Abstract
Leukaemia stem cells (LSCs) underlie cancer therapy resistance but targeting these cells remains difficult. The Wnt–β-catenin and PI3K–Akt pathways cooperate to promote tumorigenesis and resistance to therapy. In a mouse model in which both pathways are activated in stem and progenitor cells, LSCs expanded under chemotherapy-induced stress. Since Akt can activate β-catenin, inhibiting this interaction might target therapy-resistant LSCs. High-throughput screening identified doxorubicin (DXR) as an inhibitor of the Akt–β-catenin interaction at low doses. Here we repurposed DXR as a targeted inhibitor rather than a broadly cytotoxic chemotherapy. Targeted DXR reduced Akt-activated β-catenin levels in chemoresistant LSCs and reduced LSC tumorigenic activity. Mechanistically, β-catenin binds multiple immune-checkpoint gene loci, and targeted DXR treatment inhibited expression of multiple immune checkpoints specifically in LSCs, including PD-L1, TIM3 and CD24. Overall, LSCs exhibit distinct properties of immune resistance that are reduced by inhibiting Akt-activated β-catenin. These findings suggest a strategy for overcoming cancer therapy resistance and immune escape.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
ChIP–seq, RNA-seq and ATAC-seq data that support the findings of this study have been deposited in the Gene Expression Omnibus under accession code GSE105049. Original data underlying this manuscript can be accessed from the Stowers Original Data Repository at http://www.stowers.org/research/publications/LIBPB-791. Source data for Figs. 1–6 and Extended Data Figs. 1, 3 and 5–7 are presented with the paper. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
References
Kuttesch, J. F. Jr. Multidrug resistance in pediatric oncology. Invest. New Drugs 14, 55–67 (1996).
Greaves, M. & Maley, C. C. Clonal evolution in cancer. Nature 481, 306–313 (2012).
Kreso, A. & Dick, J. E. Evolution of the cancer stem cell model. Cell Stem Cell 14, 275–291 (2014).
Dick, J. E. Stem cell concepts renew cancer research. Blood 112, 4793–4807 (2008).
Eppert, K. et al. Stem cell gene expression programs influence clinical outcome in human leukemia. Nat. Med. 17, 1086–1093 (2011).
Greaves, M. Darwinian medicine: a case for cancer. Nat. Rev. Cancer 7, 213–221 (2007).
Greaves, M. Cancer stem cells renew their impact. Nat. Med. 17, 1046–1048 (2011).
Ding, L. et al. Clonal evolution in relapsed acute myeloid leukaemia revealed by whole-genome sequencing. Nature 481, 506–510 (2012).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell. 144, 646–674 (2011).
Holohan, C., Van Schaeybroeck, S., Longley, D. B. & Johnston, P. G. Cancer drug resistance: an evolving paradigm. Nat. Rev. Cancer 13, 714–726 (2013).
Peng, W. et al. Loss of PTEN promotes resistance to T cell-mediated immunotherapy. Cancer Discov. 6, 202–216 (2016).
Fruman, D. A. et al. The PI3K pathway in human disease. Cell 170, 605–635 (2017).
Cully, M., You, H., Levine, A. J. & Mak, T. W. Beyond PTEN mutations: the PI3K pathway as an integrator of multiple inputs during tumorigenesis. Nat. Rev. Cancer 6, 184–192 (2006).
Koren, S. & Bentires-Alj, M. Tackling resistance to PI3K inhibition by targeting the epigenome. Cancer Cell 31, 616–618 (2017).
Gutierrez, A. et al. High frequency of PTEN, PI3K, and AKT abnormalities in T-cell acute lymphoblastic leukemia. Blood 114, 647–650 (2009).
Hogan, L. E. et al. Integrated genomic analysis of relapsed childhood acute lymphoblastic leukemia reveals therapeutic strategies. Blood 118, 5218–5226 (2011).
Bhatla, T. et al. Epigenetic reprogramming reverses the relapse-specific gene expression signature and restores chemosensitivity in childhood B-lymphoblastic leukemia. Blood 119, 5201–5210 (2012).
