Abstract
SLC7A11-mediated cystine uptake is critical for maintaining redox balance and cell survival. Here we show that this comes at a significant cost for cancer cells with high levels of SLC7A11. Actively importing cystine is potentially toxic due to its low solubility, forcing cancer cells with high levels of SLC7A11 (SLC7A11high) to constitutively reduce cystine to the more soluble cysteine. This presents a significant drain on the cellular NADPH pool and renders such cells dependent on the pentose phosphate pathway. Limiting glucose supply to SLC7A11high cancer cells results in marked accumulation of intracellular cystine, redox system collapse and rapid cell death, which can be rescued by treatments that prevent disulfide accumulation. We further show that inhibitors of glucose transporters selectively kill SLC7A11high cancer cells and suppress SLC7A11high tumour growth. Our results identify a coupling between SLC7A11-associated cystine metabolism and the pentose phosphate pathway, and uncover an accompanying metabolic vulnerability for therapeutic targeting in SLC7A11high cancers.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Source Data for Figs. 1–6 and Extended Data Figs. 1–7 are provided with the paper. The 33 cancer-type data were derived from the TCGA Research Network: http://cancergenome.nih.gov/. The RNA-seq data from PDXs have been deposited in dbGAP under accession number phs001980.v1.p1. All data supporting the findings of this study are available from the corresponding author on reasonable request.
References
Pavlova, N. N. & Thompson, C. B. The emerging hallmarks of cancer metabolism. Cell Metab. 23, 27–47 (2016).
Boroughs, L. K. & DeBerardinis, R. J. Metabolic pathways promoting cancer cell survival and growth. Nat. Cell Biol. 17, 351–359 (2015).
DeBerardinis, R. J. & Chandel, N. S. Fundamentals of cancer metabolism. Sci. Adv. 2, e1600200 (2016).
Wolpaw, A. J. & Dang, C. V. Exploiting metabolic vulnerabilities of cancer with precision and accuracy. Trends Cell Biol. 28, 201–212 (2018).
Bi, J., Wu, S., Zhang, W. & Mischel, P. S. Targeting cancer’s metabolic co-dependencies: A landscape shaped by genotype and tissue context. Biochim. Biophys. Acta 1870, 76–87 (2018).
Narta, U. K., Kanwar, S. S. & Azmi, W. Pharmacological and clinical evaluation of l-asparaginase in the treatment of leukemia. Crit. Rev. Oncol. Hematol. 61, 208–221 (2007).
Stipanuk, M. H. Sulfur amino acid metabolism: pathways for production and removal of homocysteine and cysteine. Annu. Rev. Nutr. 24, 539–577 (2004).
Conrad, M. & Sato, H. The oxidative stress-inducible cystine/glutamate antiporter, system xc−: cystine supplier and beyond. Amino Acids 42, 231–246 (2012).
Lewerenz, J. et al. The cystine/glutamate antiporter system xc− in health and disease: from molecular mechanisms to novel therapeutic opportunities. Antioxid. Redox Signal. 18, 522–555 (2013).
Sato, H., Tamba, M., Ishii, T. & Bannai, S. Cloning and expression of a plasma membrane cystine/glutamate exchange transporter composed of two distinct proteins. J. Biol. Chem. 274, 11455–11458 (1999).
Stipanuk, M. H., Dominy, J. E. Jr., Lee, J. I. & Coloso, R. M. Mammalian cysteine metabolism: new insights into regulation of cysteine metabolism. J. Nutr. 136, 1652S–1659S (2006).
Dixon, S. J. et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell 149, 1060–1072 (2012).
Stockwell, B. R. et al. Ferroptosis: A regulated cell death nexus linking metabolism, redox biology, and disease. Cell 171, 273–285 (2017).
Zhang, Y. et al. BAP1 links metabolic regulation of ferroptosis to tumour suppression. Nat. Cell Biol. 20, 1181–1192 (2018).
