Abstract
In mouse embryonic stem cells (mESCs), chemical blockade of Gsk3α/β and Mek1/2 (2i) instructs a self-renewing ground state whose endogenous inducers are unknown. Here we show that the axon guidance cue Netrin-1 promotes naive pluripotency by triggering profound signalling, transcriptomic and epigenetic changes in mESCs. Furthermore, we demonstrate that Netrin-1 can substitute for blockade of Gsk3α/β and Mek1/2 to sustain self-renewal of mESCs in combination with leukaemia inhibitory factor and regulates the formation of the mouse pluripotent blastocyst. Mechanistically, we reveal how Netrin-1 and the balance of its receptors Neo1 and Unc5B co-regulate Wnt and MAPK pathways in both mouse and human ESCs. Netrin-1 induces Fak kinase to inactivate Gsk3α/β and stabilize β-catenin while increasing the phosphatase activity of a Ppp2r2c-containing Pp2a complex to reduce Erk1/2 activity. Collectively, this work identifies Netrin-1 as a regulator of pluripotency and reveals that it mediates different effects in mESCs depending on its receptor dosage, opening perspectives for balancing self-renewal and lineage commitment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Deep sequencing and ChIP–seq data that support the findings of this study have been deposited in the Gene Expression Omnnibus under accession code GSE102831. Previously published sequencing data that were re-analysed here are available under accession code E-MTAB-2958, E-MTAB-2959 (ref. 31), GSE81285 (ref. 30) and GSE31381 (ref. 52). All other data supporting the findings of this study are available from the corresponding author on reasonable request.
References
Evans, M. J. & Kaufman, M. H. Establishment in culture of pluripotential cells from mouse embryos. Nature 292, 154–156 (1981).
Martin, G. R. Isolation of a pluripotent cell line from early mouse embryos cultured in medium conditioned by teratocarcinoma stem cells. Proc. Natl Acad. Sci. USA 78, 7634–7638 (1981).
Dunn, S. J., Martello, G., Yordanov, B., Emmott, S. & Smith, A. G. Defining an essential transcription factor program for naive pluripotency. Science 344, 1156–1160 (2014).
Williams, R. L. et al. Myeloid leukaemia inhibitory factor maintains the developmental potential of embryonic stem cells. Nature 336, 684–687 (1988).
Sato, N., Meijer, L., Skaltsounis, L., Greengard, P. & Brivanlou, A. H. Maintenance of pluripotency in human and mouse embryonic stem cells through activation of Wnt signaling by a pharmacological GSK-3-specific inhibitor. Nat. Med. 10, 55–63 (2004).
Ying, Q. L., Nichols, J., Chambers, I. & Smith, A. BMP induction of Id proteins suppresses differentiation and sustains embryonic stem cell self-renewal in collaboration with STAT3. Cell 115, 281–292 (2003).
Kunath, T. et al. FGF stimulation of the Erk1/2 signalling cascade triggers transition of pluripotent embryonic stem cells from self-renewal to lineage commitment. Development 134, 2895–2902 (2007).
Marks, H. et al. The transcriptional and epigenomic foundations of ground state pluripotency. Cell 149, 590–604 (2012).
Ying, Q. L. et al. The ground state of embryonic stem cell self-renewal. Nature 453, 519–523 (2008).
Ficz, G. et al. FGF signaling inhibition in ESCs drives rapid genome-wide demethylation to the epigenetic ground state of pluripotency. Cell Stem Cell 13, 351–359 (2013).
Buehr, M. et al. Capture of authentic embryonic stem cells from rat blastocysts. Cell 135, 1287–1298 (2008).
Li, P. et al. Germline competent embryonic stem cells derived from rat blastocysts. Cell 135, 1299–1310 (2008).
Choi, J. et al. Prolonged Mek1/2 suppression impairs the developmental potential of embryonic stem cells. Nature 548, 219–223 (2017).
Serafini, T. et al. The netrins define a family of axon outgrowth-promoting proteins homologous to C. elegans UNC-6. Cell 78, 409–424 (1994).
Kennedy, T. E., Serafini, T., de la Torre, J. R. & Tessier-Lavigne, M. Netrins are diffusible chemotropic factors for commissural axons in the embryonic spinal cord. Cell 78, 425–435 (1994).
Cirulli, V. & Yebra, M. Netrins: beyond the brain. Nat. Rev. Mol. Cell Biol. 8, 296–306 (2007).
Grandin, M. et al. Structural decoding of the netrin-1/UNC5 interaction and its therapeutical implications in cancers. Cancer Cell 29, 173–185 (2016).
