Abstract
Loss of the fragile X protein FMRP is a leading cause of intellectual disability and autism1,2, but the underlying mechanism remains poorly understood. We report that FMRP deficiency results in hyperactivated nonsense-mediated mRNA decay (NMD)3,4 in human SH-SY5Y neuroblastoma cells and fragile X syndrome (FXS) fibroblast-derived induced pluripotent stem cells (iPSCs). We examined the underlying mechanism and found that the key NMD factor UPF1 binds directly to FMRP, promoting FMRP binding to NMD targets. Our data indicate that FMRP acts as an NMD repressor. In the absence of FMRP, NMD targets are relieved from FMRP-mediated translational repression so that their half-lives are decreased and, for those NMD targets encoding NMD factors, increased translation produces abnormally high factor levels despite their hyperactivated NMD. Transcriptome-wide alterations caused by NMD hyperactivation have a role in the FXS phenotype. Consistent with this, small-molecule-mediated inhibition of hyperactivated NMD, which typifies iPSCs derived from patients with FXS, restores a number of neurodifferentiation markers, including those not deriving from NMD targets. Our mechanistic studies reveal that many molecular abnormalities in FMRP-deficient cells are attributable—either directly or indirectly—to misregulated NMD.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Sequencing datasets (FASTQ files), including RNA-seq and RIP-seq, have been deposited in the DNA Data Bank of Japan under the accession number DRA005644. Proteomics datasets have been deposited in the ProteomeXchange Consortium under the accession number PXD014901. Source data are provided with this paper. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
Code availability
All of the scripts used for data processing and statistical analysis were written in Python, Perl or R and are available upon request.
References
Richter, J. D., Bassell, G. J. & Klann, E. Dysregulation and restoration of translational homeostasis in fragile X syndrome. Nat. Rev. Neurosci. 16, 595–605 (2015).
Hagerman, R. J. et al. Fragile X syndrome. Nat. Rev. Dis. Prim. 3, 17065 (2017).
Jaffrey, S. R. & Wilkinson, M. F. Nonsense-mediated RNA decay in the brain: emerging modulator of neural development and disease. Nat. Rev. Neurosci. 19, 715–728 (2018).
Kurosaki, T., Popp, M. W. & Maquat, L. E. Quality and quantity control of gene expression by nonsense-mediated mRNA decay. Nat. Rev. Mol. Cell Biol. 20, 406–420 (2019).
Kurosaki, T. et al. A post-translational regulatory switch on UPF1 controls targeted mRNA degradation. Genes Dev. 28, 1900–1916 (2014).
Imamachi, N., Salam, K. A., Suzuki, Y. & Akimitsu, N. A GC-rich sequence feature in the 3′ UTR directs UPF1-dependent mRNA decay in mammalian cells. Genome Res. 27, 407–418 (2017).
Chan, W.-K. et al. An alternative branch of the nonsense-mediated decay pathway. EMBO J. 26, 1820–1830 (2007).
Huang, L. et al. RNA homeostasis governed by cell type-specific and branched feedback loops acting on NMD. Mol. Cell 43, 950–961 (2011).
Purcell, R. H., Purcell, R. H., Toro, C., Gahl, W. A. & Hall, R. A. A disease-associated mutation in the adhesion GPCR BAI2 (ADGRB2) increases receptor signaling activity. Hum. Mutat. 38, 1751–1760 (2017).
Van Es, M. A. et al. Genetic variation in DPP6 is associated with susceptibility to amyotrophic lateral sclerosis. Nat. Genet. 40, 29–31 (2008).
Spillantini, M. G. & Goedert, M. Tau pathology and neurodegeneration. Lancet Neurol. 12, 609–622 (2013).
Puffenberger, E. G. et al. A homozygous missense mutation in HERC2 associated with global developmental delay and autism spectrum disorder. Hum. Mutat. 33, 1639–1646 (2012).
Irimia, M. et al. A highly conserved program of neuronal microexons is misregulated in autistic brains. Cell 159, 1511–1523 (2014).
Ascano, M. et al. FMRP targets distinct mRNA sequence elements to regulate protein expression. Nature 492, 382–386 (2012).
