notch and Delta generate mutually exclusive signalling states. Nature 465, 86–90 (2010).
Nandagopal, N. et al. Dynamic ligand discrimination in the Notch signaling pathway. Cell 172, 869–880.e19 (2018).
Ables, J. L., Breunig, J. J., Eisch, A. J. & Rakic, P. Not(ch) just development: Notch signalling in the adult brain. Nat. Rev. Neurosci. 12, 269–283 (2011).
Louvi, A. & Artavanis-Tsakonas, S. Notch signalling in vertebrate neural development. Nat. Rev. Neurosci. 7, 93–102 (2006).
Franco, S. J. & Müller, U. Shaping our minds: stem and progenitor cell diversity in the mammalian neocortex. Neuron 77, 19–34 (2013).
Das, R. M. & Storey, K. G. Apical abscission alters cell polarity and dismantles the primary cilium during neurogenesis. Science 343, 200–204 (2014).
Fujita, I. et al. Endfoot regeneration restricts radial glial state and prevents translocation into the outer subventricular zone in early mammalian brain development. Nat. Cell Biol. 22, 26–37 (2019).
Hatakeyama, J. et al. Cadherin-based adhesions in the apical endfoot are required for active Notch signaling to control neurogenesis in vertebrates. Development 141, 1671–1682 (2014).
de Strooper, B. & Karran, E. The cellular phase of Alzheimer’s disease. Cell 164, 603–615 (2016).
Selkoe, D. J. & Hardy, J. The amyloid hypothesis of Alzheimer’s disease at 25 years. EMBO Mol. Med. 8, 595–608 (2016).
O’Brien, R. J. & Wong, P. C. Amyloid precursor protein processing and Alzheimer’s disease. Annu. Rev. Neurosci. 34, 185–204 (2011).
Quintero-Monzon, O. et al. Dissociation between the processivity and total activity of γ-secretase: implications for the mechanism of Alzheimer’s disease-causing presenilin mutations. Biochemistry 50, 9023–9035 (2011).
Uemura, K. et al. N-cadherin-based adhesion enhances Aβ release and decreases Aβ42/40 ratio. J. Neurochem. 108, 350–360 (2009).
Bray, S. J. Notch signalling: a simple pathway becomes complex. Nat. Rev. Mol. Cell Biol. 7, 678–689 (2006).
J, R. W. et al. Evidence for a physical interaction between presenilin and Notch. Proc. Natl Acad. Sci. USA 96, 3263–3268 (1999).
Ray, W. J. et al. Cell surface presenilin-1 participates in the γ-secretase-like proteolysis of Notch. J. Biol. Chem. 274, 36801–36807 (1999).
Morsut, L. et al. Engineering customized cell sensing and response behaviors using synthetic Notch receptors. Cell 164, 780–791 (2016).
Roybal, K. T. et al. Precision tumor recognition by T cells with combinatorial antigen-sensing circuits. Cell 164, 770–779 (2016).
Yang, G. et al. Structural basis of Notch recognition by human γ-secretase. Nature 565, 192–197 (2018).
Collado, M. S. et al. The postnatal accumulation of junctional E-cadherin is inversely correlated with the capacity for supporting cells to convert directly into sensory hair cells in mammalian balance organs. J. Neurosci. 31, 11855–11866 (2011).
Hao, W. et al. Regulation of beta-amyloid production in neurons by astrocyte-derived cholesterol. Proc. Natl Acad. Sci. USA 118, e2102191118 (2021).
Maeda, K., Takemura, M., Umemori, M. & Adachi-Yamada, T. E-cadherin prolongs the moment for interaction between intestinal stem cell and its progenitor cell to ensure Notch signaling in adult Drosophila midgut. Genes Cells 13, 1219–1227 (2008).
Soh, B. S. et al. N-cadherin prevents the premature differentiation of anterior heart field progenitors in the pharyngeal mesodermal microenvironment. Cell Res. 24, 1420–1432 (2014).
Kwon, C. et al. A regulatory pathway involving Notch1/β-catenin/Isl1 determines cardiac progenitor cell fate. Nat. Cell Biol. 11, 951–957 (2009).
Yoshida, K., Murata, M., Yamaguchi, T., Matsuzaki, K. & Okazaki, K. Reversible human TGF-β signal shifting between tumor suppression and fibro-carcinogenesis: implications of Smad phospho-isoforms for hepatic epithelial–mesenchymal transitions. J. Clin. Med. 5, 7 (2016).
Arefin, B., Parvin, F., Bahrampour, S., Stadler, C. B. & Thor, S. Drosophila neuroblast selection is gated by Notch, Snail, SoxB, and EMT gene interplay. Cell Rep. 29, 3636–3651.e3 (2019).
Dustin, M. L. & Groves, J. T. Receptor signaling clusters in the immune synapse. Annu. Rev. Biophys. 41, 543–556 (2012).
Bakalar, M. H. et al. Size-dependent segregation controls macrophage phagocytosis of antibody-opsonized targets. Cell 174, 131-–142.e13 (2018).
