Abstract
Tissue stem cells are the cell of origin for many malignancies. Metabolites regulate the balance between self-renewal and differentiation, but whether endogenous metabolic pathways or nutrient availability predispose stem cells towards transformation remains unknown. Here, we address this question in epidermal stem cells (EpdSCs), which are a cell of origin for squamous cell carcinoma. We find that oncogenic EpdSCs are serine auxotrophs whose growth and self-renewal require abundant exogenous serine. When extracellular serine is limited, EpdSCs activate de novo serine synthesis, which in turn stimulates α-ketoglutarate-dependent dioxygenases that remove the repressive histone modification H3K27me3 and activate differentiation programmes. Accordingly, serine starvation or enforced α-ketoglutarate production antagonizes squamous cell carcinoma growth. Conversely, blocking serine synthesis or repressing α-ketoglutarate-driven demethylation facilitates malignant progression. Together, these findings reveal that extracellular serine is a critical determinant of EpdSC fate and provide insight into how nutrient availability is integrated with stem cell fate decisions during tumour initiation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Change history
12 October 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Ge, Y. & Fuchs, E. Stretching the limits: from homeostasis to stem cell plasticity in wound healing and cancer. Nat. Rev. Genet. 19, 311–325 (2018).
Martincorena, I. et al. High burden and pervasive positive selection of somatic mutations in normal human skin. Science 348, 880–886 (2015).
Martincorena, I. et al. Somatic mutant clones colonize the human esophagus with age. Science 362, 911–917 (2018).
Blokzijl, F. et al. Tissue-specific mutation accumulation in human adult stem cells during life. Nature 538, 260–264 (2016).
Jaiswal, S. et al. Age-related clonal hematopoiesis associated with adverse outcomes. N. Engl. J. Med. 371, 2488–2498 (2014).
Genovese, G. et al. Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N. Engl. J. Med. 371, 2477–2487 (2014).
Lapouge, G. et al. Identifying the cellular origin of squamous skin tumors. Proc. Natl Acad. Sci. USA 108, 7431–7436 (2011).
Barker, N. et al. Crypt stem cells as the cells-of-origin of intestinal cancer. Nature 457, 608–611 (2009).
White, A. C. et al. Defining the origins of Ras/p53-mediated squamous cell carcinoma. Proc. Natl Acad. Sci. USA 108, 7425–7430 (2011).
White, A. C. & Lowry, W. E. Refining the role for adult stem cells as cancer cells of origin. Trends Cell Biol. 25, 11–20 (2015).
Brown, S. et al. Correction of aberrant growth preserves tissue homeostasis. Nature 548, 334–337 (2017).
Dow, L. E. et al. Apc restoration promotes cellular differentiation and reestablishes crypt homeostasis in colorectal. Cancer Cell 161, 1539–1552 (2015).
Ying, Z., Sandoval, M. & Beronja, S. Oncogenic activation of PI3K induces progenitor cell differentiation to suppress epidermal growth. Nat. Cell Biol. 20, 1256–1266 (2018).
Warrell, R. P. et al. Differentiation therapy of acute promyelocytic leukemia with tretinoin (all-trans-retinoic acid). N. Engl. J. Med. 324, 1385–1393 (1991).
Boumahdi, S. et al. SOX2 controls tumour initiation and cancer stem-cell functions in squamous-cell carcinoma. Nature 511, 246–250 (2014).
Sendoel, A. et al. Translation from unconventional 5′ start sites drives tumour initiation. Nature 541, 494–499 (2017).
Maier, S. et al. SOX2 amplification is a common event in squamous cell carcinomas of different organ sites. Hum. Pathol. 42, 1078–1088 (2011).
Blanco, S. et al. Stem cell function and stress response are controlled by protein synthesis. Nature 534, 335–340 (2016).
Saxton, R. A. & Sabatini, D. M. mTOR signaling in growth, metabolism, and disease. Cell 168, 960–976 (2017).
Carey, B. W., Finley, L. W. S., Cross, J. R., Allis, C. D. & Thompson, C. B. Intracellular α-ketoglutarate maintains the pluripotency of embryonic stem cells. Nature 518, 413–416 (2014).
Flores, A. et al. Lactate dehydrogenase activity drives hair follicle stem cell activation. Nat. Cell Biol. 19, 1017–1026 (2017).
Agathocleous, M. et al. Ascorbate regulates haematopoietic stem cell function and leukaemogenesis. Nature 549, 476–481 (2017).