Bhutia, Y. D., Babu, E., Ramachandran, S. & Ganapathy, V. Amino acid transporters in cancer and their relevance to “glutamine addiction”: novel targets for the design of a new class of anticancer drugs. Cancer Res. 75, 1782–1788 (2015).
Lo, M., Wang, Y. Z. & Gout, P. W. The xc− cystine/glutamate antiporter: a potential target for therapy of cancer and other diseases. J. Cell. Physiol. 215, 593–602 (2008).
Koppula, P., Zhang, Y., Zhuang, L. & Gan, B. Amino acid transporter SLC7A11/xCT at the crossroads of regulating redox homeostasis and nutrient dependency of cancer. Cancer Commun. 38, 12 (2018).
Lee, W. N. et al. Mass isotopomer study of the nonoxidative pathways of the pentose cycle with [1,2-13C2]glucose. Am. J. Physiol. 274, E843–E851 (1998).
Fan, J. et al. Quantitative flux analysis reveals folate-dependent NADPH production. Nature 510, 298–302 (2014).
Lewis, C. A. et al. Tracing compartmentalized NADPH metabolism in the cytosol and mitochondria of mammalian cells. Mol. Cell. 55, 253–263 (2014).
Shin, C. S. et al. The glutamate/cystine xCT antiporter antagonizes glutamine metabolism and reduces nutrient flexibility. Nat. Commun. 8, 15074 (2017).
Koppula, P., Zhang, Y., Shi, J., Li, W. & Gan, B. The glutamate/cystine antiporter SLC7A11/xCT enhances cancer cell dependency on glucose by exporting glutamate. J. Biol. Chem. 292, 14240–14249 (2017).
Goji, T., Takahara, K., Negishi, M. & Katoh, H. Cystine uptake through the cystine/glutamate antiporter xCT triggers glioblastoma cell death under glucose deprivation. J. Biol. Chem. 292, 19721–19732 (2017).
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Bak, D. W., Bechtel, T. J., Falco, J. A. & Weerapana, E. Cysteine reactivity across the subcellular universe. Curr. Opin. Chem. Biol. 48, 96–105 (2019).
Hay, N. Reprogramming glucose metabolism in cancer: can it be exploited for cancer therapy? Nat. Rev. Cancer 16, 635–649 (2016).
Zhang, D. et al. 2-Deoxy-d-glucose targeting of glucose metabolism in cancer cells as a potential therapy. Cancer Lett. 355, 176–183 (2014).
Pereira, D. J., Schoolwerth, A. C. & Pais, V. M. Cystinuria: current concepts and future directions. Clin. Nephrol. 83, 138–146 (2015).
Elmonem, M. A. et al. Cystinosis: a review. Orphanet J. Rare Dis. 11, 47 (2016).
Siebeneicher, H. et al. Identification and optimization of the first highly selective GLUT1 inhibitor BAY-876. Chem. Med. Chem. 11, 2261–2271 (2016).
Jozwiak, P., Krzeslak, A., Pomorski, L. & Lipinska, A. Expression of hypoxia-related glucose transporters GLUT1 and GLUT3 in benign, malignant and non-neoplastic thyroid lesions. Mol. Med. Rep. 6, 601–606 (2012).
Krzeslak, A. et al. Expression of GLUT1 and GLUT3 glucose transporters in endometrial and breast cancers. Pathol. Oncol. Res. 18, 721–728 (2012).
Ancey, P. B., Contat, C. & Meylan, E. Glucose transporters in cancer—from tumor cells to the tumor microenvironment. FEBS J. 285, 2926–2943 (2018).
Faubert, B. et al. Lactate metabolism in human lung tumors. Cell 171, 358–371 (2017).
Hui, S. et al. Glucose feeds the TCA cycle via circulating lactate. Nature 551, 115–118 (2017).
Sabharwal, S. S. & Schumacker, P. T. Mitochondrial ROS in cancer: initiators, amplifiers or an Achilles’ heel? Nat. Rev. Cancer 14, 709–721 (2014).