Pruessmeyer, J. et al. A disintegrin and metalloproteinase 17 (ADAM17) mediates inflammation-induced shedding of syndecan-1 and −4 by lung epithelial cells. J. Biol. Chem. 285, 555–564 (2010).
Kim, J. et al. Single-cell mechanogenetics using monovalent magnetoplasmonic nanoparticles. Nat. Protoc. 12, 1871–1889 (2017).
Dudok, B. et al. Cell-specific STORM super-resolution imaging reveals nanoscale organization of cannabinoid signaling. Nat. Neurosci. 18, 75–86 (2014).
Moss, M. L. & Rasmussen, F. H. Fluorescent substrates for the proteinases ADAM17, ADAM10, ADAM8, and ADAM12 useful for high-throughput inhibitor screening. Anal. Biochem. 366, 144–148 (2007).
Lee, H. J. et al. En bloc and segmental deletions of human XIST reveal X chromosome inactivation-involving RNA elements. Nucleic Acids Res. 47, 3875–3887 (2019).
Pulido-Quetglas, C. et al. Scalable design of paired CRISPR guide RNAs for genomic deletion. PLoS Comput. Biol. 13, e1005341 (2017).
Kim, H. K. et al. SpCas9 activity prediction by DeepSpCas9, a deep learning–based model with high generalization performance. Sci. Adv. 5, eaax9249 (2019).
Ramakrishna, S. et al. Gene disruption by cell-penetrating peptide-mediated delivery of Cas9 protein and guide RNA. Genome Res. 24, 1020–1027 (2014).
Gopalappa, R., Suresh, B., Ramakrishna, S. & Kim, H. Paired D10A Cas9 nickases are sometimes more efficient than individual nucleases for gene disruption. Nucleic Acids Res. 46, e71 (2018).
Guschin, D. Y. et al. A rapid and general assay for monitoring endogenous gene modification. Methods Mol. Biol. 649, 247–256 (2010).
van der Spoel, D. et al. GROMACS: fast, flexible, and free. J. Comput. Chem. 26, 1701–1718 (2005).
de Jong, D. H. et al. Improved parameters for the Martini coarse-grained protein force field. J. Chem. Theory Comput. 9, 687–697 (2013).
Hess, B., Bekker, H., Berendsen, H. J. C. & Fraaije, J. G. E. M. LINCS: a linear constraint solver for molecular simulations. J. Comput. Chem. 18, 1463–1472 (1997).
Michaud-Agrawal, N., Denning, E. J., Woolf, T. B. & Beckstein, O. MDAnalysis: a toolkit for the analysis of molecular dynamics simulations. J. Comput. Chem. 32, 2319–2327 (2011).
Chavent, M. et al. Methodologies for the analysis of instantaneous lipid diffusion in MD simulations of large membrane systems. Faraday Discuss. 169, 455–475 (2014).
Acknowledgements
The authors thank S. Blacklow (Harvard U.), C. Miller (King’s College London) and K. Shimamura (Kumamoto U.) for the kind gifts of Notch, APP and DN-cadherin plasmids, respectively. We also thank A. Balmain, M. Moasser and E. Collison (UCSF) for sharing cell lines. D. Fletcher (UC Berkeley), A. Joffe (UC Berkeley) and D. Al-Rawi (Stanford U.) provided insightful discussion. For reagents, technical support, and discussions, we thank the Kim, Cheon, Gartner and Jun laboratories, as well as the Nikon Imaging Center and Wynton at UCSF. M.K. was supported by a Life Science Research Foundation fellowship as the Shurl and Kay Curci Foundation fellow, and by Burroughs Wellcome Travel Fund. This work was supported by NRF, NRF-2021R1F1A1063378 (M.K.), NRF-2018R1A5A1025511 (D.S.) and NRF-2017R1A2B3004198 (H.H.K.), HI17C0676 from Korean Ministry of Health and Welfare (H.H.K.), 5R01NS047229 from National Institute on Aging (NIA) and the National Institute of Health (NIH) (A.G.), 5R01AG008200 from National Institute on Aging (NIA) and the National Institute of Health (NIH) (N.K.R.), IBS-R026-D1 from IBS (M.K., H.H.K. and J.C.), NRF-2019R1A2C1085712 (Y.H.K.), the UCSF Center for Cellular Construction (an NSF Science and Technology Center, no. DBI-1548297) (Z.J.G.), U01CA244109 from the National Cancer Institute (Z.J.G.), 1R01GM112081, 1R01GM126542-01 and R35GM134948 from the National Institute of General Medical Science (NIGMS) and the NIH (Y.Jun), 1R21AG072232-01 from the National Institute on Aging (NIA) and the NIH (M.L.K. and Y.Jun), R00CA226366 from the National Cancer Institute (M.L.K) and NIH, and the UCSF Program for Breakthrough Biomedical Research (PBBR) funded in part by the Sandler Foundation (M.L.K and Y.Jun). Z.J.G. is a Chan Zuckerberg BioHub Investigator.