Intlekofer, A. M. & Finley, L. W. S. Metabolic signatures of cancer cells and stem cells. Nat. Metab. 1, 177–188 (2019).
Vardhana, S. A. et al. Glutamine independence is a selectable feature of pluripotent stem cells. Nat. Metab. 1, 676–687 (2019).
Maddocks, O. D. K. et al. Modulating the therapeutic response of tumours to dietary serine and glycine starvation. Nature 544, 372–376 (2017).
Maddocks, O. D. K. et al. Serine starvation induces stress and p53-dependent metabolic remodelling in cancer cells. Nature 493, 542–546 (2012).
Gao, X. et al. Dietary methionine influences therapy in mouse cancer models and alters human metabolism. Nature 572, 397–401 (2019).
Liu, K. et al. Sox2 cooperates with inflammation-mediated Stat3 activation in the malignant transformation of foregut basal progenitor cells. Cell Stem Cell 12, 304–315 (2013).
Hosios, A. M. et al. Amino acids rather than glucose account for the majority of cell mass in proliferating mammalian cells. Dev. Cell 36, 540–549 (2016).
Yang, M. & Vousden, K. H. Serine and one-carbon metabolism in cancer. Nat. Rev. Cancer 16, 650–662 (2016).
Michelakis, E. D., Webster, L. & Mackey, J. R. Dichloroacetate (DCA) as a potential metabolic-targeting therapy for cancer. Br. J. Cancer 99, 989–994 (2008).
Halestrap, A. P. The mitochondrial pyruvate carrier. Kinetics and specificity for substrates and inhibitors. Biochem. J. 148, 85–96 (1975).
Diehl, F. F., Lewis, C. A., Fiske, B. P. & Vander Heiden, M. G. Cellular redox state constrains serine synthesis and nucleotide production to impact cell proliferation. Nat. Metab. 1, 861–867 (2019).
Sullivan, L. B. et al. Supporting aspartate biosynthesis is an essential function of respiration in proliferating cells. Cell 162, 552–563 (2015).
Titov, D. V. et al. Complementation of mitochondrial electron transport chain by manipulation of the NAD+/NADH ratio. Science 352, 231–235 (2016).
Williamson, D. H., Lund, P. & Krebs, H. A. The redox state of free nicotinamide-adenine dinucleotide in the cytoplasm and mitochondria of rat liver. Biochem. J. 103, 514–527 (1967).
Birsoy, K. et al. An essential role of the mitochondrial electron transport chain in cell proliferation is to enable aspartate synthesis. Cell 162, 540–551 (2015).
Lechler, T. & Fuchs, E. Asymmetric cell divisions promote stratification and differentiation of mammalian skin. Nature 437, 275–280 (2005).
Asare, A., Levorse, J. & Fuchs, E. Coupling organelle inheritance with mitosis to balance growth and differentiation. Science 355, eaah4701 (2017).
Altman, B. J., Stine, Z. E. & Dang, C. V. From Krebs to clinic: glutamine metabolism to cancer therapy. Nat. Rev. Cancer 16, 619–634 (2016).
Possemato, R. et al. Functional genomics reveal that the serine synthesis pathway is essential in breast cancer. Nature 476, 346–350 (2011).
Schvartzman, J. M., Thompson, C. B. & Finley, L. W. S. Metabolic regulation of chromatin modifications and gene expression. J. Cell Biol. 217, 2247–2259 (2018).
Ezhkova, E. et al. Ezh2 orchestrates gene expression for the stepwise differentiation of tissue-specific stem cells. Cell 136, 1122–1135 (2009).
Sen, G. L., Webster, D. E., Barragan, D. I., Chang, H. Y. & Khavari, P. A. Control of differentiation in a self-renewing mammalian tissue by the histone demethylase JMJD3. Genes Dev. 22, 1865–1870 (2008).
Maddocks, O. D. K., Labuschagne, C. F., Adams, P. D. & Vousden, K. H. Serine metabolism supports the methionine cycle and DNA/RNA methylation through de novo ATP synthesis in cancer cells. Mol. Cell 61, 210–221 (2016).
Hamanaka, R. B. et al. Mitochondrial reactive oxygen species promote epidermal differentiation and hair follicle development. Sci. Signal. 6, ra8 (2013).
Morris, J. P. et al. α-ketoglutarate links p53 to cell fate during tumour suppression. Nature 573, 595–599 (2019).
