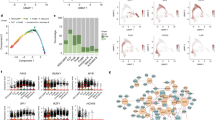
Abstract
Tumour heterogeneity encompasses both the malignant cells and their microenvironment. While heterogeneity between individual patients is known to affect the efficacy of cancer therapy, most personalized treatment approaches do not account for intratumour heterogeneity. We addressed this issue by studying the heterogeneity of nodal B-cell lymphomas by single-cell RNA-sequencing and transcriptome-informed flow cytometry. We identified transcriptionally distinct malignant subpopulations and compared their drug-response and genomic profiles. Malignant subpopulations from the same patient responded strikingly differently to anti-cancer drugs ex vivo, which recapitulated subpopulation-specific drug sensitivity during in vivo treatment. Infiltrating T cells represented the majority of non-malignant cells, whose gene-expression signatures were similar across all donors, whereas the frequencies of T-cell subsets varied significantly between the donors. Our data provide insights into the heterogeneity of nodal B-cell lymphomas and highlight the relevance of intratumour heterogeneity for personalized cancer therapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The WGS and WES data have been deposited at the European Genome-phenome Archive (EGA). The EGA study accession ID is EGAS00001004335. The scRNA-seq data that support the findings of this study have been deposited in heiDATA under accession code VRJUNV. The single-cell expression data of merged B- and T-cell UMAP plots (Fig. 2a,b and Fig. 3a,b) are available for easy-to-use interactive browsing at https://www.zmbh.uni-heidelberg.de/Anders/scLN-index.html. Genes that were differentially expressed between B-cell clusters can be browsed in an interactive html file (Supplementary Data 1). All other data supporting the findings of this study are available from the corresponding author on reasonable request.
Code availability
The R codes used for data analysis are available at our GitHub repository without further restriction (www.github.com/DietrichLab/scLymphomaExplorer).
References
Burrell, R. A., McGranahan, N., Bartek, J. & Swanton, C. The causes and consequences of genetic heterogeneity in cancer evolution. Nature 501, 338–345 (2013).
Chapuy, B. et al. Molecular subtypes of diffuse large B cell lymphoma are associated with distinct pathogenic mechanisms and outcomes. Nat. Med. 24, 679–690 (2018).
Swanton, C. Intratumor heterogeneity: evolution through space and time. Cancer Res. 72, 4875–4882 (2012).
McGranahan, N. & Swanton, C. Clonal heterogeneity and tumor evolution: past, present, and the future. Cell 168, 613–628 (2017).
Kim, I. S. & Zhang, X. H. One microenvironment does not fit all: heterogeneity beyond cancer cells. Cancer Metastasis Rev. 35, 601–629 (2016).
Fridman, W. H., Pages, F., Sautes-Fridman, C. & Galon, J. The immune contexture in human tumours: impact on clinical outcome. Nat. Rev. Cancer 12, 298–306 (2012).
Navin, N. et al. Tumour evolution inferred by single-cell sequencing. Nature 472, 90–94 (2011).
Wang, Y. & Navin, N. E. Advances and applications of single-cell sequencing technologies. Mol. Cell 58, 598–609 (2015).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Gao, R. et al. Nanogrid single-nucleus RNA sequencing reveals phenotypic diversity in breast cancer. Nat. Commun. 8, 228 (2017).
Yuan, J. & Sims, P. A. An automated microwell platform for large-scale single cell RNA-seq. Sci. Rep. 6, 33883 (2016).
Islam, S. et al. Quantitative single-cell RNA-seq with unique molecular identifiers. Nat. Methods 11, 163–166 (2014).
Milpied, P. et al. Human germinal center transcriptional programs are de-synchronized in B cell lymphoma. Nat. Immunol. 19, 1013–1024 (2018).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Ramskold, D. et al. Full-length mRNA-seq from single-cell levels of RNA and individual circulating tumor cells. Nat. Biotechnol. 30, 777–782 (2012).
Powell, A. A. et al. Single cell profiling of circulating tumor cells: transcriptional heterogeneity and diversity from breast cancer cell lines. PLoS ONE 7, e33788 (2012).