Yamashita, A., Ohnishi, T., Kashima, I., Taya, Y. & Ohno, S. Human SMG-1, a novel phosphatidylinositol 3-kinase-related protein kinase, associates with components of the mRNA surveillance complex and is involved in the regulation of nonsense-mediated mRNA decay. Genes Dev. 15, 2215–2228 (2001).
Herder, C., Swiercz, J. M., Müller, C., Peravali, R. & Quiring, R. ArhGEF18 regulates RhoA–Rock2 signaling to maintain neuro-epithelial apico-basal polarity and proliferation. Development 2797, 2787–2797 (2013).
Hunt, D., Raivich, G. & Anderson, P. N. Activating transcription factor 3 and the nervous system. Front. Mol. Neurosci. 5, 7 (2012).
Durand, S., Franks, T. M. & Lykke-Andersen, J. Hyperphosphorylation amplifies UPF1 activity to resolve stalls in nonsense-mediated mRNA decay. Nat. Commun. 7, 12434 (2016).
Tani, H. et al. Identification of hundreds of novel UPF1 target transcripts by direct determination of whole transcriptome stability. RNA Biol. 9, 1370–1379 (2012).
Yamada, T. & Akimitsu, N. Contributions of regulated transcription and mRNA decay to the dynamics of gene expression. Wiley Interdiscip. Rev. RNA 10, e1508 (2019).
Mendell, J. T., Sharifi, N. A., Meyers, J. L., Martinez-Murillo, F. & Dietz, H. C. Nonsense surveillance regulates expression of diverse classes of mammalian transcripts and mutes genomic noise. Nat. Genet. 36, 1073–1078 (2004).
Lykke-Andersen, S. et al. Human nonsense-mediated RNA decay initiates widely by endonucleolysis and targets snoRNA host genes. Genes Dev. 28, 2498–2517 (2014).
Lou, C.-H. et al. Nonsense-mediated RNA decay influences human embryonic stem cell fate. Stem Cell Rep. 6, 844–857 (2016).
Ran, F. A. et al. Genome engineering using the CRISPR–Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Ohnishi, T. et al. Phosphorylation of hUPF1 induces formation of mRNA surveillance complexes containing hSMG-5 and hSMG-7. Mol. Cell 12, 1187–1200 (2003).
Kashima, I. et al. Binding of a novel SMG-1–Upf1–eRF1–eRF3 complex (SURF) to the exon junction complex triggers Upf1 phosphorylation and nonsense-mediated mRNA decay. Genes Dev. 20, 355–367 (2006).
Isken, O. et al. Upf1 phosphorylation triggers translational repression during nonsense-mediated mRNA decay. Cell 133, 314–327 (2008).
Lee, S. R., Pratt, G. A., Martinez, F. J., Yeo, G. W. & Lykke-Andersen, J. Target discrimination in nonsense-mediated mRNA decay requires Upf1 ATPase activity. Mol. Cell 59, 413–425 (2015).
Kurosaki, T., Miyoshi, K., Myers, J. R. & Maquat, L. E. NMD-degradome sequencing reveals ribosome-bound intermediates with 3′-end non-templated nucleotides. Nat. Struct. Mol. Biol. 25, 940–950 (2018).
Liu, B. et al. Regulatory discrimination of mRNAs by FMRP controls mouse adult neural stem cell differentiation. Proc. Natl Acad. Sci. USA 115, E11397–E11405 (2018).
Xie, N. et al. Reactivation of FMR1 by CRISPR/Cas9-mediated deletion of the expanded CGG-repeat of the fragile X chromosome. PLoS ONE 11, e0165499 (2016).
Lou, C. H. et al. Posttranscriptional control of the stem cell and neurogenic programs by the nonsense-mediated RNA decay pathway. Cell Rep. 6, 748–764 (2014).
Zhang, Y. et al. Rapid single-step induction of functional neurons from human pluripotent stem cells. Neuron 78, 785–798 (2013).
Sarnat, H. B. Clinical neuropathology practice guide 5-2013: markers of neuronal maturation. Clin. Neuropathol. 32, 340–369 (2013).
Castren, M. et al. Altered differentiation of neural stem cells in fragile X syndrome. Proc. Natl Acad. Sci. USA 102, 17834–17839 (2005).