Author information
Authors and Affiliations
Contributions
M.K., K.M.S., Z.J.G. and Y.Jun conceived the ideas and designed research; M.K. and K.M.S. constructed plasmids, generated cell lines and performed confocal microscopy. M.K. performed mechanogenetics, truncation study, spatial mutation, immunoblot analysis, reporter cell assay and APP experiment. K.M.S. performed Notch exclusion and activation experiments, designed truncation study and wrote custom Python image analysis scripts. W.R.K. performed animal experiment, flotillin staining and mechanogenetic experiment. A.L. performed confocal imaging and lipid order imaging experiments. N.H.K. performed coarse-grained MD simulation. R.G. generated cadherin-KO cells. M.A. synthesized magnetic nanoparticles. H.J.L., S.H.C. and K.N. helped with confocal imaging/western blot analysis. Y.Jung helped with lipid order imaging experiment. M.K.K. performed ELISA analysis of Aβ secretion from CDH-KO cells. S.H.C. performed western blot analysis. J.F. and D.S. performed initial proof-of-concept experiments. A.G. and N.K.R. provided anti-PS1 antibodies and critically read the manuscript. M.L.K. helped with the VE-cad experiment, H.H.K., Y.H.K. and J.C. oversaw CRISPR–Cas9 KO experiment, MD simulation and magnetic nanoparticle synthesis, respectively. Z.J.G. oversaw and supervised all spatial mapping and Notch truncation experiments. Y.Jun oversaw and supervised all aspects of the study. M.K. and K.M.S. analysed data. M.K., K.M.S., Z.J.G. and Y.Jun wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Z.J.G. is an equity holder in Scribe Biosciences and Provenance Bio and is an advisor for Serotiny. Y.H.K. is a vice president of, and has a financial interest in, Imnewrun. J.F. is a co-founder and a chief technology officer of, and has a financial interest in, Serotiny. The other authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks Alf Honigmann, Alpha Yap and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Interfacial membrane compartmentalization choreographs the sequential molecular processing of Notch.
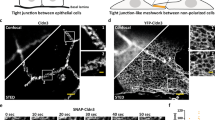
(a) Representative confocal images showing Dll1 (D1), Notch1 (N1), or presenilin1 (PS1) distributions at the interfacial junction between two cells exclusively expressing D1 or N1 in the presence of TAPI2. (left) A maximum intensity projection image of a LRE µdomains. Scale bar, 5 µm. (right) Magnified individual fluorescent channel and merged images of the yellow boxed region. The white dashed area and the white arrows indicate the cellular interface and the RIP µdomains, respectively. Scale bars, 3 µm. (b) Representative confocal images showing the interfacial junction between two cells exclusively expressing N1 and D1 in the presence of ADAM10/17 shRNA. The cell on top labeled ‘N1’ expresses N1 only, while the bottom cell labeled ‘D1’ expresses D1 only. (left) A maximum intensity projection image of a LRE µdomains. Scale bar, 5 µm. (top-right) Magnified individual fluorescence channel images of the boxed region. Scale bar, 3 µm. (bottom-right) Z-resliced images showing the sections of the cellular interfaces. Scale bar, 2 µm. (c) Representative confocal images showing N1, and PS1 distributions at the interfacial junction after washed out to remove TAPI2 and DAPT inhibition. With TAPI2 and siRNA treatment, Notch and γ-secretase compartmentalization was observed, while upon DAPT wash-out, Notch signals at the RIP µdomains disappeared. (left) A maximum intensity projection image of the cells showing enriched Notch mCherry signal at the RIP µdomains. Scale bar, 10 µm. (Top-right) Magnified individual fluorescence channel images of the boxed region. Scale bar, 2 µm. (right) Z-resliced images showing the sections of the cellular interfaces. Scale bar, 2 µm. (d) A representative confocal image showing Notch1 (N1) and presenilin1 (PS1) distribution at cellular interface. DAPT was added to inhibit S3 cleavage. (left) A maximum intensity projection image of a LRE µdomains. Scale bar, 10 µm. (right) Magnified individual fluorescent channel and merged images of the boxed region. Scale bar, 3 µm. (e) Representative confocal images showing intracellular distribution of PS1, E-cadherin (Ecad), and Notch1 (N1) in the presence of DAPT. (left) A maximum intensity projection image. Scale bar, 10 µm. (right) Magnified individual fluorescent channel and merged images of the specified intracellular region (a white dashed box). The yellow dashed circles represent intracellular puncta enriched with PS1 with no Ecad or N1 signals. The red dashed circle indicated with the red arrow represents an intracellular punctum showing all PS1, Ecad, and N1 fluorescence signals. Scale bar, 2 µm. (f) Manders’ overlap coefficients (MOCs) for quantitative assessment of PS1 with Notch1 at cell surface, PS1 with Ecad at cytosol, and PS1 with Notch1 at cytosol, respectively. Each dot represents the MOC of a selected cell surface or cytosol. In the box-whisker plot, the boxes show the 25th to 75th percentiles, and the whiskers extend to the maxima and the minima. Solid lines indicate median, respectively. n = 12 (Notch1 at surface), 7 (Ecad in cytosol), and 7 (Notch1 in cytosol) biologically independent cells across 2 independent experiments; ordinary one-way ANOVA with Tukey’s multiple comparison testing.