Shimada, K., Hayano, M., Pagano, N. C. & Stockwell, B. R. Cell-line selectivity improves the predictive power of pharmacogenomic analyses and helps identify NADPH asbiomarker for ferroptosis sensitivity. Cell Chem. Biol. 23, 225–235 (2016).
Zhang, Y., Koppula, P. & Gan, B. Regulation of H2A ubiquitination and SLC7A11 expression by BAP1 and PRC1. Cell Cycle 18, 773–783 (2019).
Zhang, Y., Zhuang, L. & Gan, B. BAP1 suppresses tumor development by inducing ferroptosis upon SLC7A11 repression. Mol. Cell. Oncol. 6, 1536845 (2019).
Sykiotis, G. P. & Bohmann, D. Stress-activated cap’n’collar transcription factors in aging and human disease. Sci. Signal. 3, re3 (2010).
Rojo de la Vega, M., Chapman, E. & Zhang, D. NRF2 and the hallmarks of cancer. Cancer Cell 34, 21–43 (2018).
Liu, X. et al. LncRNA NBR2 engages a metabolic checkpoint by regulating AMPK under energy stress. Nat. Cell Biol. 18, 431–442 (2016).
Gu, Y. et al. mTORC2 regulates amino acid metabolism in cancer by phosphorylation of the cystine–glutamate antiporter xCT. Mol. Cell 67, 128–138 (2017).
Lin, A. et al. FoxO transcription factors promote AKT Ser473 phosphorylation and renal tumor growth in response to pharmacological inhibition of the PI3K–AKT pathway. Cancer Res. 74, 1682–1693 (2014).
Dai, F. et al. BAP1 inhibits the ER stress gene regulatory network and modulates metabolic stress response. Proc. Natl Acad. Sci. USA 114, 3192–3197 (2017).
Lin, A. et al. The FoxO–BNIP3 axis exerts a unique regulation of mTORC1 and cell survival under energy stress. Oncogene 33, 3183–3194 (2014).
Liu, X. & Gan, B. lncRNA NBR2 modulates cancer cell sensitivity to phenformin through GLUT1. Cell Cycle 15, 3471–3481 (2016).
Jeon, S. M., Chandel, N. S. & Hay, N. AMPK regulates NADPH homeostasis to promote tumour cell survival during energy stress. Nature 485, 661–665 (2012).
Xiao, Z. D. et al. Energy stress-induced lncRNA FILNC1 represses c-Myc-mediated energy metabolism and inhibits renal tumor development. Nat. Commun. 8, 783 (2017).
Chauhan, A. S. et al. STIM2 interacts with AMPK and regulates calcium-induced AMPK activation. FASEB J. 33, 2957–2970 (2019).
Gan, B. et al. Lkb1 regulates quiescence and metabolic homeostasis of haematopoietic stem cells. Nature 468, 701–704 (2010).
Gan, B. et al. FoxOs enforce a progression checkpoint to constrain mTORC1-activated renal tumorigenesis. Cancer Cell 18, 472–484 (2010).
Gan, B. et al. mTORC1-dependent and -independent regulation of stem cell renewal, differentiation, and mobilization. Proc. Natl Acad. Sci. USA 105, 19384–19389 (2008).
Lu, W. et al. Metabolomic analysis via reversed-phase ion-pairing liquid chromatography coupled to a stand alone orbitrap mass spectrometer. Anal. Chem. 82, 3212–3221 (2010).
Melamud, E., Vastag, L. & Rabinowitz, J. D. Metabolomic analysis and visualization engine for LC–MS data. Anal. Chem. 82, 9818–9826 (2010).
Wallace, M. et al. Enzyme promiscuity drives branched-chain fatty acid synthesis in adipose tissues. Nat. Chem. Biol. 14, 1021–1031 (2018).
Young, J. D. INCA: a computational platform for isotopically non-stationary metabolic flux analysis. Bioinformatics 30, 1333–1335 (2014).
Heinrich, P. et al. Correcting for natural isotope abundance and tracer impurity in MS-, MS/MS- and high-resolution-multiple-tracer-data from stable isotope labeling experiments with IsoCorrectoR. Sci. Rep. 8, 17910 (2018).