Sheridan, S. D. et al. Epigenetic characterization of the FMR1 gene and aberrant neurodevelopment in human induced pluripotent stem cell models of fragile X syndrome. PLoS ONE 6, e26203 (2011).
Telias, M., Segal, M. & Ben-Yosef, D. Neural differentiation of fragile X human embryonic stem cells reveals abnormal patterns of development despite successful neurogenesis. Dev. Biol. 374, 32–45 (2013).
Doers, M. E. et al. iPSC-derived forebrain neurons from FXS individuals show defects in initial neurite outgrowth. Stem Cells Dev. 23, 1777–1787 (2014).
Halevy, T., Czech, C. & Benvenisty, N. Molecular mechanisms regulating the defects in fragile X syndrome neurons derived from human pluripotent stem cells. Stem Cell Rep. 4, 37–46 (2015).
Achuta, V. S. et al. Functional changes of AMPA responses in human induced pluripotent stem cell-derived neural progenitors in fragile X syndrome. Sci. Signal. 11, eaan8784 (2018).
Irwin, S. A. et al. Abnormal dendritic spine characteristics in the temporal and visual cortices of patients with fragile-X syndrome: a quantitative examination. Am. J. Med. Genet. 98, 161–167 (2001).
Durand, S. et al. Inhibition of nonsense-mediated mRNA decay (NMD) by a new chemical molecule reveals the dynamic of NMD factors in P-bodies. J. Cell Biol. 178, 1145–1160 (2007).
Martin, L. et al. Identification and characterization of small molecules that inhibit nonsense-mediated RNA decay and suppress nonsense p53 mutations. Cancer Res. 74, 3104–3113 (2014).
Feng, D. et al. Increase of a group of PTC+ transcripts by curcumin through inhibition of the NMD pathway. Biochim. Biophys. Acta 1849, 1104–1115 (2015).
Kamelgarn, M. et al. ALS mutations of FUS suppress protein translation and disrupt the regulation of nonsense-mediated decay. Proc. Natl Acad. Sci. USA 115, E11904–E11913 (2018).
Jackson, K. L. et al. Preservation of forelimb function by UPF1 gene therapy in a rat model of TDP-43-induced motor paralysis. Gene Ther. 22, 20–28 (2015).
Barmada, S. et al. Amelioration of toxicity in neuronal models of amyotrophic lateral sclerosis by hUPF1. Proc. Natl Acad. Sci. USA 112, 7821–7826 (2015).
Sun, Y. et al. C9orf72 arginine-rich dipeptide repeats inhibit UPF1-mediated RNA decay via translational repression. Nat. Commun. 11, 3354 (2020).
Gantois, I., Popic, J., Khourosky, A. & Sonenberg, N. Metformin for treatment of fragile X syndrome and other neurological disorders. Annu. Rev. Med. 70, 167–181 (2019).
Warren, L. et al. Highly efficient reprogramming to pluripotency and directed differentiation of human cells with synthetic modified mRNA. Cell Stem Cell 7, 618–630 (2010).
Miyoshi, N. et al. Reprogramming of mouse and human cells to pluripotency using mature microRNAs. Cell Stem Cell 8, 633–638 (2011).
Mandal, P. K. & Rossi, D. J. Reprogramming human fibroblasts to pluripotency using modified mRNA. Nat. Protoc. 8, 568–582 (2013).
Chan, E. M. et al. Live cell imaging distinguishes bona fide human iPS cells from partially reprogrammed cells. Nat. Biotechnol. 27, 1033–1037 (2009).
Bock, C. et al. Reference maps of human ES and iPS cell variation enable high-throughput characterization of pluripotent cell lines. Cell 144, 439–452 (2011).
Hosoda, N., Kim, Y. K., Lejeune, F. & Maquat, L. E. CBP80 promotes interaction of Upf1 with Upf2 during nonsense-mediated mRNA decay in mammalian cells. Nat. Struct. Mol. Biol. 12, 893–901 (2005).
Lykke-Andersen, J., Shu, M. D. & Steitz, J. A. Human Upf proteins target an mRNA for nonsense-mediated decay when bound downstream of a termination codon. Cell 103, 1121–1131 (2000).