Kim, C. et al. Chemoresistance evolution in triple-negative breast cancer delineated by single-cell sequencing. Cell 173, 879–893 (2018).
Teras L. R. et al. 2016 US lymphoid malignancy statistics by World Health Organization subtypes. CA Cancer J. Clin. 66, 443–459 (2016).
Wagner-Johnston, N. D. et al. Outcomes of transformed follicular lymphoma in the modern era: a report from the National LymphoCare Study (NLCS). Blood 126, 851–857 (2015).
Crump, M. et al. Outcomes in refractory diffuse large B-cell lymphoma: results from the international SCHOLAR-1 study. Blood 130, 1800–1808 (2017).
Philip, T. et al. High-dose therapy and autologous bone marrow transplantation after failure of conventional chemotherapy in adults with intermediate-grade or high-grade non-Hodgkin’s lymphoma. N. Engl. J. Med. 316, 1493–1498 (1987).
Bartlett, N. L. et al. Single-agent ibrutinib in relapsed or refractory follicular lymphoma: a phase 2 consortium trial. Blood 131, 182–190 (2018).
Winter A. M., et al. A multi-institutional outcomes analysis of patients with relapsed or refractory DLBCL treated with ibrutinib. Blood 130, 1676–1679 (2017).
Horna, P., Olteanu, H., Kroft, S. H. & Harrington, A. M. Flow cytometric analysis of surface light chain expression patterns in B-cell lymphomas using monoclonal and polyclonal antibodies. Am. J. Clin. Pathol. 136, 954–959 (2011).
Challa-Malladi, M. et al. Combined genetic inactivation of β2-Microglobulin and CD58 reveals frequent escape from immune recognition in diffuse large B cell lymphoma. Cancer Cell 20, 728–740 (2011).
Schreiber, R. D., Old, L. J. & Smyth, M. J. Cancer immunoediting: integrating immunity’s roles in cancer suppression and promotion. Science 331, 1565–1570 (2011).
Xu-Monette, Z. Y., Zhou, J. & Young, K. H. PD-1 expression and clinical PD-1 blockade in B-cell lymphomas. Blood 131, 68–83 (2018).
Sun, L. L. et al. Anti-CD20/CD3 T cell-dependent bispecific antibody for the treatment of B cell malignancies. Sci. Transl. Med. 7, 287ra270 (2015).
Os, A. et al. Chronic lymphocytic leukemia cells are activated and proliferate in response to specific T helper cells. Cell Rep. 4, 566–577 (2013).
Becht E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2019).
Schaerli, P. et al. CXC chemokine receptor 5 expression defines follicular homing T cells with B cell helper function. J. Exp. Med. 192, 1553–1562 (2000).
Breitfeld, D. et al. Follicular B helper T cells express CXC chemokine receptor 5, localize to B cell follicles, and support immunoglobulin production. J. Exp. Med. 192, 1545–1552 (2000).
Dorfman, D. M. & Shahsafaei, A. CD200 (OX-2 membrane glycoprotein) is expressed by follicular T helper cells and in angioimmunoblastic T-cell lymphoma. Am. J. Surg. Pathol. 35, 76–83 (2011).
Weber, J. P. et al. ICOS maintains the T follicular helper cell phenotype by down-regulating Kruppel-like factor 2. J. Exp. Med. 212, 217–233 (2015).
Yang, Z. Z. et al. PD-1 expression defines two distinct T-cell sub-populations in follicular lymphoma that differentially impact patient survival. Blood Cancer J. 5, e281 (2015).
Byford, E. T., Carr, M., Ladikou, E., Ahearne, M. J. & Wagner, S. D. Circulating Tfh1 (cTfh1) cell numbers and PD1 expression are elevated in low-grade B-cell non-Hodgkin’s lymphoma and cTfh gene expression is perturbed in marginal zone lymphoma. PLoS ONE 13, e0190468 (2018).
Yamazaki, T., Nagumo, H., Hayashi, T., Sugane, K. & Agematsu, K. CD72-mediated suppression of human naive B cell differentiation by down-regulating X-box binding protein 1. Eur. J. Immunol. 35, 2325–2334 (2005).