Extended Data Fig. 2 Adherens junctions reorganize Notch signaling biomolecules into two distinct membrane microdomains (µdomains).
(a) Representative confocal immunofluorescence images showing distribution of presenilin1 (PS1) across entire cell membranes. White and yellow arrows indicate the cell-cell interfacial membranes and the cell membranes without contact, respectively. (top) A maximum intensity projection of the wide-field confocal z-stacks. Scale bar, 20 µm. (bottom) A confocal z-resliced image along the representative membranes of two contacting cells. A white arrow indicates the cell-cell interfacial membrane where strong PS1 signals were observed. Yellow dashed lines represent the cell membranes without cell-cell contacts. Scale bar, 3 µm. (b) Maximum intensity projection images showing distribution of PS1 and E-cadherin AJs (Ecad) relative to LRE-µdomain with Notch1-Dll1 pair enrichment at the interfacial membrane in the presence of TAPI2. PS1 and Dll1 were visualized by immunostaining, Notch and AJs by expression of their respective mCherry- or EGFP-fusion constructs. AJs showed nearly identical spatial distribution with the γ-secretase enriched RIP-µdomain, but exhibited inverse distribution with the LRE- µdomain. Scale bar, 5 µm. (c) Representative wide-field confocal immunofluorescence images showing PS1 (magenta) enrichment within AJs (green). Scale bar, 50 µm (low-magnification), 10 µm (zoom-in). (d) Confocal images of U2OS cells co-expressing Ecad-GFP (green) and SNAP-NFL-mCherry (red), and immunostained with PS1 antibody (cyan). Scale bars, 10 µm, 2 µm, and 2 µm for maximum intensity projection, zoomed-in, and z-resliced images, respectively. (e) Paired analysis of Manders’ overlap coefficients of E-cadherin and Notch signals over PS1 in multiple cells (n = 9 examined across 3 independent experiments). Two-tailed paired Student’s t test. (f) Confocal z-resliced images showing PS1 (magenta) and Nicastrin (NCT, red) distribution relative to cadAJs (green). Scale bar, 2 µm. (g) Analysis of Manders’ overlap coefficients (MOC) of E-cadherin, PS1, and DAPI over Nicastrin in multiple cells (n = 6 examined across 2 independent experiments). One-way ordinary ANOVA followed by Tukey’s multiple comparison testing. (h,i) Representative confocal immunofluorescence images showing (h) ADAM17 and (i) ADAM10 distribution relative to AJs. ADAM17 exhibited no preferential localization relative to AJs. Scale bar, 20 µm. (j) Box-whisker plots showing Manders’ overlap coefficients (MOCs) of PS1 (green), Notch (red), Dll1 (purple), and ADAM10 (yellow) relative to AJs. Each dot represents the MOC of a selected AJ. Boxes and whiskers denote the inner-quartile and full ranges. Colored lines and (+) marks indicate median and mean, respectively (n = 15 (Dll1), 11 (Notch1), 14 (PS1), and 19 (ADAM10) cells examined over two independent experiments; ns, not significant; ordinary one-way ANOVA with Tukey’s test). (k) Representative confocal fluorescence images of HaCaT cells immunostained with anti-Notch1 (red) and anti-N-cadherin (green). (l) Confocal images of MDCK cells expressing Ecad-GFP (green) and SNAP-NFL-mCherry. Notch receptors were labeled with BG-Alexafluor647 (magenta). (m) Confocal images of HUVECs expressing SNAP-NF-mCherry and immunostained with vascular endothelial cadherin (VE-cad) antibody. (l-m) Scale bar, 10 µm and 2 µm for maximum intensity projection and z-resliced images, respectively. (n) Polarized MDCK cells grown on a transwell filter. Notch, actin, and nucleus were immunostained with BG-AF647, phalloidin-488, and DAPI, respectively. Scale marked every 5 µm for 3D construction. Scale bar, 2 µm.
Extended Data Fig. 3 Spatial dynamics of Notch receptors relative to AJs during cell-surface activation.