Castilho, R. M., Squarize, C. H., Chodosh, L. A., Williams, B. O. & Gutkind, J. S. mTOR mediates Wnt-induced epidermal stem cell exhaustion and aging. Cell Stem Cell 5, 279–289 (2009).
Ding, X. et al. mTORC1 and mTORC2 regulate skin morphogenesis and epidermal barrier formation. Nat. Commun. 7, 13226 (2016).
Reeves, M. Q., Kandyba, E., Harris, S., Del Rosario, R. & Balmain, A. Multicolour lineage tracing reveals clonal dynamics of squamous carcinoma evolution from initiation to metastasis. Nat. Cell Biol. 20, 699–709 (2018).
Sullivan, M. R. et al. Increased serine synthesis provides an advantage for tumors arising in tissues where serine levels are limiting. Cell Metab. 29, 1410–1421.e4 (2019).
Kruidenier, L. et al. A selective jumonji H3K27 demethylase inhibitor modulates the proinflammatory macrophage response. Nature 488, 404–408 (2012).
Min, B. M. et al. Inactivation of the p53 gene by either mutation or HPV infection is extremely frequent in human oral squamous cell carcinoma cell lines. Eur. J. Cancer. B Oral Oncol. 30, 338–345 (1994).
Reiss, M. et al. Status of the p53 tumor suppressor gene in human squamous carcinoma cell lines. Oncol. Res. 4, 349–357 (1992).
Ge, Y. et al. Stem cell lineage infidelity drives wound repair and cancer. Cell 169, 636–650.e14 (2017).
Locasale, J. W. et al. Phosphoglycerate dehydrogenase diverts glycolytic flux and contributes to oncogenesis. Nat. Genet. 43, 869–874 (2011).
Labuschagne, C. F., van den Broek, N. J. F., Mackay, G. M., Vousden, K. H. & Maddocks, O. D. K. Serine, but not glycine, supports one-carbon metabolism and proliferation of cancer cells. Cell Rep. 7, 1248–1258 (2014).
Lu, C. et al. IDH mutation impairs histone demethylation and results in a block to cell differentiation. Nature 483, 474–478 (2012).
Figueroa, M. E. et al. Leukemic IDH1 and IDH2 mutations result in a hypermethylation phenotype, disrupt Tet2 function, and impair hematopoietic differentiation. Cancer Cell 18, 553–567 (2010).
Letouzé, E. et al. SDH mutations establish a hypermethylator phenotype in paraganglioma. Cancer Cell 23, 739–752 (2013).
Xiao, M. et al. Inhibition of α-KG-dependent histone and DNA demethylases by fumarate and succinate that are accumulated in mutations of FH and SDH tumor suppressors. Genes Dev. 26, 1326–1338 (2012).
Intlekofer, A. M. et al. Hypoxia induces production of l-2-hydroxyglutarate. Cell Metab. 22, 304–311 (2015).
Pan, M. et al. Regional glutamine deficiency in tumours promotes dedifferentiation through inhibition of histone demethylation. Nat. Cell Biol. 18, 1090–1101 (2016).
Elia, I. et al. Breast cancer cells rely on environmental pyruvate to shape the metastatic niche. Nature 568, 117–121 (2019).
Raffel, S. et al. BCAT1 restricts αKG levels in AML stem cells leading to IDHmut-like DNA hypermethylation. Nature 551, 384–388 (2017).
Chang, C.-J. et al. EZH2 promotes expansion of breast tumor initiating cells through activation of RAF1-β–catenin signaling. Cancer Cell 19, 86–100 (2011).
Beck, B. et al. A vascular niche and a VEGF–Nrp1 loop regulate the initiation and stemness of skin tumours. Nature 478, 399–403 (2011).
Oshimori, N., Oristian, D. & Fuchs, E. TGF-β promotes heterogeneity and drug resistance in squamous cell carcinoma. Cell 160, 963–976 (2015).
Millard, P., Letisse, F., Sokol, S. & Portais, J.-C. IsoCor: correcting MS data in isotope labeling experiments. Bioinformatics 28, 1294–1296 (2012).
Perez, A. R. et al. GuideScan software for improved single and paired CRISPR guide RNA design. Nat. Biotechnol. 35, 347–349 (2017).