Klein, U., Rajewsky, K. & Kuppers, R. Human immunoglobulin (Ig)M+IgD+ peripheral blood B cells expressing the CD27 cell surface antigen carry somatically mutated variable region genes: CD27 as a general marker for somatically mutated (memory) B cells. J. Exp. Med. 188, 1679–1689 (1998).
Hans, C. P. et al. Confirmation of the molecular classification of diffuse large B-cell lymphoma by immunohistochemistry using a tissue microarray. Blood 103, 275–282 (2004).
Vento-Tormo, R. et al. Single-cell reconstruction of the early maternal–fetal interface in humans. Nature 563, 347–353 (2018).
Shirota, H. et al. IL4 from T follicular helper cells downregulates antitumor immunity. Cancer Immunol. Res. 5, 61–71 (2017).
Aguilar-Hernandez, M. M. et al. IL-4 enhances expression and function of surface IgM in CLL cells. Blood 127, 3015–3025 (2016).
Peter-Martin, B. et al. Systematic investigation of microenvironmental drug resistance mechanisms in chronic lymphocytic leukemia. Blood 134, 3363 (2019).
Spolski, R. & Leonard, W. J. IL-21 and T follicular helper cells. Int. Immunol. 22, 7–12 (2010).
Gu-Trantien C. et al. CXCL13-producing TFH cells link immune suppression and adaptive memory in human breast cancer. JCI Insight 2, e91487 (2017).
DiToro D. et al. Differential IL-2 expression defines developmental fates of follicular versus nonfollicular helper T cells. Science 361, eaao2933 (2018).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Dietrich, S. et al. Drug-perturbation-based stratification of blood cancer. J. Clin. Invest. 128, 427–445 (2018).
Delmore, J. E. et al. BET bromodomain inhibition as a therapeutic strategy to target c-Myc. Cell 146, 904–917 (2011).
Chapuy, B. et al. Discovery and characterization of super-enhancer-associated dependencies in diffuse large B cell lymphoma. Cancer Cell 24, 777–790 (2013).
Andor, N. et al. Single-cell RNA-seq of follicular lymphoma reveals malignant B-cell types and coexpression of T-cell immune checkpoints. Blood 133, 1119–1129 (2019).
Ledergor, G. et al. Single cell dissection of plasma cell heterogeneity in symptomatic and asymptomatic myeloma. Nat. Med. 24, 1867–1876 (2018).
Puram, S. V. et al. Single-cell transcriptomic analysis of primary and metastatic tumor ecosystems in head and neck. Cancer Cell 171, 1611–1624 (2017).
de Boer, B. et al. Prospective isolation and characterization of genetically and functionally distinct AML subclones. Cancer Cell 34, 674–689 (2018).
Li, J. et al. Tumor cell-intrinsic factors underlie heterogeneity of immune cell infiltration and response to immunotherapy. Immunity 49, 178–193 (2018).
Bankhead, P. et al. QuPath: open source software for digital pathology image analysis. Sci. Rep. 7, 16878 (2017).
Ratech, H. & Litwin, S. Surface immunoglobulin light chain restriction in B-cell non-Hodgkin’s malignant lymphomas. Am. J. Clin. Pathol. 91, 583–586 (1989).
Kaleem, Z., Zehnbauer, B. A., White, G. & Zutter, M. M. Lack of expression of surface immunoglobulin light chains in B-cell non-Hodgkin lymphomas. Am. J. Clin. Pathol. 113, 399–405 (2000).
Heining, C. et al. NRG1 fusions in KRAS wild-type pancreatic cancer. Cancer Discov. 8, 1087–1095 (2018).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Haghverdi, L., Lun, A. T. L., Morgan, M. D. & Marioni, J. C. Batch effects in single-cell RNA-sequencing data are corrected by matching mutual nearest neighbors. Nat. Biotechnol. 36, 421–427 (2018).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Mootha, V. K. et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
López, C. et al. Genomic and transcriptomic changes complement each other in the pathogenesis of sporadic Burkitt lymphoma. Nat. Commun. 10, 1459 (2019).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Kleinheinz K. et al. ACEseq—allele specific copy number estimation from whole genome sequencing. Preprint at bioRxiv https://doi.org/10.1101/210807 (2017).