Elbarbary, R. A., Miyoshi, K., Hedaya, O., Myers, J. R. & Maquat, L. E. UPF1 helicase promotes TSN-mediated miRNA decay. Gene. Dev. 39, 1483–1493 (2017).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. Sequence analysis featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2009).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Santos, A. et al. Comprehensive comparison of large-scale tissue expression datasets. Peer J. 3, e1054 (2015).
Mi, H., Muruganujan, A. & Thomas, P. D. PANTHER in 2013: modeling the evolution of gene function, and other gene attributes, in the context of phylogenetic trees. Nucleic Acids Res. 41, 377–386 (2013).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 45, D353–D361 (2017).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Tani, H. & Akimitsu, N. Genome-wide technology for determining RNA stability in mammalian cells: historical perspective and recent advantages based on modified nucleotide labeling. RNA Biol. 9, 37–41 (2012).
Dölken, L. High resolution gene expression profiling of RNA synthesis, processing, and decay by metabolic labeling of newly transcribed RNA using 4-thiouridine. Methods Mol. Biol. 1064, 91–100 (2013).
Trapnell, C. et al. Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 516–520 (2010).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Imamachi, N. et al. BRIC-seq: a genome-wide approach for determining RNA stability in mammalian cells. Methods 67, 55–63 (2014).
Acknowledgements
We thank M. Ascano for the pFRT/TO/Flag-HA-FMRP wild type, J. Lykke-Andersen for pcDNA3-Gl Norm-MS2bs and pcDNA3-Gl Ter-MS2bs, A. Paciorkowski for help generating iPSC lines 5417 and 5433, M. Lacagnina and N. Freitag for help maintaining the iPSCs, P. Jin for the FXS iPSC line and its isogenic control, F. Lejeune and D. Bedwell for helpful advice on the NMD inhibitors, S. Velu for synthesizing NMDI-1, M. Hoque (Genomics Center, Rutgers New Jersey Medical School) and the University of Rochester Genomic Research Center for generating libraries and performing RNA-seq and RIP-seq footprinting, K. Kawata for RNA-seq and RIP-seq data deposition, and M. Popp, X. Rambout, H. Sakano and J. Darnell for comments on the manuscript. Liquid chromatography/mass spectrometry analyses were performed by E. Spooner at the MIT Biopolymers and Proteomics Core Facility. This work was supported by R01 GM05696 (to L.E.M.) and MEXT KAKENHI (221S0002 to N.A.). C.P. was partially supported by 1R21NS104878 and Link Foundation. T.K. was partially supported by a post-doctoral fellowship from the FRAXA Research Foundation and funds from University of Rochester School of Medicine and Dentistry Pilot Funding in Stem Cell and Regenerative Medicine.
Author information
Authors and Affiliations
Contributions
T.K. and L.E.M. conceived of the project. T.K. performed all of the wet-bench experiments with technical help from R.N., except for the TRIC-seq experiments and analyses, which were performed by N.A. and N.I. C.P. helped with the generation, characterization and neuronal differentiation of iPSCs. N.I., S.M. and T.K. performed the computational analyses. T.K. and L.E.M. wrote the manuscript and the other authors edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Cell Biology thanks Miles Wilkinson and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Identifying SH-SY5Y NMD targets and their possible contributions to neurological diseases.