Bell, C. H. et al. Structure of the repulsive guidance molecule (RGM)-neogenin signaling hub. Science 341, 77–80 (2013).
Rajagopalan, S. et al. Neogenin mediates the action of repulsive guidance molecule. Nat. Cell Biol. 6, 756–762 (2004).
Xu, K. et al. Neural migration. Structures of netrin-1 bound to two receptors provide insight into its axon guidance mechanism. Science 344, 1275–1279 (2014).
Ko, S. Y., Dass, C. R. & Nurgali, K. Netrin-1 in the developing enteric nervous system and colorectal cancer. Trends Mol. Med. 18, 544–554 (2012).
Hong, K. et al. A ligand-gated association between cytoplasmic domains of UNC5 and DCC family receptors converts netrin-induced growth cone attraction to repulsion. Cell 97, 927–941 (1999).
Lu, X. et al. The netrin receptor UNC5B mediates guidance events controlling morphogenesis of the vascular system. Nature 432, 179–186 (2004).
Ozmadenci, D. et al. Netrin-1 regulates somatic cell reprogramming and pluripotency maintenance. Nat. Commun. 6, 7398 (2015).
Rajasekharan, S. & Kennedy, T. E. The netrin protein family. Genome Biol 10, 239 (2009).
Wray, J. et al. Inhibition of glycogen synthase kinase-3 alleviates Tcf3 repression of the pluripotency network and increases embryonic stem cell resistance to differentiation. Nat. Cell Biol. 13, 838–845 (2011).
Skarnes, W. C., Moss, J. E., Hurtley, S. M. & Beddington, R. S. Capturing genes encoding membrane and secreted proteins important for mouse development. Proc. Natl Acad. Sci. USA 92, 6592–6596 (1995).
Kumar, R. M. et al. Deconstructing transcriptional heterogeneity in pluripotent stem cells. Nature 516, 56–61 (2014).
Guo, G. et al. Serum-based culture conditions provoke gene expression variability in mouse embryonic stem cells as revealed by single-cell analysis. Cell Rep. 14, 956–965 (2016).
Bulut-Karslioglu, A. et al. Inhibition of mTOR induces a paused pluripotent state. Nature 540, 119–123 (2016).
Boroviak, T. et al. Lineage-specific profiling delineates the emergence and progression of naive pluripotency in mammalian embryogenesis. Developmental Cell 35, 366–382 (2015).
Galonska, C., Ziller, M. J., Karnik, R. & Meissner, A. Ground state conditions induce rapid reorganization of core pluripotency factor binding before global epigenetic reprogramming. Cell Stem Cell 17, 462–470 (2015).
Habibi, E. et al. Whole-genome bisulfite sequencing of two distinct interconvertible DNA methylomes of mouse embryonic stem cells. Cell Stem Cell 13, 360–369 (2013).
von Meyenn, F. et al. Impairment of DNA methylation maintenance is the main cause of global demethylation in naive embryonic stem cells. Molecular Cell 62, 848–861 (2016).
Beurel, E., Grieco, S. F. & Jope, R. S. Glycogen synthase kinase-3 (GSK3): regulation, actions, and diseases. Pharmacol. Ther. 148, 114–131 (2015).
Guenebeaud, C. et al. The dependence receptor UNC5H2/B triggers apoptosis via PP2A-mediated dephosphorylation of DAP kinase. Molecular Cell 40, 863–876 (2010).
Ren, X. R. et al. Focal adhesion kinase in netrin-1 signaling. Nat. Neurosci. 7, 1204–1212 (2004).
Liu, G. et al. Netrin requires focal adhesion kinase and Src family kinases for axon outgrowth and attraction. Nat. Neurosci. 7, 1222–1232 (2004).
Moore, S. W. & Kennedy, T. E. Protein kinase A regulates the sensitivity of spinal commissural axon turning to netrin-1 but does not switch between chemoattraction and chemorepulsion. J. Neurosci. 26, 2419–2423 (2006).
Qu, C. et al. c-Jun N-terminal kinase 1 (JNK1) is required for coordination of netrin signaling in axon guidance. J. Biol. Chem. 288, 1883–1895 (2013).
Gao, C. et al. FAK/PYK2 promotes the Wnt/β-catenin pathway and intestinal tumorigenesis by phosphorylating GSK3β. eLife 4, e10072 (2015).
Sangodkar, J. et al. All roads lead to PP2A: exploiting the therapeutic potential of this phosphatase. FEBS J. 283, 1004–1024 (2016).