Talevich, E., Shain, A. H., Botton, T. & Bastian, B. C. CNVkit: genome-wide copy number detection and visualization from targeted DNA sequencing. PLoS Comput. Biol. 12, e1004873 (2016).
Acknowledgements
T.R. was supported by a physician scientist fellowship from the Medical Faculty of University Heidelberg. M.Seiffert was supported by a grant from the Deutsche Forschungsgemeinschaft (DFG). S.D. was supported by a grant from the Hairy Cell Leukemia Foundation, Heidelberg Research Centre for Molecular Medicine (HRCMM) and an e:med BMBF junior group grant and DFG through the SFB873 (project B7). The Anders laboratory was supported by the DFG via Collaborative Research Centre (CRC) 1366 (project number 394046768). For the data management, we thank the Scientific Data Storage Heidelberg (SDS@hd), which is funded by the state of Baden-Württemberg and a DFG grant (grant no. INST 35/1314-1 FUGG). We thank C. Kolb (University Hospital Heidelberg), M. Knoll (University Hospital Heidelberg), A. Lenze (University Hospital Heidelberg), the EMBL flow core facility and the EMBL gene core facility for their excellent technical assistance. We also thank the DKFZ Single-Cell Open Lab (scOpenLab) for the experimental assistance and J. Apel (heiDATA, University of Heidelberg) for assistance with data management. This study was also supported by the Heidelberg Centre for Personalized Oncology (DKFZ-HIPO). We thank the DKFZ Omics IT and Data Management Core Facility (ODCF) and the DKFZ Genomics and Proteomics Core Facility (GPCF) for their technical support.
Author information
Authors and Affiliations
Contributions
T.R., M.B., M. Stolarczyk, J.-P.M., SA.H. and P.-M.B. performed experiments. T.R., J.S., A.U., F.F., M.B., N.A., H.B.-W., M.P., M. Schlesner and S.A. analysed the data. T.R., M.H., K.R., B.G., M. Seiffert, B.B., G.M., C.M.-T., S.F., W.H., S.A. and S.D. interpreted the data. T.R., T.Z., S.F. and S.D. designed the study. T.R., J.S., K.R., B.C., M. Schlesner, W.H., S.A. and S.D. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Classification of B and T cells by scRNA-seq.
a) Representative lymph node sample demonstrating how lymph node-derived B and T cells were classified by scRNA-seq (CD79B vs. CD3), flow cytometry (CD19 vs. CD3), or immunohistochemistry (PAX5 vs. CD3). Frequencies of B and T cells were determined for each approach in 12 (scRNA-seq, flow cytometry) or 7 (immunohistochemistry) biologically independent samples and correlated pairwise with each other (see main Fig. 1b and Fig. 1c). b) ScRNA-seq data of 12 biologically independent lymph node samples were subjected to SNN-based clustering and visualized by t-SNE. Each t-SNE represents one individual sample as indicated. Different B cell or T cell clusters are illustrated by different shades of green or blue, respectively. SNN: Shared-nearest-neighbour. See Source Data Extended Data Fig. 1.
Extended Data Fig. 2 Classification of malignant versus healthy B cells by means of kappa and lambda light chain expression at the scRNA level.
scRNA-seq data of 12 independent lymph node samples were subjected to SNN-based clustering and visualized by t-SNE. Each t-SNE represents one individual sample as indicated. Cells are coloured by light chain kappa fraction IGKC/(IGKC + IGLC2) of each single cell to demonstrate light chain restriction of each cluster. Non-B cells are coloured in grey. The sample DLBCL1 showed only marginal light chain expression on single cell RNA level. Therefore, these cells were regarded as malignant cells (see Method section for details). The same is true for the larger cluster of sample FL4. SNN: Shared-nearest-neighbour. See Source Data Extended Data Fig. 2.