Bolouri, H. et al. The molecular landscape of pediatric acute myeloid leukemia reveals recurrent structural alterations and age-specific mutational interactions. Nat. Med. 24, 103–112 (2018).
Griffiths, E. A. et al. Acute myeloid leukemia is characterized by Wnt pathway inhibitor promoter hypermethylation. Leuk. Lymphoma 51, 1711–1719 (2010).
Dandekar, S. et al. Wnt inhibition leads to improved chemosensitivity in paediatric acute lymphoblastic leukaemia. Br. J. Haematol. 167, 87–99 (2014).
Kandoth, C. et al. Mutational landscape and significance across 12 major cancer types. Nature 502, 333–339 (2013).
Huang, J., Nguyen-McCarty, M., Hexner, E. O., Danet-Desnoyers, G. & Klein, P. S. Maintenance of hematopoietic stem cells through regulation of Wnt and mTOR pathways. Nat. Med. 18, 1778–1785 (2012).
Korkaya, H. et al. Regulation of mammary stem/progenitor cells by PTEN/Akt/β-catenin signaling. PLoS Biol. 7, e1000121 (2009).
Huang, J. et al. Pivotal role for glycogen synthase kinase-3 in hematopoietic stem cell homeostasis in mice. J. Clin. Invest. 119, 3519–3529 (2009).
Conley, S. J. et al. Antiangiogenic agents increase breast cancer stem cells via the generation of tumor hypoxia. Proc. Natl Acad. Sci. USA 109, 2784–2789 (2012).
He, X. C. et al. PTEN-deficient intestinal stem cells initiate intestinal polyposis. Nat. Genet. 39, 189–198 (2007).
Perry, J. M. et al. Cooperation between both Wnt/β-catenin and PTEN/PI3K/Akt signaling promotes primitive hematopoietic stem cell self-renewal and expansion. Genes Dev. 25, 1928–1942 (2011).
Knapp, D. J. et al. Distinct signaling programs control human hematopoietic stem cell survival and proliferation. Blood 129, 307–318 (2017).
Fruman, D. A. & Rommel, C. PI3K and cancer: lessons, challenges and opportunities. Nat. Rev. Drug Discovery 13, 140–156 (2014).
Zhou, H. et al. Combined inhibition of β-catenin and Bcr-Abl synergistically targets tyrosine kinase inhibitor-resistant blast crisis chronic myeloid leukemia blasts and progenitors in vitro and in vivo. Leukemia 31, 2065–2074 (2017).
Tenbaum, S. P. et al. β-catenin confers resistance to PI3K and AKT inhibitors and subverts FOXO3a to promote metastasis in colon cancer. Nat. Med. 18, 892–901 (2012).
Guo, W. et al. Multi-genetic events collaboratively contribute to Pten-null leukaemia stem-cell formation. Nature 453, 529–533 (2008).
Roderick, J. E. et al. c-Myc inhibition prevents leukemia initiation in mice and impairs the growth of relapsed and induction failure pediatric T-ALL cells. Blood 123, 1040–1050 (2014).
Schubbert, S. et al. Targeting the MYC and PI3K pathways eliminates leukemia-initiating cells in T-cell acute lymphoblastic leukemia. Cancer Res. 74, 7048–7059 (2014).
Huang, W., Chang, H. Y., Fei, T., Wu, H. & Chen, Y. G. GSK3β mediates suppression of cyclin D2 expression by tumor suppressor PTEN. Oncogene 26, 2471–2482 (2007).
Lechman, E. R. et al. Attenuation of miR-126 activity expands HSC in vivo without exhaustion. Cell Stem Cell 11, 799–811 (2012).
Xu, C. et al. beta-Catenin/POU5F1/SOX2 transcription factor complex mediates IGF-I receptor signaling and predicts poor prognosis in lung adenocarcinoma. Cancer Res. 73, 3181–3189 (2013).