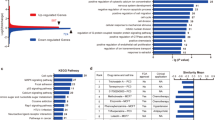
a, Schematic of transcriptome-wide approaches used to define NMD targets in human SH-SY5Y neuroblastoma cells: RNA-seq to quantitate mRNAs upregulated by UPF1 siRNA relative to Ctl siRNA (left), and RIP-seq footprinting (right) to identify mRNAs enriched in anti(α)-p-UPF1 IP relative to rabbit (r)IgG IP after increasing cellular p-UPF1 abundance using okadaic acid. PAGE, polyacrylamide gel electrophoresis. b, Representative western blots showing downregulation of UPF1 (that is UPF1-KD) relative to Ctl siRNA in samples used for RNA-seq. Here and elsewhere, left-most lanes under the wedge analyze serial 3-fold dilutions of lysate, and results represent 3 independent biological replicates. c, Western blots of lysates of SH-SY5Y cells cultured with okadaic acid for 2 hr to increase the cellular abundance of p-UPF1 before (−) or after IP using anti-p-UPF1 S1116 or rIgG. Results are representative of 3 independent biological replicates. d, As in b, but using UPF3X siRNA. e, Scatter plot of the log2 fold-change in mRNA abundance in the presence of UPF1 siRNA normalized to the abundance in the presence of Ctl siRNA (x-axis) and UPF3X siRNA normalized to the abundance of Ctl siRNA (y-axis). r and P values were calculated using Pearson’s correlation coefficient test. f, Cumulative fraction of log2 fold-change in mRNA abundance upon UPF3X-KD relative to Ctl-KD of “All mRNAs” (black line) or neuronal NMD targets defined in Fig. 1a (red line). P values were calculated using the two-sided Wilcoxon rank-sum test. g, Number and localization in brain of human NMD targets identified in this study. h, List of 22 proteins encoded by human neuronal-cell NMD targets that function in axon guidance. i, Diagram of proteins encoded by human neuronal-cell NMD targets that function in synaptic signaling (pink). Statistical source data and unprocessed blots are provided in Source Data Extended Data Fig. 1.
Extended Data Fig. 2 Confirmation of SH-SY5Y NMD targets and their change in abundance in neurological disease perturbation analyses.
a, As in Fig. 1e, but analyzing NMD targets that encode NMD factors or ATF3. Results derive from 3 independent biological replicates. b, Histogram representing disease perturbation analysis, illustrating that NMD targets are more often downregulated than upregulated in neurodevelopmental, psychiatric or neurodegenerative diseases (red). P values are calculated using the two-sided Fisher’s exact test. Statistical source data are provided in Source Data Extended Data Fig. 2.
Extended Data Fig. 3 Liquid chromatography-tandem mass spectrometry, p-UPF1 preferentially co-immunoprecipitates with NMD targets, and a ~3-fold increased SMG1 level typifying FMRP-KD cells recapitulates hyperactivated NMD.
a, Silver-stained polyacrylamide gel of lysates of HEK293T cells eluted from beads alone (beads-bound), or beads bound by rIgG- or anti-p-UPF1 (α-p-UPF1) used in LC-MS/MS. b, Proteins in anti-p-UFP1 IP identified using LC-MS/MS and ranked (x-axis) by the number of unique mass spectra (y-axis). FMRP and its FXR paralogs, red; proteins that function in NMD, blue. See also Supplementary Table 2. c, Coomassie Blue staining of human FMRP purified from E. coli (left) or FLAG-tagged human UPF1 purified from baculovirus (right). d, Quantitations of mRNAs, normalized to the level of ß-actin mRNA. Means with S.D., n = 3 independent biological replicates. (*)P < 0.05, (**)P < 0.01 or (***)P < 0.001 pertains to comparisons to Ctl-siRNA samples (two-sided t-test). e, Western blots of lysates of HEK293T cells transiently transfected with the denoted siRNA, eluted before (−) or after α-UPF1 or NRS IP. f, Quantitations of the specified RNA, normalized to the level of ß-actin mRNA, using lysates from e. Means with S.D., n = 3 independent biological replicates. P values compare Ctl siRNA- vs. specified siRNA-treated samples (two-sided t-test). g, Pertaining to Fig. 2h, western blots of lysates of HEK293T cells transfected with the specified siRNA, before or after α-p-UPF1 or rIgG IP. h, Western blot of lysates of HEK293T cells transfected with pcDNA-HA or pcDNA-HA-SMG1, the latter to approximate the increased level of SMG1 in FMRP siRNA-transfected cells. i, Quantitations of mRNAs, normalized to the level of their pre-mRNA, using lysates from h. Means with S.D., n = 3 independent biological replicates. P values compare HA-SMG1- vs. HA-transfected samples (two-sided t-test). j, Western blot of lysates of HEK293T cells transfected with the specified siRNA eluted before or after α-UPF1 or NRS IP. Results in a, e, g, h, j are representative of 3 independent biological replicates. Statistical source data and unprocessed blots are provided in Source Data Extended Data Fig. 3.