(a) A schematic to capture the spatial distribution of Notch intermediates during the cell-surface activation pathway. (b) Confocal z-resliced images showing Notch distribution (red) relative to AJ (green) from the cells without Dll4 activation (i), treated with Dll4 and TAPI2 (ii), treated with Dll4 and DAPT (iii), and washed out to remove DAPT inhibition (iv). Scale bar, 3 µm. (c) Quantification of Notch signal enrichment at the AJs during the activation. Notch enrichment (IIN/IOUT) is calculated as the ratio of average Notch fluorescence intensity within AJs (IIN) and outside AJ (IOUT). The enrichment factor of Dil is present as a control showing AJ-independent distribution. In the box-whisker plot, the boxes show the 25th to 75th percentiles, and the whiskers extend to the 10th and 90th percentiles, with individual data points above the whiskers shown for the lowest and highest 10% of each dataset. Solid lines and (+) marks indicate median and mean, respectively. n = (left to right) 13, 4, 25, 17 cells analyzed across three independent experiments. *** P = 0.0005, ****P < 0.0001, ns: non-significant, one-way ordinary ANOVA followed by Tukey’s multiple comparison testing. (d) Representative time-course confocal z-resliced images showing S2-cleaved Notch at AJs as a function of time after DAPT removal. The NICD signal (red) at the AJ gradually decreases, indicating NICD release. Images shown here are not from identical cells, but represent a general trend of NICD signal at AJs for each time point. Scale bar, 5 µm (e) Quantification IIN/IOUT ratio as a function of time after DAPT washout. Data are the mean ± s.d of n = 25 (+DAPT), 9 (0 hr), 10 (0.5 hr), 6 (1.5 hr), 8 (3 hr), 17 (12 hr), and 14 (-Dll4) biological replicates examined across 3 independent experiments.
Extended Data Fig. 4 Interrogation of the mechanism underlying γ-secretase recruitment into AJs.
(a) Representative confocal fluorescence images showing the PS1 and Flot1 distribution relative to native cell-cell AJs. (left) A maximum intensity projection image of merged channels. Scale bar, 10 µm. (center) Magnified images showing greater details of the boxed region. Scale bar, 2 µm. (right) Z-resliced images showing the sections of the AJs. Scale bar, 2 µm. Line profiles of fluorescence signals from E-cadherin, PS1, and Flot1 along the white dashed lines in the z-resliced images. (b) Cross-correlation analysis of E-cadherin and PS1 over Flot1. Both Flot1 and PS1 fluorescence intensities exhibited strong positive correlation with the AJ. The solid curves and the shades indicate means and s.e.m, respectively. n = 7. (c) Confocal fluorescence images showing PS1 and Flot1 localization at artificial AJs by mechanogenetics. E-cadherin and Flot1 were labeled with fluorescent tags. Endogenous PS1 was immunostained after fixation. Scale bar, 20 µm. (d) Magnified confocal images showing PS1 and Flot1 localization before (0 min) and after (30 min) the formation of artificial AJs via mechanogenetics. Scale bar, 20 µm. (e) Magnified confocal images showing strong accumulation of Actin at the artificial AJ by mechanogenetics. Scale bar, 5 µm. (f) Representative confocal fluorescence images showing no enrichment of Actin signal at the artificial AJs during MβCD treatment. Scale bar, 5 µm. (g) Fluorescence amplification factors (I/Io) of Ecadherin, Flot1, and PS1 localization in response to the mechanogenetic formation of AJs quantified for multiple replicates. The effect of MβCD treatment on Flot1 and PS1 relocalization was assessed. Data are the mean ± s.e.m. of n = 3 (Ecad, -MβCD), 5 (Ecad, +MβCD), 3 (Flot1, -MβCD), 5 (Flot, +MβCD), 2 (PS1, -MβCD), and 2 (PS1, + MβCD) biologically independent mechanogenetic experiments. One-way analysis of variance (ANOVA) with Tukey’s multiple comparisons test. (h) Representative fluorescence image showing the mechanogenetic formation of Ecad-ΔICD cluster. A white dashed box indicates the subcellular region where the µMT was applied. Scale bar, 10 µm. (i) Confocal image showing that the mechanogenetically induced Ecad-ΔICD clusters did not recruit Flot1. Scale bar, 20 µm. Zoom-in of the white frame-marked region of µMT application is shown on the right. Scale bar, 5 µm. (j) Spatial distribution of F-actin and PS1 in response to Ecad-ΔICD clustering. Neither F-actin nor PS1 were localized at the cluster region. Scale bar, 5 µm.
Extended Data Fig. 5 AJ-induced membrane juxtaposition drives Notch exclusion via size-dependent protein segregation.