Extended Data Fig. 3 Frequencies of T cells, healthy B cells and malignant B cells perfectly correlate between flow cytometry and scRNA-seq.
a) Stacked bar graph of the relative frequencies of malignant B cells (B), T cells (T), healthy B cells (hB) and myeloid cells (Other) calculated on the basis of scRNA expression profiles. Shown are N = 12 biologically independent samples. b) Lymph node derived cells from those samples passed to scRNA-seq (A) were stained for viability, CD19, CD3, kappa light chain and lambda light chain. The proportion of healthy (hB) and malignant B cells (B) were estimated based on the ratio of light chain restricted CD19+ tumour cells and CD19+ non-tumour cells (malignant B cells = light chain restricted B cells – non-restricted B cells). T cells (T) refer to CD3+CD19− cells, whereas Other refers to CD19−CD3− cells. Pearson’s correlation coefficients (r) are given. c) Lymph node derived cells were stained and analysed as described in B. Shown are the frequencies for each subpopulation for n = 9 (rLN), N = 4 (MCL), N = 11 (FL), N = 9 (DLBCL) and N = 7 (CLL) biologically independent samples. Box plots show the minimum, first quartile, median, second quartile and maximum. Outliers defined as values higher or lower than 1.5 interquartile ranges from median are shown as individual dots. rLN: Reactive lymph node. MCL: Mantle cell lymphoma. FL: Follicular lymphoma. DLBCL: Diffuse large B cell lymphoma, CLL: Chronic lymphocytic leukaemia. See Source Data Extended Data Fig. 3.
Extended Data Fig. 4 Proliferative capacity is preserved at the scRNA level and correlates with immunohistochemical and flow cytometry-based staining of Ki67.
a) Dot plots show G2M and S score for the B cells of four representative samples. Cells with positive G2M or S score were marked as proliferating (please see method section for details). b) The proportion of proliferating cells based on scRNA-seq (A) was correlated with the percentage of Ki67+ cells determined either by flow cytometry (orange) or immunohistochemistry (green). R values represent Pearson correlation coefficients. N = 12 biologically independent samples are shown. See Source Data Extended Data Fig. 4.
Extended Data Fig. 5 intratumour heterogeneity of malignant B cells.
a–i) ScRNA expression profiles of malignant and non-malignant B cells only were subjected to SNN-based clustering and visualized by t-SNE. Shown are 9 biologically independent samples, whereby each t-SNE represents one individual sample as indicated. Cells were coloured by cluster. SNN: Shared-nearest-neighbour. See Source Data Extended Data Fig. 5.
Extended Data Fig. 6 Dissecting four distinct subpopulations of malignant B cells in FL4 sample by means of scRNA-seq-informed flow cytometry.
a) ScRNA expression profiles of B cells from the FL4 sample only (2367 cells) were subjected to SNN-based clustering and visualized by t-SNE. b) Heatmap shows a selection of differentially expressed surface markers used for cluster differentiation. Gene expression values were scaled to the maximum of each row. c) T-SNE plot of panel A coloured by the light chain kappa fraction IGKC/(IGKC + IGLC2) of each single cell for FL4 sample only. C1 contains cells either expressing IGKC or IGLC2 predominantly (benign B cells), C2 contains only cells expressing predominantly IGLC2, whereas C3 to C5 hardly express both IGKC and IGLC2. d–f) Cells derived from sample FL4 were stained for viability, CD3, CD19, kappa, lambda, CD44, CD24, CD22 and CD27. Shown is the stepwise gating strategy to comprehend the scRNA-seq-based clusters of panel A. g) Lymph node derived cells from the FL4 sample were incubated for 48 hours with 58 different drugs and 5 concentrations. Cells were stained for viability, CD3, CD19, kappa, lambda and CD27. Viability was normalized to DMSO controls for each subpopulation separately. Shows are only those drugs with significantly differential drug response between the two subpopulations. SNN: Shared-nearest-neighbour. DMSO: Dimethyl sulfoxide. See Source Data Extended Data Fig. 6.