Acknowledgements
We thank J. Que for the R26-LSL-Sox2-IRES-eGFP mice and S. Lowe for sharing the LT3-GEPIR vector. We thank M. Nikolova and E. Wong for technical assistance, J. Levorse for in utero lentiviral injections, and M. Sribour, L. Polak and L. Hidalgo for mouse care and special diet experiments. We thank S. Mazel, S. Semova, S. Han and S. Tadesse at Rockefeller University’s Flow Cytometry core for conducting FACS sorting. We thank all members of the Fuchs and Finley labs for discussions. We thank A. Intlekofer for discussion and shared equipment, and S. Vardhana, S. Ellis, N. Infarinato and A. Siliciano for critical assessment of the manuscript. E.F. is a Howard Hughes Medical Investigator. L.W.S.F. is a Searle Scholar. S.C.B. is a Ruth Kirschstein NIH Predoctoral fellow (F31CA236465); S.G.-C. is a Postdoctoral Fellow of the Human Frontiers Science Program (LT001519/2017) and the European Molecular Biology Organization (ALTF 1239- 2016); B.H. is a Ruth Kirschstein NIH Predoctoral Fellow (F30CA236239); M.T.T. is a Ruth Kirstein NIH Postdoctoral Fellow (1F32AR073105); and B.H., J.S.S.N. and S.C.B. are predoctoral fellows of the Weill Cornell/Rockefeller/Sloan Kettering Tri-Institutional Medical Scientist Training Program (T32GM007739). This research was supported by grants to E.F. from the National Institutes of Health (R01-AR31737), NYSTEM (C32585GG) and a collaborative grant from The Starr Foundation (I11-0039 to E.F. and L.W.S.F.). This work was additionally supported by grants to L.W.S.F. from the Damon Runyon Cancer Research Foundation (DFS-23-17), the Concern Foundation, the Anna Fuller Fund, The Edward Mallinckrodt, Jr. Foundation, The Starr Foundation (I12-051) and the Memorial Sloan Kettering Cancer Center Support Grant P30 CA008748.
Author information
Authors and Affiliations
Contributions
S.C.B, E.F. and L.W.S.F. conceptualized the study, designed the experiments, interpreted the data and wrote the manuscript. S.C.B. and S.G.-C. performed the experiments and collected the data. P.K.T. performed and analysed the metabolic assays. B.H. generated Phgdh knockout lines and contributed to the allografting experiments. Y.G. generated the enhancer reporter SCC lines. J.S.S.N. contributed to the immunohistochemistry staining and analyses for the human tissue arrays. M.T.T. contributed to the generation of shPhgdh lines. J.d.C.-R. prepared serum and media for the metabolic profiling experiments. All authors provided input on the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
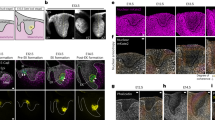
Extended Data Fig. 1 Pre-malignant EpdSCs are serine auxotrophs.
a. Representative immunofluorescence of progenitor markers α6 and K14 and tumor SC markers CD44 and SOX2 in K14-CreER;SOX2+ pre-tumorigenic lesions in second telogen mice two weeks after tamoxifen administration. Three mice per genotype were analyzed with similar results. Scale bars = 50 μm. b, Relative levels of amino acids from conditioned medium relative to unconditioned medium measured by GC-MS (n = 6 biologically independent samples). Data are mean ±SEM. c, Fractional labeling of intracellular serine from [U-13C]serine (n = 3 biologically independent samples). Data are mean ±SD. d, Population doublings of H-RasG12V-expressing pre-malignant keratinocytes following 48 h of Ser/Gly starvation (n = 3 biologically independent samples). Data are mean ±SD. e, Immunoblot of serine synthesis enzymes in WT and SOX2+ cells following 24 h of culture in control or Ser/Gly-free medium. See Supplementary Table 1 for quantification of immunoblot from triplicate independent experiments. f, Labeling of intracellular serine from [U-13C]glycine (left) and intracellular glycine from [U-13C]serine (right) (n = 3 biologically independent samples). Data are mean ±SD. Statistical significance was determined using a two-way ANOVA with Sidak’s multiple comparison test for panels b, c, and f, and an unpaired two-tailed student’s t-test for panel d. Scanned images of unprocessed blots are shown in Source Data Extended Fig. 1. Numerical data are provided in Statistics Source Data Extended Fig. 1.