Levine, D. A. & The Cancer Genome Atlas Research Network,. Integrated genomic characterization of endometrial carcinoma. Nature 497, 67–73 (2013).
Kaveri, D. et al. β-Catenin activation synergizes with Pten loss and Myc overexpression in Notch-independent T-ALL. Blood 122, 694–704 (2013).
Al-Dhfyan, A., Alhoshani, A. & Korashy, H. M. Aryl hydrocarbon receptor/cytochrome P450 1A1 pathway mediates breast cancer stem cells expansion through PTEN inhibition and β-Catenin and Akt activation. Mol. Cancer 16, 14 (2017).
Lee, G. et al. Phosphoinositide 3-kinase signaling mediates β-catenin activation in intestinal epithelial stem and progenitor cells in colitis. Gastroenterology 139, 869–881 (2010). 881 e861-869.
Sharma, P., Hu-Lieskovan, S., Wargo, J. A. & Ribas, A. Primary, adaptive, and acquired resistance to cancer immunotherapy. Cell 168, 707–723 (2017).
Spranger, S., Bao, R. & Gajewski, T. F. Melanoma-intrinsic β-catenin signalling prevents anti-tumour immunity. Nature 523, 231–235 (2015).
Spranger, S. & Gajewski, T. F. Impact of oncogenic pathways on evasion of antitumour immune responses. Nat. Rev. Cancer 18, 139–147 (2018).
Galluzzi, L., Buque, A., Kepp, O., Zitvogel, L. & Kroemer, G. Immunological effects of conventional chemotherapy and targeted anticancer agents. Cancer Cell 28, 690–714 (2015).
Gothert, J. R. et al. In vivo fate-tracing studies using the Scl stem cell enhancer: embryonic hematopoietic stem cells significantly contribute to adult hematopoiesis. Blood 105, 2724–2732 (2005).
Ciraolo, E., Morello, F. & Hirsch, E. Present and future of PI3K pathway inhibition in cancer: perspectives and limitations. Curr. Med. Chem. 18, 2674–2685 (2011).
Casares, N. et al. Caspase-dependent immunogenicity of doxorubicin-induced tumor cell death. J. Exp. Med. 202, 1691–1701 (2005).
Hsu, J. M. et al. STT3-dependent PD-L1 accumulation on cancer stem cells promotes immune evasion. Nat. Commun. 9, 1908 (2018).
Malta, T. M. et al. Machine learning identifies stemness features associated with oncogenic dedifferentiation. Cell 173, 338–354 e315 (2018).
Jinesh, G. G., Manyam, G. C., Mmeje, C. O., Baggerly, K. A. & Kamat, A. M. Surface PD-L1, E-cadherin, CD24, and VEGFR2 as markers of epithelial cancer stem cells associated with rapid tumorigenesis. Sci. Rep. 7, 9602 (2017).
Chen, G. Y., Tang, J., Zheng, P. & Liu, Y. CD24 and Siglec-10 selectively repress tissue damage-induced immune responses. Science 323, 1722–1725 (2009).
Hong, D. et al. Initiating and cancer-propagating cells in TEL-AML1-associated childhood leukemia. Science 319, 336–339 (2008).
Castor, A. et al. Distinct patterns of hematopoietic stem cell involvement in acute lymphoblastic leukemia. Nat. Med. 11, 630–637 (2005).
Kong, Y. et al. CD34+CD38+CD19+ as well as CD34+CD38−CD19+ cells are leukemia-initiating cells with self-renewal capacity in human B-precursor ALL. Leukemia 22, 1207–1213 (2008).
Wilson, K. et al. Flow minimal residual disease monitoring of candidate leukemic stem cells defined by the immunophenotype, CD34+CD38lowCD19+ in B-lineage childhood acute lymphoblastic leukemia. Haematologica 95, 679–683 (2010).
Eguchi, M., Eguchi-Ishimae, M. & Ishii, E. Recent progress in leukemic stem cell research for childhood leukemia. Rinsho Ketsueki 56, 1871–1881 (2015).