Extended Data Fig. 7 Intratumour heterogeneity of tFL1 characterized by differential cell cycle states, gene expression signatures and copy number alterations.
a, b) ScRNA expression profiles of B cells from the tFL1 sample only (492 cells) were subjected to t-SNE and coloured by S-Score (a) or G2M-Score (b, see Methods section for details). c–e) Gene set enrichment analysis was performed between CD32Low and CD32High cluster. Shown are enrichment plots for hallmark MYC targets (c), MTORC1 signalling (d) and hallmark G2M targets (e). Given are the false-positive detection rate (FDR) and the normalized enrichment score (NES). FDR and NES were calculated using GSEA desktop application (see Method section for details). f) T-SNE as shown in panel A/B was coloured by FCGR2B expression. g) Line plot shows cluster-specific total copy number estimation for all chromosomes inferred from whole exome sequencing. See Source Data Extended Data Fig. 7.
Extended Data Fig. 8 Intratumour heterogeneity of DLBCL1 sample.
a–c) ScRNA profiles of malignant B cells from the DLBCL1 sample only (3114 cells) were subjected to t-SNE and coloured by CD48 (a), SELL (b) and MYC (c) expression. d) DLBCL1 derived lymph node cells were stained for viability, CD19, CD3, CD48, CD62L and MYC or respective isotype control. Histograms show fluorescence intensity of MYC for isotype control, T cells, CD48HighCD62L+ and CD48LowCD62L− subpopulations. Experiment was repeated three times with similar results. E-G) DLBCL1 derived lymph node cells were incubated with DMSO, I-BET-762 (e) at two concentrations (1 µM, 5 µM), OTX015 (f) at two concentrations (1 µM, 5 µM), or ibrutinib (g) at two concentrations (0.2 µM, 1 µM). After 24 hours, cells were harvested and stained as described in panel d. Histograms show fluorescence intensity for CD48HighCD62L+ subclone. Experiment was repeated three times with similar results. H) Line plot shows total cluster-specific copy number estimation of DLBCL1 sample for all chromosomes as indicated using whole genome sequencing. DMSO: Dimethyl sulfoxide. See Source Data Extended Data Fig. 8.
Supplementary information
Supplementary Information
Supplementary Figs. 1–3.
Supplementary Tables
Supplementary Tables 1–8.
Supplementary Data 1
Interactive html file to browse differentially expressed genes between B-cell clusters in scRNA-seq data.
Source data
Source Data Fig. 1
Statistical source data
Source Data Fig. 2
Statistical source data
Source Data Fig. 3
Statistical source data
Source Data Fig. 4
Statistical source data
Source Data Fig. 5
Statistical source data
Source Data Fig. 6
Statistical source data
Source Data Extended Data Fig. 1
Statistical source data
Source Data Extended Data Fig. 2
Statistical source data
Source Data Extended Data Fig. 3
Statistical source data
Source Data Extended Data Fig. 4
Statistical source data
Source Data Extended Data Fig. 5
Statistical source data
Source Data Extended Data Fig. 6
Statistical source data
Source Data Extended Data Fig. 7
Statistical source data
Source Data Extended Data Fig. 8
Statistical source data
Rights and permissions
About this article
Cite this article
Roider, T., Seufert, J., Uvarovskii, A. et al. Dissecting intratumour heterogeneity of nodal B-cell lymphomas at the transcriptional, genetic and drug-response levels. Nat Cell Biol 22, 896–906 (2020). https://doi.org/10.1038/s41556-020-0532-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-0532-x
This article is cited by
-
Simple but powerful interactive data analysis in R with R/LinekdCharts
Genome Biology (2024)
-
Diffuse large B-cell lymphoma: the significance of CD8+ tumor-infiltrating lymphocytes exhaustion mediated by TIM3/Galectin-9 pathway
Journal of Translational Medicine (2024)
-
Multimodal and spatially resolved profiling identifies distinct patterns of T cell infiltration in nodal B cell lymphoma entities
Nature Cell Biology (2024)
-
Advances in single-cell RNA sequencing and its applications in cancer research
Journal of Hematology & Oncology (2023)
-
Quantitative PET-based biomarkers in lymphoma: getting ready for primetime
Nature Reviews Clinical Oncology (2023)