Badur, M. G. et al. Oncogenic R132 IDH1 mutations limit NADPH for de novo lipogenesis through (D)2-hydroxyglutarate production in fibrosarcoma cells. Cell Rep. 25, 1018–1026 (2018).
Lu, W., Wang, L., Chen, L., Hui, S. & Rabinowitz, J. D. Extraction andquantitation of nicotinamide adenine dinucleotide redox cofactors. Antioxid. Redox Signal. 28, 167–179 (2018).
Su, X., Lu, W. & Rabinowitz, J. D. Metabolite spectral accuracy on orbitraps. Anal. Chem. 89, 5940–5948 (2017).
Yan, X. et al. Inhibition of thioredoxin/thioredoxin reductase induces synthetic lethality in lung cancers with compromised glutathione homeostasis. Cancer Res. 79, 125–132 (2019).
Lee, H. et al. BAF180 regulates cellular senescence and hematopoietic stem cell homeostasis through p21. Oncotarget 7, 19134–19146 (2016).
Gan, B. et al. Role of FIP200 in cardiac and liver development and its regulation of TNFα and TSC–mTOR signaling pathways. J. Cell Biol. 175, 121–133 (2006).
Acknowledgements
We thank R. DePinho for critical reading and insightful comments. This research has been supported by the Andrew Sabin Family Fellow Award and Bridge Fund from The University of Texas MD Anderson Cancer Center, Career Enhancement Award from University of Texas Specialized Program of Research Excellence in Lung Cancer National Institutes of Health/National Cancer Institute 5P50CA070907, KC180131 from Department of Defense Kidney Cancer Research Program (to B.G.), grants from the National Institutes of Health (R01CA181196 to B.G. and R01CA188652 to C.M.M.). B.G. is an Andrew Sabin Family Fellow. Y.Z. and P.K. were Scholars at the Center for Cancer Epigenetics at The University of Texas MD Anderson Cancer Center. P.K. is also supported by the CPRIT Research Training Grant (RP170067) and Dr. John J. Kopchick Research Award from The MD Anderson UTHealth Graduate School of Biomedical Sciences. E.W.L. is supported by National Institutes of Health grant T32EB009380. PDX generation and annotation were supported by the University of Texas MD Anderson Cancer Center Moon Shots Program, Specialized Program of Research Excellence grant CA070907 and University of Texas PDX Development and Trial Center grant U54CA224065. This research was also supported by the National Institutes of Health Cancer Center Support Grant P30CA016672 to The University of Texas MD Anderson Cancer Center.
Author information
Authors and Affiliations
Contributions
X.L. and Y.Z. performed most of the experiments with assistance from P.K., G.L, L.Z. and H.L. K.O. conducted most metabolomic and isotope-tracing analyses except 3-[2H]glucose-tracing analyses. E.W.L. performed 3-[2H]glucose tracing analyses under the direction of C.M.M. J.S. conducted bioinformatics analysis under the direction of W.L. X.Z. and B.F. provided PDXs. J.Z. processed tumour and tissue samples. M.J.Y. performed histopathological analysis. K.O. and M.V.P. provided KL-11743 and designed and interpreted pharmacokinetic analysis. B.G. conceived and supervised the study and wrote most of the manuscript. All authors commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
K.O. and M.V.P. are full-time employees of Kadmon Corporation. The other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The effect of SLC7A11 overexpression on glutamate, TCA cycle and glycolysis metabolites, and the expression levels of PPP enzymes.