Extended Data Fig. 2 NAD+ regeneration regulates serine auxotrophy.
a, Schematic of glucose metabolism via glycolysis and the TCA cycle, including associated inhibitors. b, Isotopologues of citrate formed from [U-13C]glucose (n = 3 biologically independent samples). Data are mean ±SD. M+0 represents the fraction of citrate not labeled by glucose-derived carbons. Heavier isotopologues are formed when glucose is used to generate citrate through either PDH (M+2) or PC (M+3). Higher weight isotopologues are derived from successive turns around the TCA cycle. c, RT-qPCR for pyruvate metabolism genes (n = 3 independent experiments). Data are mean ±SEM. d, e, Relative cell number with dichloroacetate (DCA) (n = 3 biologically independent samples) (d) or UK-5099 (n = 3 biologically independent samples) (e). Data are mean ±SD. f, Mechanism of action of α-ketobutyrate (AKB). g, Proliferation with AKB (n = 3 biologically independent samples). Data are mean ±SD. h, Relative cell number with indicated compounds (n = 3). Data are mean ±SEM. i, Schematic of pyruvate mechanism of action. j, Percentage of pyruvate consumed after 24 h (n = 5 for SOX2+ –Ser/Gly, otherwise n = 6 biologically independent samples). Data are mean ±SEM. k, 24 hr lactate secretion into medium (n = 6 biologically independent samples). Data are mean ±SEM. l, Representative immunofluorescence of transduced EpdSCs, performed in triplicate independent experiments with similar results. m, Quantification of the whole cell NAD+/NADH ratio (n = 3 biologically independent samples). Data are mean ±SEM. n, [U-13C]glucose labeling of serine following 16 h of Ser/Gly starvation (n = 3 biologically independent samples). Data are mean ±SEM. Scale bars = 10 μm. Statistical significance was determined using an unpaired two-tailed student’s t-test for panels m, an unpaired two-tailed student’s t-test using the Holm-Sidak method for multiple comparisons in panels d, e and n, a two-way ANOVA with Sidak’s multiple comparison test for panels b, c, g and k, and Dunnett’s multiple comparison test for panel h. Numerical data are provided in Statistics Source Data Extended Fig. 2.
Extended Data Fig. 3 Effects of serine starvation on epidermal growth and differentiation.
a, Serum serine and glycine in female mice maintained on indicated chow for 2 weeks (n = 6 mice per condition). Data are mean ±SD. b, Representative immunofluorescence of cleaved caspase 3 in P0 WT and SOX2+ mice. Analysis was performed on 3 mice/condition with similar results. c, Growth curve in indicated media (n = 3 biologically independent samples). d, e, Intracellular serine pools (d) and 4-hr fractional labeling from glucose (e) following 24 h of low Ser/Gly culture (n = 3 biologically independent samples). f, Schematic of in vivo LbNOX expression experiment. g, Proliferation in P0 mice on control or Ser/Gly-free diet (n = 6 WT control, 5 WT –Ser/Gly, 6 SOX2+ control, 10 SOX2+ –Ser/Gly mice). h, Immunofluorescence of cell division classes based on Survivin staining (left), quantification of spindle angle relative to the basement membrane in P0 WT and SOX2+ SCs on control or Ser/Gly-free diet (middle, data are mean), and binning of spindle axes in WT control (n = 21 mitoses), WT –Ser/Gly (n = 25 mitoses), SOX2+ control (n = 32 mitoses) and SOX2+ -Ser/Gly (n = 35 mitoses) (right). Mitoses were counted across three animals per condition. Scale bar = 5 μm. SB = suprabasal. i, j, Immunofluorescence (i) and quantification (j) of K14 in indicated media (n = 3 independent experiments). Scale bar = 50 μm k, Immunofluorescence of K14 in indicated media (n = 3 independent experiments). Scale bar = 50 μm. Unless indicated all data are mean ±SEM. Statistical analysis was performed by an unpaired two-tailed student’s t-test for panels a and d, a two-way ANOVA with Sidak’s multiple comparison test for panels e and g and Tukey’s multiple comparison test in panel k, a parts of whole Chi-Square analysis in panel h, and a one-way ANOVA with Tukey’s multiple comparison test in panel j. Numerical data are provided in Statistics Source Data Extended Fig. 3.