Kikushige, Y. et al. TIM-3 is a promising target to selectively kill acute myeloid leukemia stem cells. Cell Stem Cell 7, 708–717 (2010).
Jan, M. et al. Prospective separation of normal and leukemic stem cells based on differential expression of TIM3, a human acute myeloid leukemia stem cell marker. Proc. Natl Acad. Sci. USA 108, 5009–5014 (2011).
Miao, Y. et al. Adaptive immune resistance emerges from tumor-initiating stem cells. Cell 177, 1172–1186 (2019).
Lesche, R. et al. Cre/loxP-mediated inactivation of the murine Pten tumor suppressor gene. Genesis 32, 148–149 (2002).
Harada, N. et al. Intestinal polyposis in mice with a dominant stable mutation of the β-catenin gene. EMBO J. 18, 5931–5942 (1999).
Hu, Y. & Smyth, G. K. ELDA: extreme limiting dilution analysis for comparing depleted and enriched populations in stem cell and other assays. J. Immunol. Methods 347, 70–78 (2009).
Tran, T. H. et al. Long circulating self-assembled nanoparticles from cholesterol-containing brush-like block copolymers for improved drug delivery to tumors. Biomacromolecules 15, 4363–4375 (2014).
Freireich, E. J., Gehan, E. A., Rall, D. P., Schmidt, L. H. & Skipper, H. E. Quantitative comparison of toxicity of anticancer agents in mouse, rat, hamster, dog, monkey, and man. Cancer Chemother. Rep. 50, 219–244 (1966).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 26, 139–140 (2010).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: A method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21–29 (2015).
Acknowledgements
We thank H. Marshall, K. Zapien, J. McCann, S. Billinger, J. Haug, A. Box, C. Semerad, J. Park, L. Blunk, S. Chen, T. Parmely, A. Peak, K. Hall, B. Slaughter and J. Unruh for technical support; K. Tannen for proofreading and editing; H. Wu for providing Pten conditional-mutant mice; J. Goethert for providing HSC-SCL-Cre-ERT mice; M. Taketo for providing the conditional β-catAct mutants; and A. McMahon for providing FLAG:β-catenin ES cells. D. Xu is supported by NIH (R01 GM100701). The synthesis and characterization of polymers for NanoDXR was partly supported by a grant from the National Science Foundation NSF CAREER Award to R.M.K. (DMR-0748398) and ACS PRF 5247200 to R.M.K., managed by the American Chemical Society. The NanoDXR preparation was supported by the Academic Plan Grant from the University of Connecticut. We acknowledge support from the University of Kansas Cancer Center’s Biospecimen Repository Core Facility staff for helping obtain human specimens and the University of Kansas Cancer Center’s Cancer Center Support Grant (P30 CA168524). Research reported in this publication was supported by Stowers Institute for Medical Research, Children’s Mercy Kansas City, Braden’s Hope for Childhood Cancer, the Leukemia and Lymphoma Society (LLS), Kansas Bioscience Authority, Hall Family Foundation and the University of Kansas Cancer Center (KUCC) National Cancer Institute Cancer Center Support Grant (NCI-CCSG) P30 CA168524 and used the KUCC Lead Development and Optimization Shared Resource.
Author information
Authors and Affiliations
Contributions
J.M.P. designed and conducted the primary experiments and wrote the manuscript. F.T. conducted ChIP–seq and ATAC-seq experiments. A.R. and G.S.S. conducted high-throughput screening. T.L. conducted the clinical trial. X.C.H., A.M. and D.D. conducted transplantation and drug treatments. X.L., R.M.K., T.-H.T., P.D. and C.T.N. designed and synthesized nanoDXR. S.J.W., E.G., K.A., A.S.G., R.R., and M.B. provided insights into clinical treatment. A.G. oversaw patient biospecimen acquisition. Z.H. and D.X. conducted computational simulation. S.D. provided β-catenin inhibitor. J.N., L.R., X.Y., J.P., K.S., M.Z., A.V., P.Q., Z.L. and M.H. helped in scientific discussion and facilitated some experiments. S.C. and A.P. conducted bioinformatics analysis. L.L. provided overall supervision of the project. All authors reviewed and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 HTS screening and in vitro analysis shows DXR preferentially inhibits LSC expansion.