Batut, J. et al. Two highly related regulatory subunits of PP2A exert opposite effects on TGF-β/Activin/Nodal signalling. Development 135, 2927–2937 (2008).
Dominici, C. et al. Floor-plate-derived netrin-1 is dispensable for commissural axon guidance. Nature 545, 350–354 (2017).
Bin, J. M. et al. Complete loss of netrin-1 results in embryonic lethality and severe axon guidance defects without increased neural cell death. Cell Rep. 12, 1099–1106 (2015).
Nakamura, T. et al. A developmental coordinate of pluripotency among mice, monkeys and humans. Nature 537, 57–62 (2016).
Heffner, C. S. et al. Supporting conditional mouse mutagenesis with a comprehensive cre characterization resource. Nat. Commun. 3, 1218 (2012).
Hayashi, K., Ohta, H., Kurimoto, K., Aramaki, S. & Saitou, M. Reconstitution of the mouse germ cell specification pathway in culture by pluripotent stem cells. Cell 146, 519–532 (2011).
ten Berge, D. et al. Embryonic stem cells require Wnt proteins to prevent differentiation to epiblast stem cells. Nat. Cell Biol. 13, 1070–1075 (2011).
Mitra, S. K. & Schlaepfer, D. D. Integrin-regulated FAK–Src signaling in normal and cancer cells. Curr. Opin. Cell Biol. 18, 516–523 (2006).
Nichols, J., Chambers, I., Taga, T. & Smith, A. Physiological rationale for responsiveness of mouse embryonic stem cells to gp130 cytokines. Development 128, 2333–2339 (2001).
Correa-Cerro, L. S. et al. Generation of mouse ES cell lines engineered for the forced induction of transcription factors. Sci. Rep. 1, 167 (2011).
Acknowledgements
We are grateful to the PBES Lyon for technical assistance. We thank V. Azuara and D. Stupack for critical reading of the manuscript and L. Favre-Louis for technical assistance. This work was supported by institutional grants from INSERM/CNRS, Atip-avenir, Plan cancer, La ligue contre le cancer nationale et régionale (F.L.), INCa (F.L.), Fondation ARC (F.L., G.F. and D.O.), Centre Léon Bérard (F.L. and A.H.), Fondation pour la recherche médicale (F.L.), National Institutes of Health (R01-HD081534 (B.J.M.)), ANR (P.M.), ERC (P.M.) Max Planck Society (A.M.) and the DFG Forschergruppe 2722 (M.K.).
Author information
Authors and Affiliations
Contributions
A.H. and G.F. performed most of the experiments in Figs. 1–7. D.O. performed experiments in Figs. 1, 5 and 7. X.G. and N.C. performed experiments in Figs. 1, 3 and 5. A.M., C.G. and C.R. performed and analysed ChIP–seq experiments. J.C. and N.R. carried out the bioinformatic analyses. M.K. and T.I. produced recombinant Netrin-1 used in Figs. 4 and 5. P.W. performed teratoma experiments. F.L., A.H., G.F. and D.O. designed experiments. F.L. initiated, designed and supervised the study. F.L. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
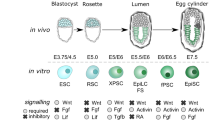
Extended Data Fig. 1 Netrin-1 is expressed in naive pluripotent cells in vitro.
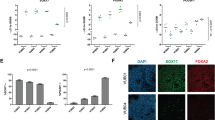
(a) Data present FPKM values for Ntn1, Ntn4, Ntn5, NtnG1 and NtnG2 in serum/Lif mESCs treated or not with Mek1/2-inh (PD), Gsk3α/β-inh (CHIR) or both (2i). 2 independent experiments. (b) Western blot depicting netrin-1 levels in human iPS cells treated similarly as (a) (3 independent experiments). (c) Netrin-1 transcripts level in indicated mESCs. Q-RTPCR data are expressed relative to mESCs as the mean ± s.d. (n = 3 independent experiments). Student’s t-test was used and two-tailed p-values are indicated. (d) Netrin-1 western blot in indicated mESCs (3 independent experiments). (e) Netrin-1 expression in single mESCs in Serum/Lif and 2i. Single-cell transcriptomic data are extracted from Kumar et al., 2014, ref. 28. n = number of cells analysed in each condition. (f) Netrin-1 mean expression in single mESCs. Data are extracted from Kumar et al., 2014, ref. 28. n = number of cells analysed in each condition. The bar represents the mean ± s.d. of netrin-1 expression in the 2 conditions. Student t-test was used and two-sided p-value is indicated.
