Abstract
Many germ cells are eliminated during development, long before oogenesis or spermatogenesis. In mouse fetal testes, the majority of germ cell apoptosis coincides with the onset of male differentiation, suggesting coordination of these processes. We studied fetal germ-cell fates and discovered that both apoptosis and differentiation initiate in clonally related clusters. Lineage tracing confirmed that germ cells die as clones independent of intercellular bridges, suggesting that shared intrinsic properties are apoptotic determinants. We identified transcriptional heterogeneity among fetal germ cells that included an apoptosis-susceptible population characterized by failure to differentiate, whereas successful differentiation to prospermatogonia occurred through the expression of epigenetically regulated genes, including LINE1. Our results indicate that the fetal germ-cell fate is based on discrete cell-heritable identities. Elevated DNA methylation in the apoptosis-susceptible subpopulation supports our hypothesis that earlier errors in germ-cell epigenetic reprogramming derail differentiation in cellular progeny, leading to fetal apoptotic selection that ultimately improves the gamete quality.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The scRNA-seq and BS-seq data that support the findings of this study have been deposited in the Gene Expression Omnibus under the accession codes GSE119045 and GSE155122. All other data supporting the findings of this study are available from the corresponding author on reasonable request. Gene-ontology analysis on MSigDB computed overlaps with GO:BP (https://www.gsea-msigdb.org/gsea/msigdb/genesets.jsp?collection=GO:BP). Source data are provided with this paper.
Code availability
Code for the scRNA-seq analysis is available at https://github.com/dnucsf/NatureCellBiology2020.
References
Lehtonen, J. & Parker, G. A. Gamete competition, gamete limitation, and the evolution of the two sexes. Mol. Hum. Reprod. 20, 1161–1168 (2014).
Gardner, R. L. & Rossant, J. Investigation of the fate of 4–5 day post-coitum mouse inner cell mass cells by blastocyst injection. J. Embryol. Exp. Morphol. 52, 141–152 (1979).
Anderson, R., Copeland, T. K., Schöler, H., Heasman, J. & Wylie, C. The onset of germ cell migration in the mouse embryo. Mech. Dev. 91, 61–68 (2000).
Cantú, A. V., Altshuler-Keylin, S. & Laird, D. J. Discrete somatic niches coordinate proliferation and migration of primordial germ cells via Wnt signaling. J. Cell. Biol. 214, 215–229 (2016).
Saitou, M., Kagiwada, S. & Kurimoto, K. Epigenetic reprogramming in mouse pre-implantation development and primordial germ cells. Development 139, 15–31 (2012).
Menke, D. B., Koubova, J. & Page, D. C. Sexual differentiation of germ cells in XX mouse gonads occurs in an anterior-to-posterior wave. Dev. Biol. 262, 303–312 (2003).
Ohta, K. et al. Male differentiation of germ cells induced by embryonic age-specific Sertoli cells in mice. Biol. Reprod. 86, 112 (2012).
Aitken, R. J., Findlay, J. K., Hutt, K. J. & Kerr, J. B. Apoptosis in the germ line. Reproduction 141, 139–150 (2011).
Runyan, C. et al. Steel factor controls midline cell death of primordial germ cells and is essential for their normal proliferation and migration. Development 133, 4861–4869 (2006).
Knudson, C. M., Tung, K. S., Tourtellotte, W. G., Brown, G. A. & Korsmeyer, S. J. Bax-deficient mice with lymphoid hyperplasia and male germ cell death. Science 270, 96–99 (1995).
Rodriguez, I., Ody, C., Araki, K., Garcia, I. & Vassalli, P. An early and massive wave of germinal cell apoptosis is required for the development of functional spermatogenesis. EMBO J. 16, 2262–2270 (1997).
Perez, G. I., Knudson, C. M., Leykin, L., Korsmeyer, S. J. & Tilly, J. L. Apoptosis-associated signaling pathways are required for chemotherapy-mediated female germ cell destruction. Nat. Med. 3, 1228–1232 (1997).
Coucouvanis, E. C., Sherwood, S. W., Carswell-Crumpton, C., Spack, E. G. & Jones, P. P. Evidence that the mechanism of prenatal germ cell death in the mouse is apoptosis. Exp. Cell Res. 209, 238–247 (1993).
Wang, R. A., Nakane, P. K. & Koji, T. Autonomous cell death of mouse male germ cells during fetal and postnatal period. Biol. Reprod. 58, 1250–1256 (1998).