Extended Data Fig. 4 Serine synthesis promotes differentiation.
a, Western blot of Phgdh knockdown in WT EpdSCs. shPhgdh2.1 and shPhgdh2.2 represent independent transductions with same shRNA and in subsequent experiments, shPhgdh2.2 is referred to as shPhgdh2. Experiment was performed twice with similar results. shRNA sequences can be found in Supplementary Table 5. b, 48 hour population doublings of shPhgdh WT cells (n = 3 biologically independent samples). Data are mean ±SEM. c, Intracellular serine pools upon Ser/Gly starvation (n = 3 biologically independent samples). Data are mean ±SEM. d, Involucrin immunofluorescence in shPhgdh lines cultured with DMSO or DM-αKG following 24 hours of Ser/Gly starvation (n = 3 independent experiments). Data are mean ±SEM. e-f, Representative immunofluorescence (e) and quantification (f) of Involucrin in indicated conditions (n = 3 independent experiments). Data are mean ±SEM. Statistical significance was determined using a two-way ANOVA with Sidak’s multiple comparison test in panels b and f, Tukey’s multiple comparison test in panel d, and an ordinary one-way ANOVA with Dunnett’s multiple comparison test in panel c. Scanned images of unprocessed blots are shown in Source Data Extended Fig. 4. Numerical data are provided in Statistics Source Data Extended Fig. 4.
Extended Data Fig. 5 Glucose-derived serine synthesis supports the TCA cycle and αKG-driven differentiation.
a, Metabolite pools following 16 h of Ser/Gly starvation (n = 3 biologically independent samples). Data are mean ±SEM. b, Estimation of serine synthesis contribution to αKG pools (n = 6 biologically independent samples, serine uptake data from Fig. 1c) See Methods and Supplementary Tables 2 and 3 for more information. αKGSSP = αKG produced from SSP; SerSSP = serine produced from SSP; SerEC +SG = serine consumed in the presence of extracellular Ser/Gly; SerEC –SG = serine secreted in the absence of extracellular Ser/Gly; GlnIC = available intracellular glutamine pool; GlnEC +SG = glutamine consumed in the presence of extracellular Ser/Gly; GluEC +SG = glutamate secreted in the presence of extracellular Ser/Gly. Data are mean ±SEM. c, Intracellular fumarate and malate in WT EpdSCs expressing indicated hairpins following 16 h Ser/Gly deprivation (n = 3 biologically independent samples). Data are mean ±SEM. d, Involucrin staining in cells supplemented with 1 mM formate, 4 mM DM-αKG or 4 mM DM-succinate (n = 3 independent experiments). Data are mean ±SEM. e, Representative immunoblot for H3K27me3 levels. See Supplementary Table 4 for quantification of immunoblot in triplicate independent experiments. f, Immunofluorescence for H3K27me3 upon culture in low Ser/Gly or Ser/Gly-free medium (n = 3 independent experiments). Data are mean ±SEM. g, Immunofluorescence for H3K27me3 upon culture in serine free, glycine free, or Ser/Gly-free medium (n = 3 independent experiments). Data are mean ±SEM. Statistical significance was determined using a two-way ANOVA with Sidak’s multiple comparison test for panel a, with Tukey’s multiple comparison test for panel d and g, and an ordinary one-way ANOVA with Tukey’s multiple comparison test for panels c and f. Scanned images of unprocessed blots are shown in Source Data Extended Fig. 5. Numerical data are provided in Statistics Source Data Extended Fig. 5.
Extended Data Fig. 6 Role of ROS, p53 and mTORC1 signaling in serine starvation response.
a-c, Representative immunofluorescence (a) and quantification of K14 (b) and H3K27me3 (c) in WT and SOX2+ cells cultured with cell-permeable esterified reduced glutathione (eGSH), the antioxidant Trolox, the JMJD3 inhibitor GSK-J4, or the EZH2 inhibitor GSK343, performed in triplicate. (n = 3 independent experiments). d–f, Representative immunofluorescence (d) and quantification of K14 (e) and H3K27me3 (f) in WT and SOX2+ cells in indicated conditions, (n = 3 independent experiments). g, Representative immunoblot of p53 stabilization upon 24 h Ser/Gly starvation or treatment with 10 μM of the MDM2 inhibitor Nutlin-3a, performed in duplicate. Experiment was performed in triplicate with similar results. h, RT-qPCR for expression of canonical p53 target genes (n = 3 independent experiments). i, Representative immunoblot for phosphorylation of the mTORC1 target S6K following 24 h Ser/Gly starvation, performed in duplicate. Experiment was performed in triplicate with similar results. j, S6 phosphorylation in P0 mice on a control or Ser/Gly-free diet (n = 3 animals analyzed per condition). Scale bar = 50 μm. All data are mean ±SEM. Statistical significance was determined using an ordinary one-way ANOVA with Tukey’s multiple comparison test for panels b, c, e and f (P-values are relative to control) and a two-way ANOVA with Sidak’s multiple comparison test for panel h. Scanned images of unprocessed blots are shown in Source Data Extended Fig. 6. Numerical data are provided in Statistics Source Data Extended Fig. 6.