(a) Additional artificial AJs showing Notch recruitment. Scale bar, 2 µm. (b) Time-lapse epifluorescence images (were acquired before micromagnetic tweezer (µMT) stimulation and then at 10, 20, 30 minutes of the µMT application. Gradual MFN and E-cadherin clustering was clearly seen, followed by Notch accumulation at the AJ. Scale bar, 2 µm. (c) Kinetics of signal enrichments at the artificial AJ shown in the panel (b). This is a representative result from n = 3 artificial AJs from 3 independent experiments. (d) Representative western blot for total Notch ICD from the U2OS cells stably expressing Notch1 truncation variants. The blot was probed with anti-Notch1-ICD. The same lysates were used in (Fig. 3F). The asterisk (*, upper band) represent the intact Notch truncation variants. Expected molecular weight of NFL, NΔEGF1-25, NΔEGF, and NEXT are 250 kD, 150 kD, 110 kD, and 95 kD, respectively. The cross (+, lower) represents the reduced protein band of 70 kD. All variants contain the SDS/DTT-sensitive link that produces the protein band corresponding to the polypeptide of Notch ECD and transmembrane-intracellular domain (TMICD). The number shown in each lane indicates the quantified band intensity of the corresponding lane normalized to that of NEXT variant. The intensity is calculated by summing the intensities measured from two bands detected in each lane. (e) Western blot quantification of cleaved NICD levels over total Notch levels. Data are the mean ± s.e.m. of n = 3 experiments. One-way analysis of variance (ANOVA) with Tukey’s multiple comparisons test. (f) Method to quantify Notch enrichment. Please see methods for more details. (g) Representative confocal images and enrichment factors (IIN/IOUT) of Dil membrane staining dye distribution relative to AJs. Scale bars, 10 µm (maximum intensity projection), 3 µm (z resliced images). (h) Time series of confocal z-resliced images showing the enrichment of NΔEGF (red) at the AJ (green) under DAPT treatment (t = 0), and the dissipation during DAPT washout (t ≥ 2). Scale bar, 3 µm. (i) Single-cell traces showing the time-course of the decline of NΔEGF enrichment factor at the AJs during DAPT washout (mean ± s.e.m.; n = 4 independent single-cell experiments). (j) Quantification of changes in NICD signal from these four cells at the AJs and non-AJ membrane, at t = 0 (green, before washout) and t = 6 hr (red, after DAPT washout). AJs and non-AJ membrane were detected based on thresholding and automatic segmentation using the custom-built script. Intracellular-mCherry signal significantly decreased at the AJs, but not at non-AJ membranes (*P = 0.035, ns: P = 0.075, Student’s t and Wilcoxon test, n = 4 cells examined across 2 independent experiments). (k) Confocal z-resliced images showing the distribution of extracellular SNAP (purple) and intracellular mCherry (red) tags of NΔEGF relative to AJs (green) after DAPT removal. Scale bar, 3 µm. (l, m) Nuclear location of NICD released from cell membrane that recombinantly expresses NΔEGF. (l) Confocal fluorescence images of U2OS cells expressing SNAP-NΔEGF-mCherry and Ecad-GFP. (Upper) Cells treated with TAPI2 only. White arrowheads indicate the cells with nuclear NICD-mCherry accumulation. (Lower) Cells treated with both TAPI2 and DAPT. Scale bar, 20 µm. (m) Quantification of the ratio of nucleus-to-cytosolic mCherry signals in cells with DAPT (n = 39 AJs) and those without DAPT (n = 21 AJs) from 3 independent experiments. A box and a whisker indicate the interquartile and the full range, respectively. Colored lines indicate median. Two tailed unpaired Student’s t-test.
Extended Data Fig. 6 Spatial mutations of Notch to study the effect of membrane compartmentalization on the signaling.
A representative western blot of lysate from cells expressing NΔEGF and Halo-Ecad-GFP after 2 hr incubation with or without DNA crosslinkers. The blot was labelled with anti-SNAP (a) and anti-Ecadherin (b) antibodies. The expected mass of NΔEGF, E-cadherin monomer, and the complex with the Notch construct and E-cadherin forming a heterodimer are 90 kD, 158 kD, and 230 kD, respectively. β-actin detection was used to assess protein loading. In both blots, predicted bands representing Notch-E-cadherin heterodimers (solid black lines) and SNAP-NΔEGF-mCherry or Halo-Ecad-GFP monomers (dashed black lines) are indicated. (c) A representative maximum intensity projection of XY images (i) and a XZ-resliced image (ii) showing exclusion of full-length Notch1 (SNAP-NFL-mCherry) from the AJs (green) after the DNA crosslinking. (d) Representative confocal maximum intensity projection images showing the distribution of NΔEGF relative to the AJs after crosslinking. Cells were treated with or without DAPT. Scale bar, 10 µm. (e) Single-cell confocal z-resliced images showing intracellular mCherry signal at the AJ under DNA and DAPT treatment (left) and after washing out DAPT (right). Removing DAPT elicited a significant reduction in mCherry signal intensity from the AJ. Scale bar, 5 µm. (f) Paired analysis of multiple cells expressing NΔEGF in enrichment factor (IIN/IOUT) after DAPT washout. Each dot represents IIN/IOUT value before and after DAPT washout from a single cell. Each line corresponds to the IIN/IOUT changes before and after DAPT washout in a same single cell (paired two-tailed Student’s t test; n = 6 cells examined across 2 independent experiments). (g) Larger area (1 ×1 mm2) confocal fluorescence images shown in Fig. 4f. Scale bar, 200 µm. (h) Dynamic light scattering spectra of BG-modified macromolecules used in the experiment to induce spatial mutation of NEXT in Fig. 4d-g. (i) Western blot analyses showing that spatial mutations of NEXT alter the level of Notch activation. Representative western blot from three independent experiments. The blot was probed with specific antibodies for cleaved NICD (Val1744) and β-actin. Each lane was loaded with the lysates from NEXT-expressing cells incubated with different BG-modified polymers or proteins for 20 h. The lysate from NFL was used as control. (j,k) Representative images of artificial AJs formed in live cells. Cells treated with both TAPI2 and DAPT (j) or with only TAPI2 but no DAPT (k). Magnified images were shown in lower panels. An intense mCherry signal was observed at the artificial AJ with TAPI2 and DAPT treatment, while no enrichment of Notch1 signal was seen from cells without DAPT. Scale bar, 5 µm (low-magnification), 2 µm (zoomed-in images). (l) Representative time-lapse images showing Notch signal activation in UAS-Gal4 reporter cells with artificial AJs (white arrows). Cells were cultured in the presence of TAPI2 and no source of S2 cleavage. Neighboring cells without magnetic stimulation were used as internal negative controls. Images were acquired using epifluorescence imaging every 2 hr for 24 hr. Scale bar, 50 µm.