Extended Data Fig. 4 Tethering MS2CP-tagged FMRP fails to promote UPF1 recruitment, and UPF1 downregulation reduces FMRP binding to NMD targets.
a, As in Fig. 2l, but tethering MS2CP-FMRP in place of MS2CP-UPF1. b, Western blotting as in Fig. 2m but pertaining to a. c, Histogram representation as in Fig. 2n but using lysates analyzed in b. Results are represented as means with S.D., where n = 3 independent biological replicates. P values were calculated in comparisons of MS2CP-FMRP vs. MS2CP-transfected samples (two-sided t-test). d, Western blot of lysates of HEK293T cells, treated with the specified siRNA, before or after IP using anti-FMRP or, as a negative control, mIgG. Results are representative of 3 independent biological replicates. e, Histogram representation of RT-qPCR quantitations of the specified mRNA, as in Extended Data Fig. 3f, using lysates analyzed in d. Results are represented as means with S.D., where n = 3 independent biological replicates. P values were calculated in comparisons of Ctl siRNA vs. UPF1#1 siRNA-transfected samples (two-sided t-test). Statistical source data and unprocessed blots are provided in Source Data Extended Data Fig. 4.
Extended Data Fig. 5 Verification of FMRP-KD in TRIC-seq experiments, generation of FMR1-KO SH-SY5Y cells, and demonstration that ~2-fold overexpression of UPF1 in SH-SY5Y cells recapitulates the hyperactivated NMD that typifies FMR1-KO SH-SY5Y cells.
a, Scheme used to pulse-label SH-SY5Y-cell transcripts with 4-thiouridine (TU), chase for the indicated times in fresh medium lacking TU, biotinylate purified TU RNA, and capture biotinylated RNAs using streptavidin beads followed by RT-qPCR quantitations. b, Quantification of FMR1 mRNA relative to GAPDH mRNA in SH-SY5Y cells transfected with the specified siRNA, where n = 2 independent biological replicates. c, Western blots of lysates analyzed in b. Results are representative of 3 independent biological replicates. d, Cumulative fraction of log2 fold-change in mRNA half-life in FMRP#1-KD (upper) or FMRP#2-KD (lower) relative to Ctl-KD of “All mRNAs” (dotted line) or neuronal NMD targets (solid line). P values were calculated using the two-sided Wilcoxon rank-sum test. e, As in Fig. 3d, but for 33 NMD targets previously defined for non-neuronal cells. f, Diagram of relevant FMR1 gene exons, showing the sequences (blue) in exon 4 or 8 targeted by each pair of guide RNAs and CRISPR-Cas9n. Nicked sites, arrowheads; deleted or inserted sequences (red). g, As in Fig. 3f, but normalizations were to GAPDH mRNA. Means with S.D., n = 3 independent biological replicates. (*) P < 0.05 or (**) P < 0.01 is relative to WT samples (two-sided t-test). h, Western blots of lysates of WT SH-SY5Y cells expressing FLAG or FLAG-UPF1, the latter to bring the cellular abundance of UPF1 to approximate the increased level of UPF1 in FMR1-KO cells. Results are representative of 3 independent biological replicates. i, Quantitations of the specified mRNAs, normalized to the level of their pre-mRNA, using lysates from h. Means with S.D.,where n = 3 independent biological replicates. P values compare FLAG-UPF1 vs. FLAG-transfected samples (two-sided t-test). Statistical source data and unprocessed blots are provided in Source Data Extended Data Fig. 5.
Extended Data Fig. 6 Transcriptome-wide analyses of UPF1 siRNA downregulation in WT or FMR1-KO SHSY5Y cells show that NMD targets are comparably upregulated, and demonstration that the frequency of FMRP and p-UPF1 footprints on NMD targets is comparable.