Extended Data Fig. 7 Mouse SCCs are sensitive to Ser/Gly starvation in vivo.
a. H3K27me3 levels in mouse SCCs grown in Nude mice on control or Ser/Gly-free diet (n = 8 control tumors, n = 5 –Ser/Gly tumors) Data are mean ±SEM. b, K10 expression in SCCs grown in Nude mice on control or Ser/Gly-free diet treated with 10 mg/kg GSK-J4 (n = 6 tumors per condition). Data are mean ±SEM. c, Representative hematoxylin & eosin (H&E) of SCCs. Asterisks denote keratin pearls, arrows denote keratohyalin granules, arrowheads denote intercellular bridges, all signs of differentiation in SCCs. Three tumors were analyzed per condition with similar results. d, pS6 staining in SCCs (n = 5 tumors per condition). Data are mean ±SEM. e-f, pS6 staining (e) and K10 staining (f) in SCCs grown in Nude mice treated with 4 mg/kg rapamycin. Three tumors were analyzed per condition with similar results. g, H3K27me3 staining in SCCs expressing RFP or LbNOX-Flag (n = 3 tumors per condition). Data are mean ±SEM. h, K14 and K10 staining in SCCs expressing RFP or LbNOX-Flag (n = 3 tumors per condition). Data are mean ±SEM. i, Tumor growth of SCCs expressing RFP or LbNOX-Flag (n = 20 tumors per condition). Data are mean ±SEM. P-values are comparison to RFP control at end point. Statistical significance was determined using an unpaired two-tailed student’s t-test for panels a and d, and a two-way ANOVA with Tukey’s multiple comparison test for panels b, g, h and i. Numerical data are provided in Statistics Source Data Extended Fig. 7.
Extended Data Fig. 8 Human SCCs are sensitive to Ser/Gly starvation regardless of p53 status in vivo.
a-c, K14/K10 (a), H3K27me3 (b) and growth (c) of p53 mutant A431 human SCCs grown in Nude mice (n = 4 tumors per condition for immunofluorescence analysis, n = 20 tumors per condition for growth). Data are mean ±SEM. d-f K14/K10 (d), H3K27me3 (e) and growth (f) of p53 null SCC9 human SCCs grown in Nude mice (n = 4 tumors per condition for immunofluorescence analysis, n = 20 tumors per condition for growth). Data are mean ±SEM. g, Super-enhancer epicenter reporter expression in SCC9 SCCs grown in Nude mice (n = 2 control reporter tumors, 6 mir21 tumors, 6 Klf5 tumors per condition). Data are mean. h, Immunohistochemistry of H3K27me3 in normal human skin and tongue. Statistical analysis was determined using an unpaired two-tailed student’s t-test for panels a, b, d and e, a two-way ANOVA with Tukey’s multiple comparison test for panels c and f, and with Sidak’s multiple comparison test for panel g. Numerical data are provided in Statistics Source Data Extended Fig. 8.
Extended Data Fig. 9 Glucose-derived serine synthesis suppresses tumorigenesis.
a, Representative PHGDH knockdown efficiency in mouse SCC cells. Experiment performed in duplicate independent experiments with similar results. b, Population doublings of shPhgdh SCC cells in vitro during 48 h of Ser/Gly starvation (n = 3 biologically independent samples). c, Representative H&E of shScramble and shPhgdh SCCs. Asterisks denote keratin pearls, arrows denote keratohyalin granules, arrowheads denote intercellular bridges, all signs of differentiation in SCCs. Three tumors analyzed per condition with similar results. d–l, Immunofluorescence and analysis of SOX2 (d, e), K14 (f, g), Involucrin (h, i) and Ki67 (j–l) in shScramble and shPhgdh SCCs (n = 3 tumors for shPhgdh-1 –Ser/Gly, n = 4 tumors for all other conditions). m, Growth of SCCs (n = 4 tumors per condition). n, Verification of Phgdh knockout by immunoblot in SCC cells. Experiment was performed in duplicate with similar results. sgRNA sequence information can be found in Supplementary Table 6. o, Population doublings of sgPhgdh SCC cells in vitro during 48 h of Ser/Gly starvation (n = 3 biologically independent samles). p, Growth of sgPhgdh SCC cells orthotopically grafted into Nude mice (n = 8 tumors for days 1-30, n = 24 tumors for days 1-24). Scale bar = 50 μm. All data are mean ±SEM. Statistical significance was determined using a two-way ANOVA with Sidak’s multiple comparison test in panels b and o, with Tukey’s multiple comparison test in e, g and i, with Dunnett’s multiple comparison test in panel p, and an ordinary one-way ANOVA with Tukey’s multiple comparison test in panels k and l. Scanned images of unprocessed blots are shown in Source Data Extended Fig. 9. Numerical data are provided in Statistics Source Data Extended Fig. 9.