a, Flow chart showing HTS design. b, Vector designs for cells expressing Akt and β-catenin and TCF reporter activity. c, FRET verification between Akt and β-catenin. While FRET was observed in mCherry-β-catenin + EGFP-AKT transfected cells and could be inhibited by DXR, essentially no discernible FRET occurred when mCherry-β-catenin was transfected with EGFP alone (see also Fig. 2h-i). Two biologically independent experiments with n=62, 19, 27, and 36 independent cells for vehicle, [Low]DXR, and vehicle (negative), [Low]DXR (negative), respectively; mean ± SD is indicated. d-e, BM isolated from leukemic Pten:β-catAct mice at 8 wpi was cultured in HSC expansion media (see Methods). Doxorubicin (white), 103506 (light grey), and thioguanosine (dark grey) were added to 11, 33 or 100 nM and cultured for 72 hours and analyzed by flow cytometry for LSCs (d) and HSPCs (e) as in Fig. 1. Fold change before and after culture for each population is indicated relative to equivalent vehicle control concentrations. n=3 biologically independent replicates for each condition; mean ± SEM is indicated. Statistics determined by unpaired two-tailed t-test.
Extended Data Fig. 2 Schematic representation of experimental setup and treatment scheme for leukemic mice.
See Methods for additional detail.
Extended Data Fig. 3 Low-dose DXR inhibits downstream Wnt signaling.
a, Heatmap of Hallmark MYC target genes (V1) up and downregulated in HSPCs and LSCs treated with vehicle and LSCs treated with low-dose DXR. Data from two biological replicates of each, differing by <0.3 standard deviations, are shown. b, c, Gene ontology enrichment analysis using -log10 of the uncorrected p value as x axis. The upregulated or downregulated enriched terms are shown in red or blue, respectively. Numbers correspond to the same term upregulated in b but downregulated in c. b, Upregulated terms in LSCs vs. HSPCs in leukemic mice (treated only with vehicle control). c, Downregulated terms in LSCs from low-dose DXR vs. vehicle control treated leukemic mice. n=2 biologically independent replicates for each (see Methods). d, TCF/Lef:H2B-GFP mice were stained with CD3 and c-Kit and analyzed by FACS. Percent of TCF/Lef:H2B-GFP+ cells for each population are indicated. Mean of n=2 biologically independent replicates for each is indicated. e, TCF/Lef:H2B-GFP mice were treated with vehicle or low-dose nanoDXR and analyzed by FACS at 3 hours post-injection for TCF/Lef:H2B-GFP+ cells. n=5 biologically independent replicates for each condition; mean ± SD is indicated. Statistics determined by unpaired two-tailed t-test. f, Representative FACS plots for data quantified in (e) and including untreated, TCF/Lef:H2B-GFP negative littermate control analysis. n=4,5, and 5 biologically independent replicates as indicated; mean ± SD is indicated. Statistics determined by unpaired two-tailed t-test.
Extended Data Fig. 4 β-catenin binds multiple immune checkpoint gene loci; low-dose DXR has differential effects on IC genes in LSCs and blast cells.
a, CHIP-seq was performed on 2×107 cells from the β-catenin-3xFlag mouse ES cell line. Genome browser view of β-catenin binding density at the Pdcd1 (Pd-1), Havcr2 (Tim-3), Cd24a, and Ctla4 gene loci promoter regions and/or intergenic regions. b, ATAC-seq was used to show chromatin accessibility profiles of Wnt target genes observed in blast cells (~30k cells per replicate) and leukemic stem cells (~15k cells per replicate). c, Accessibility profiles of immuno-checkpoint genes observed by ATAC-seq in blast cells (~30k cells per replicate) and LSCs (~15k cells per replicate). Cells were sorted from BM pooled from 20 leukemia mice treated with low-dose DXR and 8 leukemia mice treated with vehicle control (b-c). Experiment was repeated 1 time with similar results.