a, Western blots of lysates of SH-SY5Y cells transiently transfected with Ctl or UPF1#1 siRNA using wild-type (WT) or FMR1-KO SH-SY5Y-cell lysates. b, Cumulative fraction of log2 fold-change in mRNA abundance upon UPF1#1-KD relative to Ctl-KD of “All mRNAs” (dotted line) or neuronal NMD targets (solid line) using WT (blue line) or FMR1-KO (red line) SH-SY5Y cell lysates, analyzed in a. P values were calculated using the two-sided Wilcoxon signed-rank test. c, Western blots of lysates of SH-SY5Y cells before (−) or after immunoprecipitation (IP) of using anti-FMRP or mouse (m)IgG. Results are representative of 3 independent biological replicates in a and c. d, Scatter plot of the abundance of FMRP footprints normalized to Input RNAs (x-axis) and p-UPF1 footprints normalized to Input RNAs (y-axis) on the specified regions (5′UTRs, CDSs and 3′UTRs) of NMD targets. r and P values were calculated using Pearson’s correlation coefficient test. Statistical source data and unprocessed blots are provided in Source Data Extended Data Fig. 6.
Extended Data Fig. 7 NMD is hyperactivated in FXS lymphoblasts, and characterization of iPSCs derived from healthy or FXS cells.
a, As in Fig. 4a, dot plots of RT-qPCR quantitations of the specified mRNA, normalized to the level of its pre-mRNA, using lysates from healthy (i.e. normal) or FXS lymphoblasts. Red bars represent means with S.D, where n = 6 independent biological cell lines. P values were calculated in comparisons of normal vs. FXS samples (two-sided Wilcoxon rank-sum test). b, as in a, but normalized to the level of ß-actin mRNA. Red bars, means with S.D., n = 6 independent biological cell lines. P values were calculated in comparisons of normal vs. FXS samples (two-sided Wilcoxon rank-sum test). c, Bright-field and IF images of iPSCs from a representative normal or a representative FXS patient-derived cell line for validation purposes. For IF, FXS iPSCs were stained for TRA-1-60 (green) or OCT4 (red), each of which is a pluripotent marker. Scale bar, 200 μm. Results are representative of 3 independent biological replicates. d, as in Fig. 4b, but normalized to the level of ß-actin mRNA. Means with S.D., where n = 3 biologically independent replicates. (***) P < 0.001 are relative to Normal #1 samples (one-way ANOVA and Dunnett’s multiple comparison test). e, As in Fig. 4b but for isogenic iPSCs + or − CGG repeat expansion. Means with S.D., where n = 3 independent biological replicates. P values were calculated in comparisons of + vs. − CGG repeat expansion (two-sided t-test). f, Western blots of lysates of normal and FXS iPSCs from Fig. 4b. Results are representative of 3 independent biological replicates. g, Protocol of neural differentiation – or + an NMD inhibitor. h, Quantitations of western blots shown in Fig. 4c. i, FXS iPSC-derived neurons exhibit deficient neurite formation on day 7 of differentiation. βIII-tubulin staining of normal and FXS neurons derived from iPSCs was used to assess neurite formation, average neurite length, neurite ramification, and neuronal differentiation efficacy. Means with S.D., where n = 8 independent biological replicates. P values were calculated in comparisons of normal vs. FXS samples (two-sided t-test). j, Quantitations of IF shown in Fig. 4d. Results are means with S.D., where n = 5 independent biological replicates. P values were calculated in comparisons of normal vs. FXS samples (two-sided t-test). Statistical source data and unprocessed blots are provided in Source Data Extended Data Fig. 7.
Extended Data Fig. 8 RT-qPCR and IF demonstrating that NMD inhibitors partially normalize FXS-derived iPSC differentiation.