Extended Data Fig. 10 Serine starvation drives αKG-dependent demethylation in vivo.
a, b, OGDH (a) and SDHA (b) protein knockdown efficiency in SCC cells following 24 h in culture with or without doxycycline. Western blots performed in duplicate with similar results. shRNA sequences can be found in Supplementary Table 5. c, H&E of SCCs. Asterisks denote keratin pearls, arrows denote keratohyaline granules, arrowheads denote intercellular bridges, all signs of differentiation in SCCs. Three tumors were analyzed per condition with similar results. d, Bulk H3K27me3 levels in shRenilla, shOgdh and shSdha SCCs grafted into Nude mice on control or Ser/Gly-free diet. Three tumors were analyzed per condition with similar results. Scale bar = 50 μm. Scanned images of unprocessed blots are shown in Source Data Extended Fig. 10.
Supplementary information
Supplementary Tables 1–7
Supplementary Table 1: Serine synthesis pathway western blot quantification normalized to tubulin and WT control; Supplementary Table 2: Serine standard curve data; Supplementary Table 3: Extracellular glutamine, glutamate and serine data; Supplementary Table 4: H3K27me3 immunoblot quantification normalized to H3 within genotype; Supplementary Table 5: shRNA sequences; Supplementary Table 6: sgRNA sequences; Supplementary Table 7: RT–qPCR primers.
Source data
Statistical Source Data Fig. 1
Statistical source data.
Statistical Source Data Fig. 2
Statistical source data.
Statistical Source Data Fig. 3
Statistical source data.
Statistical Source Data Fig. 4
Statistical source data.
Statistical Source Data Fig. 5
Statistical source data.
Statistical Source Data Fig. 6
Statistical source data.
Statistical Source Data Fig. 7
Statistical source data.
Statistical Source Data Extended Data Fig. 1
Statistical source data.
Unprocessed Blots Extended Data Fig. 1
Unprocessed western blots.
Statistical Source Data Extended Data Fig. 2
Statistical source data.
Statistical source Data Extended Data Fig. 3
Statistical source data.
Statistical source Data Extended Data Fig. 4
Statistical source data.
Unprocessed Blots Extended Data Fig. 4
Unprocessed western blots.
Statistical source Data Extended Data Fig. 5
Statistical source data.
Unprocessed Blots Extended Data Fig. 5
Unprocessed western blots.
Statistical source Data Extended Data Fig. 6
Statistical source data.
Unprocessed Blots Extended Data Fig. 6
Unprocessed western blots.
Statistical source Data Extended Data Fig. 7
Statistical source data.
Statistical source Data Extended Data Fig. 8
Statistical source data.
Statistical source Data Extended Data Fig. 9
Statistical source data.
Unprocessed Blots Extended Data Fig. 9
Unprocessed western blots.
Unprocessed Blots Extended Data Fig. 10
Unprocessed western blots.
Rights and permissions
About this article
Cite this article
Baksh, S.C., Todorova, P.K., Gur-Cohen, S. et al. Extracellular serine controls epidermal stem cell fate and tumour initiation. Nat Cell Biol 22, 779–790 (2020). https://doi.org/10.1038/s41556-020-0525-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-0525-9
This article is cited by
-
Inhibition of mitochondrial folate metabolism drives differentiation through mTORC1 mediated purine sensing
Nature Communications (2024)
-
Serine metabolism in macrophage polarization
Inflammation Research (2024)
-
Metabolic determinants of tumour initiation
Nature Reviews Endocrinology (2023)
-
Molecular mechanisms of cellular metabolic homeostasis in stem cells
International Journal of Oral Science (2023)
-
Metabolic pathway analysis using stable isotopes in patients with cancer
Nature Reviews Cancer (2023)