Extended Data Fig. 5 Single-dose DXR-loaded nanoparticles substitute for multiple doses of free DXR in reducing LSCs. [Low]NanoDXR treatment reduces functional LSCs in vivo.
a, b, Leukemic mice established as described in Extended Data. 2 were treated with 5 daily injections of free DXR at 0.5 or 0.15 µg/g with and without chemotherapy. Alternatively, a single injection on day 1 of 0.8 or 2.5 µg/g of DXR-loaded nanoparticles (NanoDXR) was given with and without chemotherapy. At 10 days post-treatment, BM was analyzed by flow cytometry to determine frequency of LSCs (a) and HSPCs (b). n=6,10,3,6,10,6,3 (a) and n=6,10,8,6,10,6,3 (b) (left to right, respectively) biologically independent replicates for each condition; mean ± SEM is indicated. Note that 5 doses of 0.15 µg/g DXR is ineffective; however, a single NanoDXR injection with a similar cumulative dose (0.8 µg/g) is most effective at reducing LSCs while allowing for HSPC recovery. c-d, Cohorts of leukemic mice were prepared and treated as in Figure S2 but with low-dose NanoDXR. At 12 days post-treatment, BM was harvested from treated mice and transplanted into sub-lethally irradiated NSG recipients. c, Treatment schematic and Kaplan-Meier curves of recipient mice. The free low-dose DXR treatment group (solid line) from Fig. 5f is shown for comparison (n = 30 per group). d, Recipients of BM from low-dose DXR and low-dose NanoDXR treated leukemic mice were analyzed by flow cytometry at 6 months post-transplant for Blast cells, HSPCs and LSCs. n=27 (free_ and 24 (nano) biologically independent replicates; mean ± SD is indicated. Statistics determined by unpaired two-tailed t-test.
Extended Data Fig. 6 Low-dose DXR treatment reduces leukemia-initiating activity only in human leukemia exhibiting chemoresistant pS552-β-cat+ LSCs.
a, b, Summary of pediatric leukemia patients analyzed by FACS for LSCs and pS552-β-cat+ LSCs. B- and T-lymphoid LSCs were identified as enriched in CD45+ CD34+ CD19+ and CD45+ c-Kit+ CD3+ cells, respectively. BM samples at diagnosis and at day 29 post-chemotherapy treatment. Pt samples 019 and 034, B and T-lymphoid leukemias exhibiting chemoresistant pS552-β-cat+ LSCs, were subjected to further in vivo treatment and analysis. Pt 024 and 031, B and T-lymphoid acute leukemias lacking chemoresistant pS552-β-cat+ LSCs, were also tested. c, Experimental schematic of establishment and treatment of patient-derived xenografts (PDX) (see Methods). d, e, Kaplan-Meier survival curve of 1° PDX recipients of diagnostic and post-chemotherapy BM from Pt 019 (d) and Pt 034 (e). n=3 biologically independent replicates each. f, g, Kaplan-Meier survival curve for 2° PDX recipients treated with vehicle or low-dose nanoDXR. BM from 1° recipients of diagnostic (d) or post-chemotherapy (e) was transplanted into 2° PDX recipients and treated with low-dose NanoDXR 14 days post-transplant. h, i, FACs analysis of 2° PDX recipient BM from (f-g). Human CD45+ and human LSC engraftment was determined after succumbing to leukemia or at experimental endpoint. n=10 (each, h) and n=10 and 9 (vehicle and low-dose NanoDXR, respectively, i) biologically independent replicates; mean ± SEM is indicated. Statistics determined by unpaired two-tailed t-test. j–o, Similar analysis to D-I but of patient-derived xenografts (PDX) from Pt 024 and Pt 031, which lack chemoresistant pS552-β-cat+ LSCs. n=3 biologically independent replicates each (j-k). n = biologically independent replicates as indicated (f, g, l, m). n=9,10 (vehicle and low-dose NanoDXR, respectively, n) and n=10,9 (vehicle and low-dose NanoDXR, respectively, o) biologically independent replicates; mean ± SEM is indicated. Statistics determined by unpaired two-tailed t-test. (N.S. = not significant). Statistics determined by Log-rank (Mantel-Cox) test for survival curves.