a, Quantitations demonstrating that NMD was inhibited by each of the 3 NMD inhibitors. Samples derived from day 0, that is 24-hr after culturing iPSCs in the presence of each inhibitor as shown in Fig. 4e, b, Results are means with S.D., where n = 3 independent biological replicates. (*) P < 0.05 or (**) P < 0.01 is relative to RNA samples without NMD inhibitor (two-sided t-test). b, As in Extended Data Fig. 7j, but with or without an NMD inhibitor. Means with S.D., where n = 5 independent biological replicates. (*) P < 0.05, (**) P < 0.01 or (***) P < 0.01 is relative to RNA samples without an NMD inhibitor (two-sided t-test). c, As in Fig. 4e, but staining for MAP2 (red). d, As in Fig. 4e, but staining for BRN2/POU3F2 (green). Results are representative of 3 independent biological replicates in (c) and (d). e, Histogram representations of quantitations of western blots shown in Fig. 4f. Results represent n = 3 independent biological replicates, except for SYN1 analyses in FXS neurons, BRN2 analysis in FXS neurons treated with NMDI-1, and DCX analysis in FXS neurons treated with curcumin, where n = 2 independent biological replicates. f, Histogram representation of neurite outgrowth manifested by representative normal or FXS neurons on day 15 after differentiation. Results represent 2 independent biological replicates for FXS neurons and 3 independent biological replicates for normal neurons. All cells were stained for viability, and fluorescent signals derive from minimally 4 fields per well, where the extent of fluorescence for FXS neurons in the absence (−) of inhibitor is defined as 1. Statistical source data are provided in Source Data Extended Data Fig. 8.
Supplementary information
Supplementary Information
Supplementary Fig. 1.
Supplementary Tables 1–7
Supplementary Table 1: Human SH-SY5Y neuronal cell NMD targets. Supplementary Table 2: Liquid chromatography–tandem mass spectrometry identification of HEK293T cell proteins that co-immunoprecipitate with p-UPF1. Supplementary Table 3: FMRP siRNA-mediated mRNA half-life changes of human SH-SY5Y cell NMD targets. Supplementary Table 4: UPF1 siRNA-mediated changes in mRNA abundance in wild-type and FMR1 KO SH-SY5Y cells. Supplementary Table 5: Differentially expressed transcripts in neurons derived from FXS iPSCs with or without NMDI-1 treatment. Supplementary Table 6: Reagents and resources used in this study. Supplementary Table 7: Karyotype analysis of the iPSCs used in this study.
Source data
Source Data Fig. 1
Data quantitation and statistical source data.
Source Data Fig. 1
Unprocessed blots/gels.
Source Data Fig. 2
Data quantitation and statistical source data.
Source Data Fig. 2
Unprocessed blots/gels.
Source Data Fig. 3
Data quantitation and statistical source data.
Source Data Fig. 3
Unprocessed blots/gels.
Source Data Fig. 4
Data quantitation and statistical source data.
Source Data Fig. 4
Unprocessed blots/gels.
Source Data Extended Data Fig. 1
Data quantitation and statistical source data.
Source Data Extended Data Fig. 1
Unprocessed blots/gels.
Source Data Extended Data Fig. 2
Data quantitation and statistical source data.
Source Data Extended Data Fig. 3
Data quantitation and statistical source data.
Source Data Extended Data Fig. 3
Unprocessed blots/gels.
Source Data Extended Data Fig. 4
Data quantitation and statistical source data.
Source Data Extended Data Fig. 4
Unprocessed blots/gels.
Source Data Extended Data Fig. 5
Data quantitation and statistical source data.
Source Data Extended Data Fig. 5
Unprocessed blots/gels.
Source Data Extended Data Fig. 6
Data quantitation and statistical source data.
Source Data Extended Data Fig. 6
Unprocessed blots/gels.
Source Data Extended Data Fig. 7
Data quantitation and statistical source data.
Source Data Extended Data Fig. 7
Unprocessed blots/gels.
Source Data Extended Data Fig. 8
Data quantitation and statistical source data.
Rights and permissions
About this article
Cite this article
Kurosaki, T., Imamachi, N., Pröschel, C. et al. Loss of the fragile X syndrome protein FMRP results in misregulation of nonsense-mediated mRNA decay. Nat Cell Biol 23, 40–48 (2021). https://doi.org/10.1038/s41556-020-00618-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-00618-1
This article is cited by
-
FMRP-mediated spatial regulation of physiologic NMD targets in neuronal cells
Genome Biology (2024)
-
Fmr1 exon 14 skipping in late embryonic development of the rat forebrain
BMC Neuroscience (2022)
-
Sustained correction of hippocampal neurogenic and cognitive deficits after a brief treatment by Nutlin-3 in a mouse model of fragile X syndrome
BMC Medicine (2022)
-
NMD abnormalities during brain development in the Fmr1-knockout mouse model of fragile X syndrome
Genome Biology (2021)