Extended Data Fig. 7 Generation of U2OS SNAP-NFL-Gal4 fluorescence reporter cell lines lacking E-cadherin via CRISPR/Cas9.
(a,b) Schematic representation of human CDH1 (a) and CDH2 (b) gene structure and targeted segmental deletion sites. The sixteen exons are shown in orange boxes (E1-E16). Red arrowheads indicate the sgRNA-binding sites (E13LT1, E13LT2, E14RT1 and E14RT2 for CDH1, 2E1LT1 and 2E2RT1 for CDH2). The targeted segmental deletions of 4.6 kb for CDH1 and 940 bp for CDH2 knockout are shown with a black line with red arrow tips, respectively. Purple arrows represent PCR primers used for the T7E1 assay and detection of alleles with targeted deletions, respectively. (c) CDH1 mRNA expression levels in CDH1 KO clone #3 and #4 were determined using qRT-PCR. CDH1 expression levels in the selected clones containing a segmental deletion were quantified relative to CDH1 mRNA levels of the wild-type U2OS SNAP-NFL-Gal4 cells. Clone #3 was used for subsequent experiments. Data are the mean ± s.d. of n = 3 biologically independent samples. (d) qRT-PCR analysis of CDH2 mRNA expression levels in U2OS SNAP-NFL-Gal4 reporter cells (WT), CDH1 KO clone #3 (Ecad-KO), and a negative control sample (no primer pair added). CDH2 mRNA levels in both WT and Ecad-KO cells were quantified relative to the negative control sample. Both WT and Ecad-KO cells showed negligible CDH2 mRNA levels, indicating that Ecad-KO cells have minimal mRNA expression of both CDH1 and CDH2. Data are the mean ± s.e.m. of n = 3 biologically independent samples; one-way ordinary ANOVA test. (e) Western blot analyses of cleaved NICD levels in the wild-type SNAP-NFL-Gal4 cells, CDH1 knock-out (Ecad-KO) cells, and Ecad-KO cells with recombinant E-cadherin transfection. (top) A representative image of immunoblotting. (bottom) Quantification of cleaved NICD levels. The average intensity of NICD bands relative to β-actin bands was quantified and then normalized to that of Ecad-KO cells. Data are the mean ± s.d. of n = 5 biological replicates; one-way ANOVA followed by Tukey’s test). (f,g) Representative epi-fluorescence images showing Notch activation in co-culture of Ecad-KO cells with Ecad-KO + Ecad cells (f) or with Ecad-KO + Ncad cells (g). Ecad-KO cells shows no GFP signal (green) while Ecad-KO + Ecad or Ecad-KO + Ncad cells show robust GFP signal indicative of reintroduction of E- or N-cadherin. Scale bar, 50 µm.
Extended Data Fig. 8 Additional immunofluorescence images showing that the AJ-mediated membrane compartmentalization modulates neural progenitor cell (NPC) differentiation.
(a) Immunostaining of the subventricular zone (SVZ) in the lateral ventricle (LV) of the E13.5 DAPT-treated mouse brain. Notch was colocalized at NAJ, visualized by immunostaining with anti-N-cadherin and anti-Notch1 antibodies. (i) Representative lower magnification image. The indicated area (a white box) is magnified and rotated 90° clockwise in the lower panel. Scale bar, 100 µm. (ii) Magnified view of the region indicated with a white dashed box in the (i) lower panel. Scale bar, 2.5 µm. (iii) Line profiles of N-cadherin and Notch distributions. (b) Additional confocal images of coronal sections of developing mouse brain retrovirally infected with dominant negative form of E-cadherin vector (DN-Ecad-EGFP). Transduced cells differentiated into post-mitotic neurons can be identified as EGFP + /Tuj1 + , while those remained as NPCs with plasmid transfection are only EGFP + . (left) Low-magnification images. Insets show the magnified image of a representative single cell immunostained for myc-tag. (right) Magnified view of the boxed region. Scale bar, 50 µm. (c) Ratio of GFP/Tuj1-double positive cells to total Tuj1-positive post-mitotic neurons in these two conditions. Data are the mean ± s.e.m. of n = 3 (vector control) and 5 (DN-Ecad-EGFP) biologically independent animals. Two-tailed unpaired Student’s t test.