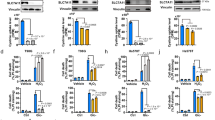
a, Western blotting showing Myc-tagged SLC7A11 expression in 786-O cells. The experiment was repeated five times, independently, with similar results. b, Bar graph showing relative fold changes of glutamate and TCA cycle metabolites in EV and SLC7A11-overexpressing 786-O cells. n=3 independent experiments. c, Western blotting showing indicated protein levels in EV and SLC7A11-overexpressing 786-O cells. The experiment was repeated twice, independently, with similar results. d, Bar graph showing relative fold changes of glycolysis metabolites in EV and SLC7A11-overexpressing 786-O cells. n=3 independent experiments. e,f, Bar graph showing the fold changes of PPP and PPP-derived intermediates induced by SLC7A11 overexpression in RCC4 or ACHN cells. n=3 independent experiments. g, Simplified schematic of glycolysis and the PPP, showing 13C labeling patterns resulting from 1,2-13C2 glucose. Red fills indicate 13C atoms. h, Glucose consumption rates in EV and SLC7A11-overexpressing 786-O cells. n=5 independent experiments. i, Simplified schematic showing the sequential transfer of deuterium labels at position 3 of glucose to NADPH and then newly synthesized palmitic acid. Red circles indicate positional deuterium labels. j, Newly synthesized deuterium labelled palmitate in EV and SLC7A11-overexpressing 786-O cells. n=3 independent experiments. In (j), data are plotted as mean ±95% confidence interval (CI). Other error bars are mean ± s.d. All p values were calculated using two-tailed unpaired Student’s t-test. Detailed statistical tests are described in the Methods. Scanned images of unprocessed blots are shown in Source Data Extended Data Fig. 1. Numeral data are provided in Statistics Source Data Extended Data Fig. 1.
Extended Data Fig. 2 G6PD knockdown sensitizes cancer cells to glucose limitation and SLC7A11 expression correlates with PPP gene expression in human cancers.
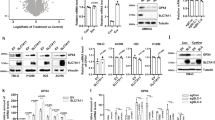
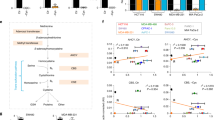
a, c, G6PD protein levels in control shRNA (shCtrl) and G6PD knockdown (shG6PD) UMRC6 (a) and A498 cells (c). The experiments were repeated twice, independently, with similar results. Scanned images of unprocessed blots are shown in Source Data Extended Data Fig. 2. b, d, Cell death analysed by PI staining in indicated cells cultured in 25 or 1 mM glucose for 24 hours. Error bars are mean± s.d., n=3 independent experiments, p values were calculated using two-tailed unpaired Student’s t-test. e, Compared to other glucose metabolism genes, PPP genes show significant positive correlations with SLC7A11 in LUAD(n=514), BLCA(n=407), HNSC(n=520), CHOL(n=36), ESCA(n=184), LUSC(n=502), and LIHC(n=371). f, Scatter plots showing the correlations between SLC7A11 and 4 PPP genes (G6PD, PGD, TALDO1, and TKT) in KIRC(n=533), LUAD(n=514), and LUSC(n=502), respectively. g, Scatter plots showing the correlations between SLC7A11 and SLC2A1 in KIRP(n=290).h, Kaplan–Meier plots of KIRP patients stratified by SLC7A11 and SLC2A1 expression levels, respectively (left 2 panels); Kaplan–Meier plots of KIRP patients stratified by unsupervised clustering on SLC7A11 and SLC2A1 expression (right 2 panels). Group 1 has lower SLC7A11 and SLC2A1 expression, while Group 2 has higher SLC7A11 and SLC2A1 expression. Detailed statistical tests of b, d and f-h are described in the Methods. Error bars are mean ± s.d, all bar graphs have 3 independent repeats. Numeral data are provided in Statistics Source Data Extended Data Fig. 2.
Extended Data Fig. 3 High expression of SLC7A11 promote disulfide stress, deplete NADPH and causes redox system collapse under glucose deprivation.