Extended Data Fig. 2 Netrin-1 triggers pluripotency features partially overlapping with the ground state.
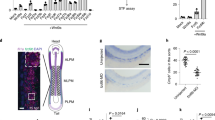
(a) Netrin-1 receptors transcript levels in mouse ES and iPS cells. RNA-seq data are presented as FPKM values and expressed as the mean ± s.d. (n = 3 independent experiments). (b) Netrin-1 receptors expression during early mouse development. Data, extracted from Boroviak. et al., 2015, ref. 31, present transcripts level in FPKM. Data are presented as the mean ± s.d. (n = 3 independent experiments). (c) Western blot performed in the indicated cell lines (3 independent experiments). (d) Western blot performed in the indicated mESCs grown in serum/Lif (3 independent experiments). (e) FACS analysis (FSC/SSC) of the different populations grown in serum/Lif. (f) netrin-1 expression in ES cells subpopulations. Data are extracted from Guo et al. 2016, ref. 29. Esrrb expression is used to distinguish quartiles of Esrrbhigh (>Q1) and Esrrblow (<Q1) cells. netrin-1 expression is analysed in the corresponding quartile. n = 48 total cells were analysed, 12 cells for <Q1 and 36 cells for >Q1. Student’s t-test was used and two-tailed p-value is indicated. (g) Scheme depicting exit from pluripotency assays. (h) Pictures of a single experiment representative of three independent ones. (i) Colony counting. Data are the mean ± s.d. (n = 3 independent experiments). Student’s t-test was used and two-sided p-values are indicated. (j) Q-RTPCR depicts transcript level in mESCs and EBs generated with indicated cell lines. Data are the mean ± s.d. n = 3 independent experiments with exclusion of outliers. (k) Statistical overrepresentation analysis. Panther DB was used to detect overrepresented GO within differentially expressed genes. A Fisher’s exact two-sided test was used to calculate p-values. n = 3 independent samples. (l) Proliferation curves. Data are the mean ± s.d. of 2 independent experiments. (m) Cell cycle features. Data are the mean ± s.d. (n = 2 independent experiments). (n) Violin plots displaying methylation levels for n = 1.3 M matched CpGs. Bold line indicates 25-75th percentile, white dot indicates median. (o) Dnmt3A and Dnmt3B expression levels. Data are the mean ± s.d. (n = 3 independent experiments) of normalized counts. Student’s T test was used and two-sided p-values are indicated.
Extended Data Fig. 3 Molecular cascade downstream of Netrin-1 in mESCs.
(a) Knockdown efficiency of Fak in mESCs. Q-RTPCR depicts Fak transcript level following transfection of netrin-1 mESCs with independent siRNA. Data, normalised to si#control mESCs, are the mean +/- sd of n = 3 independent experiments. Student T-test was used and two-sided p-values are indicated. (b) Effect of netrin-1 signalling on Lif sensitivity. Control and netrin-1 (WT) mESCs were serum-starved ON and stimulated with Lif for 10 mins prior to samples collection (3 independent experiments). (c) Knockdown efficiency of Ppp2ca and Ppp2r2c in mESCs. Similar settings as (a). Data, normalised to si#control mESCs, are the mean +/− sd of n = 3 independent experiments. Student T-test was used and two-sided p-values are indicated.
Extended Data Fig. 4 Endogenous Netrin-1 controls pluripotency features.
(a) Western blot in Ntn1fl/fl mESCs treated or not with 4′OH-tamoxifen (TAM) for 3 days before collection (3 independent experiments). (b) Q-RTPCR depicts Fgf5, Otx2, Gata4 and Gata6 transcript level following netrin-1 depletion in mESCs. Data are the mean +/− sd of n = 3 independent experiments. Student T-test was used and two-sided p-values are indicated. (c) Proliferation curves. The Ntn1fl/fl mESCs, treated or not with TAM, were counted at each passage in serum/Lif. Data are the mean +/- sd of n = 3 independent experiments. (d) Cell death analysis. The Ntn1fl/fl mESCs, treated or not with TAM, were grown for 2 days in N2B27+Lif before PI-AnnexinV staining was performed. The left panel presents a representative FACS profile and the right panel a graph of mean data ± s.d. (n = 3 independent experiments). Value 100% is given to the percentage of live cells in untreated Ntn1fl/fl mESCs. Student T-test was used and two-sided p-values are indicated. (e) Brightfield pictures of Ntn1fl/fl mESCs treated or not with 4’OH-tamoxifen and subsequently maintained in culture for 22 passages. Bars: 50 µm. 3 independent experiments. (f) Scheme of the crispr/cas9 guides. The grey boxes correspond to exons, and pink arrows indicate the 2 independent guides for each locus. (g) Self-renewal assay. Control and netrin-1 KO mESCs are plated at clonal density in serum+Lif (left panel) or serum+Lif+2i (right panel) for 7 days before AP positive colonies was scored. Data are mean ± s.d. (n = 3 independent experiments). Student’s t-test was used and two-sided p-values are indicated. (h) Gene expression in single blastomeres. Data, extracted from Nakamura et al., 2016 (ref. 46), correspond to FPKM values. n = 9 nanog positive cells n = 12 gata6 positive blastomeres. Each dot corresponds to a cell, the bar is the mean ± s.d. Student T-test was used and two-sided p-values are indicated.