Extended Data Fig. 7 Low-dose Daunorubicin performs similarly to Low-dose Doxorubicin.
a, b, Daunorubicin (DNR) dose-response and FACs analysis of LSC, HSPC and Blast Cell frequency in leukemic mice treated with chemotherapy with either low-dose DXR or low-dose DNR. Experiment was repeated 1 time with similar results (a). n=5 biologically independent replicates for each; mean ± SD is indicated. Statistics determined by unpaired two-tailed t-test.
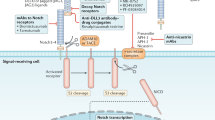
Extended Data Fig. 8 Targeting Akt-activated β-catenin dependent immune escape in LSCs.
The cooperative role of the Wnt/β-catenin and PI3K/Akt pathway in resistance to anti-cancer therapies, including immune escape, made the Pten:β-catAct double mutant mice served as an ideal model to study cancer therapy resistance. We found that cooperative Akt:β-catenin signaling is particularly critical for therapy-resistant LSCs. a, Investigating the mechanism underlying this resistance, we unexpectedly found that Akt-activated β-catenin binds to multiple IC genes, which are expressed on LSCs. b, In identifying DXR as an inhibitor of Akt:β-catenin interaction at low doses, we found that DXR could be repurposed as a targeted therapy for resistant LSCs, in part by inhibiting multiple ICs, particularly PD-1/PD-L1. Notably, LSCs but not blast cells exhibit unique properties of immune resistance, which can be reduced with low-dose DXR.
Supplementary information
Supplementary Information
Supplementary ATAC-seq QC analysis.
Supplementary Tables
Supplementary Table 1, related to Fig. 3. Limiting Dilution Assays. Recipients of sorted blast cells or LSCs (see Figure 3) from chemotherapy or [Low]DXR treated mice were analyzed at 10-12 weeks post-transplant. Recipient mice exhibiting ≥ 1% CD45Hi blast cells in bone marrow were considered engrafted. Upper and lower confidence intervals for the estimated CRU frequency were obtained using ELDA software (http://bioinf.wehi.edu.au/software/elda/index.html). Supplementary Table 2, related to Extended Data 6. Pediatric ALL Sample Characteristics and Analysis. Characteristics of pediatric acute leukemia samples examined. Supplementary Table 3, related to Figure 6l-n. Relapsed/Refactory AML Patient Characteristics. Characteristics of adult relapsed/refractory AML patients taking part in the study (Fig. 6l-n).
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 2
Unprocessed western blots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Rights and permissions
About this article
Cite this article
Perry, J.M., Tao, F., Roy, A. et al. Overcoming Wnt–β-catenin dependent anticancer therapy resistance in leukaemia stem cells. Nat Cell Biol 22, 689–700 (2020). https://doi.org/10.1038/s41556-020-0507-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-0507-y
This article is cited by
-
Crosstalk between colorectal CSCs and immune cells in tumorigenesis, and strategies for targeting colorectal CSCs
Experimental Hematology & Oncology (2024)
-
SHP-1 inhibition targets leukaemia stem cells to restore immunosurveillance and enhance chemosensitivity by metabolic reprogramming
Nature Cell Biology (2024)
-
Therapeutic role of PTEN in tissue regeneration for management of neurological disorders: stem cell behaviors to an in-depth review
Cell Death & Disease (2024)
-
Down-regulation of Musashi-2 exerts antileukemic effects on acute lymphoblastic leukemia cells and increases sensitivity to dexamethasone
Annals of Hematology (2024)
-
PRMT3-mediated arginine methylation of IGF2BP1 promotes oxaliplatin resistance in liver cancer
Nature Communications (2023)