a, Simplified schematic of how SLC7A11 can be linked to NADPH and the PPP. b, c, Measurement of intracellular GSSG (b) and GSH (c) concentrations in EV and SLC7A11-overexpressing 786-O cells cultured with (+Glc) or without glucose (-Glc). d, Diagrams illustrating the structures of γ-glutamylcysteine, γ-glutamyl-cystine, GSH, and glutathionyl-cysteine. Glu: glutamate; Gly: glycine; Cys: cysteine. e, f, The relative levels of intracellular γ-glutamyl-cystine (e) and glutathionyl-cysteine (f) in EV and SLC7A11-overexpressing 786-O cells cultured with (+Glc) or without glucose (-Glc). g, Representative phase-contrast images of indicated cells cultured with or without glucose.h, Western blotting analysis of SLC7A11 protein levels in the control (sgCtrl) and SLC7A11 knockout (sgSLC-1/2) UMRC6 cells. i-l, Measurement of intracellular GSSG (i) and GSH (j) concentrations and the relative levels of intracellular γ-glutamyl-cystine (k) and glutathionyl-cysteine (l) in control (sgCtrl) and SLC7A11 knockout (sgSLC-1/2) UMRC6 cells cultured with (+Glc) or without glucose (-Glc). m, Representative phase-contrast images of indicated cells cultured with (+Glc) or without glucose (-Glc). n, o, Cystine uptake levels in EV and SLC7A11- overexpressing 786-O cells (n) or UMRC6 cells (o) upon treatment with 1 mM sulfasalazine (SAS). p-u, Cell death with or without representative phase-contrast images (p, s), NADP+/NADPH ratios (q, t), and ROS levels (r, u) of EV and SLC7A11- overexpressing 786-O or UMRC6 cells cultured in glucose-containing or glucose free medium with or without treatment of 1 mM SAS. Error bars are mean ± s.d, all bar graphs have 3 independent repeats. All scale bars=100 μm. The experiment (g, h, m, p) was repeated twice, independently, with similar results. All p values were calculated using two-tailed unpaired Student’s t-test. Scanned images of unprocessed blots are shown in Source Data Extended Data Fig. 3. Numeral data are provided in Statistics Source Data Extended Data Fig. 3.
Extended Data Fig. 4 Cystine deprivation or 2DG reverses redox defects and prevents cell death upon glucose starvation.
a-d, Measurement of intracellular GSSG (a) and GSH (b) concentrations, and the relative levels of intracellular γ-glutamyl-cystine (c) and glutathionyl-cysteine (d) in UMRC6 cells cultured with normal (+Glc), glucose free (-Glc), glucose/cystine-double free (-Glc-Cystine), or cystine free (-Cystine) medium. e, f, Measurement of NADP+/NADPH ratios (e), and ROS levels (f) in EV and SLC7A11-overexpressing 786-O cells cultured with indicated medium. g-i, Representative phase-contrast images and cell death of indicated cells cultured with indicated medium. j, k, Diagrams illustrating the structure (j) and metabolism (k) of glucose and 2DG. l-n, The relative levels of intracellular 2-deoxyglucose-6-phosphate (l), 2-deoxy-6-phosphogluconolactone (m) and 2-deoxy-6-phosphogluconate (n) in UMRC6 cells cultured in glucose-containing or glucose free medium with or without treatment of 2 mM 2DG. o-r, Measurement of intracellular GSSG (o) and GSH (p) concentrations, and the relative levels of intracellular γ-glutamyl-cystine (q) and glutathionyl-cysteine (r) in UMRC6 cells cultured in glucose-containing or glucose free medium with or without treatment of 2 mM 2DG. s, Representative phase-contrast images of UMRC6 cells cultured in glucose-containing or glucose free medium with or without treatment of 2 mM 2DG.t-w, Measurement of NADP+/NADPH ratios (t), ROS levels (u), cell death (v) and the representative phase-contrast images (w) of EV and SLC7A11-overexpressing 786-O cells cultured in glucose-containing or glucose-free medium with or without treatment of 2 mM 2DG. The experiments (g, h, i, s, w) were repeated twice, independently, with similar results. All error bars are mean± s.d., n=3 independent experiments. All scale bars=100 μm. All p values were calculated using two-tailed unpaired Student’s t-test. Numeral data are provided in Statistics Source Data Extended Data Fig. 4.