Extended Data Fig. 5 Netrin-1 controls coordinated differentiation.
(a) Neo1 and Unc5B expression in epiblast-like cells (EpiLC). Q-RTPCR data are expressed relative to mESCs as the mean ± s.d. (n = 3 independent experiments). (b) Pictures of teratoma obtained following injection of Ntn1fl/fl mESCs treated (right panel) or not (left panel) with TAM 24 hours prior to injection. 4 independent teratoma per condition were analysed. (c) Q-RTPCR depicts Nestin and βIII-tubulin levels at day 8 of differentiation in N2B27-Lif. Data are normalized to housekeeping genes and value 1 is given to day8 Ctrl mESCs. Data are the mean ± s.d. (n = 3 independent experiments). Student’s t-test was used and two-sided p-values are indicated. (d) Cell death analysis. The Ntn1fl/fl mESCs, treated or not with TAM, were grown for 2 days in N2B27-Lif before PI-AnnexinV staining was performed. The left panel present a representative FACS profile and the right panel a graph of mean data ± s.d. (n = 3 independent experiments). Value 100% is given to the percentage of live cells in untreated Ntn1fl/fl mESCs. Student T-test was used and two-sided p-values are indicated.
Supplementary information
Supplementary Table 1
Sequences, antibodies and primers.
Source data
Source Data Fig. 1
Statistical source data
Source Data Fig. 1
Unprocessed western blots
Source Data Fig. 2
Statistical source data
Source Data Fig. 2
Unprocessed western blots
Source Data Fig. 3
Statistical source data
Source Data Fig. 3
Unprocessed western blots
Source Data Fig. 4
Statistical source data
Source Data Fig. 4
Unprocessed western blots
Source Data Fig. 5
Statistical source data
Source Data Fig. 5
Unprocessed western blots
Source Data Fig. 6
Statistical source data
Source Data Fig. 7
Statistical source data
Source Data Fig. 7
Unprocessed western blots
Source Data Extended Data Fig. 1
Statistical source data
Source Data Extended Data Fig. 1
Unprocessed western blots
Source Data Extended Data Fig. 2
Statistical source data
Source Data Extended Data Fig. 2
Unprocessed western blots
Source Data Extended Data Fig. 3
Statistical source data
Source Data Extended Data Fig. 3
Unprocessed western blots
Source Data Extended Data Fig. 4
Statistical source data
Source Data Extended Data Fig. 4
Unprocessed western blots
Source Data Extended Data Fig. 5
Statistical source data
Rights and permissions
About this article
Cite this article
Huyghe, A., Furlan, G., Ozmadenci, D. et al. Netrin-1 promotes naive pluripotency through Neo1 and Unc5b co-regulation of Wnt and MAPK signalling. Nat Cell Biol 22, 389–400 (2020). https://doi.org/10.1038/s41556-020-0483-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-0483-2
This article is cited by
-
Netrin-1 Alleviates Early Brain Injury by Regulating Ferroptosis via the PPARγ/Nrf2/GPX4 Signaling Pathway Following Subarachnoid Hemorrhage
Translational Stroke Research (2024)
-
Shaping the brain vasculature in development and disease in the single-cell era
Nature Reviews Neuroscience (2023)
-
Molecular versatility during pluripotency progression
Nature Communications (2023)
-
Netrin-1 blockade inhibits tumor associated Myeloid-derived suppressor cells, cancer stemness and alleviates resistance to chemotherapy and immune checkpoint inhibitor
Cell Death & Differentiation (2023)
-
Dichotomous role of Shp2 for naïve and primed pluripotency maintenance in embryonic stem cells
Stem Cell Research & Therapy (2022)