Extended Data Fig. 9 Amyloid precursor proteins (APPs) with intact YENPTY motif show size-dependent spatial segregation and membrane proteolysis, consistent with APP lacking the YENPTY motif.
(a) Representative confocal maximum intensity projection (right) and z-resliced (left) images of U2OS cells co-expressing N-cadherin (green) and full-length APP (red). To capture the spatial distribution of the APP intermediates, cells were cultured with a combination of α-, β-, and γ-secretase inhibitors. Scale bar, 3 µm (max. projection) and 2 µm (z-resliced). (b) The spatial redistribution of APP relative to the NAJs was quantified using Manders’ overlap coefficient (MOC). Data are presented as boxes and whiskers, representing interquartile and min-to-max ranges, respectively; n = 7 (i), 4 (ii), and 10 (iii) NAJs examined over two independent experiments. each detected from a single cell. One-way ANOVA followed by Tukey’s multiple comparison test.
Supplementary information
Supplementary Video 1
Supplementary Video S1. Time lapse of Notch activation in cell groups versus solitary cells. Left column: representative movies of U2OS cells expressing SNAP-NFL-Gal4 and UAS-H2B-mC with robust cadAJs within high-density culture. The cells in this group exhibited robust increase in nuclear mCherry fluorescence signal. Right columns: representative movies of U2OS cells having no prior contact with other cells (n = 9). The solitary cells exhibited negligible increase in nuclear mCherry fluorescence signals. Each movie is digitally cropped and centred on the specific representative cells. SNAP-NFL-Gal4 expression and mCherry fluorescence are pseudo-coloured as green and red, respectively. Time stamps are relative to doxycycline addition.
Supplementary Video 2
Supplementary Video S2. Time-lapse Notch activation in solitary cells cultured on a substrate with Ecad-Fc. Left column: representative movies of solitary cells cultured on a substrate coated with Ecad-Fc but not with Dll4-Fc (n = 9). Negligible increase of mCherry fluorescence signal was observed. Right column: representative movies of solitary cells cultured on a substrate coated with Ecad-Fc and Dll4-Fc (n = 9). Gradual increase of bright mCherry fluorescence signal was observed. SNAP-NFL-Gal4 expression and mCherry fluorescence are pseudo-coloured as green and red, respectively. Time stamps are relative to doxycycline addition.
Supplementary Table 1
Supplementary Table 1. Cell lines and genetic constructs used in this study. Supplementary Table 2. Primers and oligonucleotides
Source data
Source Data Fig. 1
Statistical source data for Fig. 1.
Source Data Fig. 2
Statistical source data for Fig. 2.
Source Data Fig. 3
Statistical source data for Fig. 3.
Source Data Fig. 3
Unprocessed western blots for Fig. 3.
Source Data Fig. 4
Statistical source data for Fig. 4.
Source Data Fig. 4
Unprocessed western blots for Fig. 4.
Source Data Fig. 5
Statistical source data for Fig. 5.
Source Data Fig. 6
Statistical source data for Fig. 6.
Source Data Fig. 7
Statistical source data for Fig. 7.
Source Data Extended Data Fig. 1
Statistical source data for Extended Data Fig. 1.
Source Data Extended Data Fig. 2
Statistical source data for Extended Data Fig. 2.
Source Data Extended Data Fig. 3
Statistical source data for Extended Data Fig. 3.
Source Data Extended Data Fig. 4
Statistical source data for Extended Data Fig. 4.
Source Data Extended Data Fig. 5
Statistical source data for Extended Data Fig. 5.
Source Data Extended Data Fig. 5
Unprocessed western blots for Extended Data Fig. 5.
Source Data Extended Data Fig. 6
Statistical source data for Extended Data Fig. 6.
Source Data Extended Data Fig. 6
Unprocessed western blots for Extended Data Fig. 6.
Source Data Extended Data Fig. 7
Statistical source data for Extended Data Fig. 7.
Source Data Extended Data Fig. 7
Unprocessed western blots for Extended Data Fig. 7.
Source Data Extended Data Fig. 8
Statistical source data for Extended Data Fig. 8.
Source Data Extended Data Fig. 9
Statistical source data for Extended Data Fig. 9.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kwak, M., Southard, K.M., Kim, W.R. et al. Adherens junctions organize size-selective proteolytic hotspots critical for Notch signalling. Nat Cell Biol 24, 1739–1753 (2022). https://doi.org/10.1038/s41556-022-01031-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-022-01031-6
This article is cited by
-
Adherens junctions as molecular regulators of emergent tissue mechanics
Nature Reviews Molecular Cell Biology (2024)
-
A magnetically powered nanomachine with a DNA clutch
Nature Nanotechnology (2024)
-
The blood–brain barrier: Structure, regulation and drug delivery
Signal Transduction and Targeted Therapy (2023)