Extended Data Fig. 5 Preventing disulfide but not ROS accumulation rescues redox defects and cell death in SLC7A11-overexpressing cells under glucose starvation.
a, b, Measurement of cell death of UMRC6 or 786-O cells cultured in glucose-containing, glucose-free medium or cystine-free medium with or without treatment of 100 μM DFO or 10 μM Ferrostatin-1. c–h, Measurement intracellular levels of cysteine (c), the relative levels of intracellular γ-glutamyl-cystine (d), glutathionyl-cysteine (e), NAC-cysteine (f), GSSG/GSH ratio (g) and ROS levels (h) of UMRC6 cells cultured in glucose-containing or glucose-free medium with or without treatment of 2 mM NAC. i, The solubility of different amino acids. j–o, Measurement intracellular levels of cysteine (j), the relative levels of intracellular γ-glutamyl-cystine (k), glutathionyl-cysteine (l), GSSG/GSH ratio (m), ROS levels (n) and Cysteine-penicillamine (o) of UMRC6 cells cultured in glucose-containing or glucose-free medium with or without treatment of 2 mM D-Penicillamine or L-Penicillamine. p–t, Measurement intracellular levels of cysteine (p), the relative levels of intracellular γ-glutamyl-cystine (q), glutathionyl-cysteine (r), GSSG/GSH ratio (s) and ROS levels (t) of UMRC6 cells cultured in glucose-containing or glucose-free medium with or without treatment of TCEP. u-y, Measurement intracellular levels of cysteine (u), the relative levels of intracellular γ-glutamyl-cystine (v), glutathionyl-cysteine (w), GSSG/GSH ratio (x) and ROS levels (y) of UMRC6 cells cultured in glucose-containing or glucose-free medium with or without treatment of 1 mM 2ME. Except i, all other error bars are mean± s.d., n=3 independent experiments. All p values were calculated using two-tailed unpaired Student’s t-test. Detailed statistical tests are described in the Methods. Numeral data are provided in Statistics Source Data Extended Data Fig. 5.
Extended Data Fig. 6 Cancer cells with high SLC7A11 expression are sensitive to GLUT inhibition.
a, Cell death of EV and SLC7A11- overexpressing 786-O cells treated with 0.125-0.5 mM 6-AN. b, Cell death of EV and SLC7A11- overexpressing 786-O cells treated with 0.1 mM epiandrosterone (EA). c, Quantification of NADP+/NADPH ratios in EV and SLC7A11- overexpressing 786-O cells treated with normal (+Glc), glucose free (-Glc) medium, or normal medium containing 0.1 mM EA. d, Quantification of NADP+/NADPH ratios in UMRC6 cells treated with normal (+Glc), glucose free (-Glc), glucose/cystine double free medium (-Glc-Cystine), or normal medium containing 0.1 mM EA. e, SLC7A11 protein levels in control (sgCtrl) and SLC7A11 knockout (sgSLC7A11) NCI-H226 cells were measured by western blotting. The experiment was repeated twice, independently, with similar results. f, Measurement of GSSG/GSH ratios in EV and SLC7A11-overexpressing 786-O cells treated with KL-11743, BAY-876 or cultured in glucose free medium. g, Western blotting analysis of indicated proteins in ACHN cells with SLC7A11 and/or G6PD overexpression. The experiment was repeated twice, independently, with similar results. All error bars are mean± s.d., n=3 independent experiments. All p values were calculated using two-tailed unpaired Student’s t-test. Detailed statistical tests are described in the Methods. Scanned images of unprocessed blots are shown in Source Data Extended Data Fig. 6. Numeral data are provided in Statistics Source Data Extended Data Fig. 6.
Extended Data Fig. 7 SLC7A11-high tumors are sensitive to GLUT inhibitor.
a, Plasma levels of GLUT inhibitor KL-11743 were measured in mice at different time points after intraperitoneal injection. Error bars are mean ± s.d, n=4 independent repeats. b, End-point weights of NCI-H226 xenograft tumors with indicated genotypes treated with KL-11743 or vehicle. Error bars are mean ± s.d., n=9 independent repeats. c, End-point weights of ACHN xenograft tumors with indicated genotypes treated with BAY-876, KL-11743, or vehicle. Error bars are mean ± s.d., n=8 independent repeats. d-h, End-point weights of PDX xenograft tumors with indicated genotypes treated with KL-11743 or vehicle. . Error bars are mean ± s.d., n=6 (d: KL-11743, f-h) or7 (d: vehicle, e) independent repeats. i, Representative hematoxylin and eosin staining of major organs from mice treated with vehicle or GLUT inhibitors. The experiment was repeated twice, independently, with similar results. Scale bars=50 μm. j-p, Mice weights of indicated cell line-xenografts or PDXs at different time points treated with KL-11743 or vehicle. Error bars are mean ± s.d., n=6 (l: KL-11743, n-p), 7 (l: vehicle, m), 8 (k) or 9 (j) independent repeats. All p values were calculated using two-tailed unpaired Student’s t-test. Detailed statistical tests are described in the Methods. Numeral data are provided in Statistics Source Data Extended Data Fig. 7.
Extended Data Fig. 8 The working model depicting how SLC7A11 regulates pentose phosphate pathway dependency and glucose-deprivation-induced cell death.
See discussion for detailed description. PPP: pentose phosphate pathway; GLUT: glucose transporter.
Extended Data Fig. 9 An example for the gating strategy of Flow Cytometry.
Initial cell population gating (FSC-Area VS FSC-Height) was adopted to make sure only single cells were used for analysis.
Supplementary information
Supplementary Information
Supplementary Table 1: glucose metabolism related genes. Supplementary Table 2: summary of various approaches on rescuing redox defects and cell death in SLC7A11-high cancer cells under glucose starvation. Supplementary Table 3: oligos and shRNA sequences. Supplementary Table 4: information on human research participants (age, gender, genotypic information, diagnosis and treatment categories) of PDXs in this study.
Source data
Source Data Fig. 1
Statistical source data
Source Data Fig. 2
Statistical source data
Source Data Fig. 2
Unprocessed western blots
Source Data Fig. 3
Statistical source data
Source Data Fig. 4
Statistical source data
Source Data Fig. 5
Statistical source data
Source Data Fig. 6
Statistical source data
Source Data Fig. 6
Unprocessed western blots
Source Data Extended Data Fig. 1
Statistical source data
Source Data Extended Data Fig. 1
Unprocessed western blots
Source Data Extended Data Fig. 2
Statistical source data
Source Data Extended Data Fig. 2
Unprocessed western blots
Source Data Extended Data Fig. 3
Statistical source data
Source Data Extended Data Fig. 3
Unprocessed western blots
Source Data Extended Data Fig. 4
Statistical source data
Source Data Extended Data Fig. 5
Statistical source data
Source Data Extended Data Fig. 6
Statistical source data
Source Data Extended Data Fig. 6
Unprocessed western blots
Source Data Extended Data Fig. 7
Statistical source data
Rights and permissions
About this article
Cite this article
Liu, X., Olszewski, K., Zhang, Y. et al. Cystine transporter regulation of pentose phosphate pathway dependency and disulfide stress exposes a targetable metabolic vulnerability in cancer. Nat Cell Biol 22, 476–486 (2020). https://doi.org/10.1038/s41556-020-0496-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-0496-x
This article is cited by
-
Molecular map of disulfidptosis-related genes in lung adenocarcinoma: the perspective toward immune microenvironment and prognosis
Clinical Epigenetics (2024)
-
Establishment of a prognostic risk prediction model incorporating disulfidptosis-related lncRNA for patients with prostate cancer
BMC Cancer (2024)
-
Serine synthesis and catabolism in starved lung cancer and primary bronchial epithelial cells
Cancer & Metabolism (2024)
-
Disulfidptosis-related signature predicts prognosis and characterizes the immune microenvironment in hepatocellular carcinoma
Cancer Cell International (2024)
-
Disulfidptosis-related prognostic signature correlates with immunotherapy response in colorectal cancer
Scientific Reports (2024)