Goriely, A., McVean, G. A. T., Röjmyr, M., Ingemarsson, B. & Wilkie, A. O. M. Evidence for selective advantage of pathogenic FGFR2 mutations in the male germ line. Science 301, 643–646 (2003).
Ueno, H., Turnbull, B. B. & Weissman, I. L. Two-step oligoclonal development of male germ cells. Proc. Natl Acad. Sci. USA 106, 175–180 (2009).
Cook, M. S., Munger, S. C., Nadeau, J. H. & Capel, B. Regulation of male germ cell cycle arrest and differentiation by DND1 is modulated by genetic background. Development 138, 23–32 (2011).
Foley, J. G. D. & Bard, J. B. L. Apoptosis in the cortex of the developing mouse kidney. J. Anat. 201, 477–484 (2002).
Laird, D. J., Altshuler-Keylin, S., Kissner, M. D., Zhou, X. & Anderson, K. V. Ror2 enhances polarity and directional migration of primordial germ cells. PLOS Genet. 7, e1002428 (2011).
Mendis, S. H. S., Meachem, S. J., Sarraj, M. A. & Loveland, K. L. Activin A balances Sertoli and germ cell proliferation in the fetal mouse testis. Biol. Reprod. 84, 379–391 (2011).
Lei, L. & Spradling, A. C. Mouse primordial germ cells produce cysts that partially fragment prior to meiosis. Development 140, 2075–2081 (2013).
Ventelä, S., Toppari, J. & Parvinen, M. Intercellular organelle traffic through cytoplasmic bridges in early spermatids of the rat: mechanisms of haploid gene product sharing. Mol. Biol. Cell 14, 2768–2780 (2003).
Greenbaum, M. P. et al. TEX14 is essential for intercellular bridges and fertility in male mice. Proc. Natl Acad. Sci. USA 103, 4982–4987 (2006).
Lu, K. L. & Yamashita, Y. M. Germ cell connectivity enhances cell death in response to DNA damage in the Drosophila testis. eLife 6, e27960 (2017).
Snippert, H. J. et al. Intestinal crypt homeostasis results from neutral competition between symmetrically dividing Lgr5 stem cells. Cell 143, 134–144 (2010).
Rinkevich, Y., Lindau, P., Ueno, H., Longaker, M. T. & Weissman, I. L. Germ-layer and lineage-restricted stem/progenitors regenerate the mouse digit tip. Nature 476, 409–413 (2011).
Greder, L. V. et al. Analysis of endogenous Oct4 activation during induced pluripotent stem cell reprogramming using an inducible Oct4 lineage label. Stem Cells 30, 2596–2601 (2012).
Western, P. S., Miles, D. C., van den Bergen, J. A., Burton, M. & Sinclair, A. H. Dynamic regulation of mitotic arrest in fetal male germ cells. Stem Cells 26, 339–347 (2008).
Bondar, T. & Medzhitov, R. p53-Mediated hematopoietic stem and progenitor cell competition. Cell Stem Cell 6, 309–322 (2010).
Bowling, S. et al. P53 and mTOR signalling determine fitness selection through cell competition during early mouse embryonic development. Nat. Commun. 9, 1763 (2018).
Haupt, S., Berger, M., Goldberg, Z. & Haupt, Y. Apoptosis—the p53 network. J. Cell Sci. 116, 4077–4085 (2003).
de Felici, M. D. et al. Bcl-2 and Bax regulation of apoptosis in germ cells during prenatal oogenesis in the mouse embryo. Cell Death Differ. 6, 908–915 (1999).
Rucker, E. B. et al. Bcl-x and Bax regulate mouse primordial germ cell survival and apoptosis during embryogenesis. Mol. Endocrinol. 14, 1038–1052 (2000).
Maclean, J. A. et al. Rhox: a new homeobox gene cluster. Cell 120, 369–382 (2005).
Spiller, C. M. et al. Endogenous Nodal signaling regulates germ cell potency during mammalian testis development. Development 139, 4123–4132 (2012).
Saba, R., Kato, Y. & Saga, Y. NANOS2 promotes male germ cell development independent of meiosis suppression. Dev. Biol. 385, 32–40 (2014).
Molaro, A. et al. Two waves of de novo methylation during mouse germ cell development. Genes Dev. 28, 1544–1549 (2014).
Suzuki, A. & Saga, Y. Nanos2 suppresses meiosis and promotes male germ cell differentiation. Genes Dev. 22, 430–435 (2008).
Milholland, B. et al. Differences between germline and somatic mutation rates in humans and mice. Nat. Commun. 8, 15183 (2017).
Seisenberger, S. et al. The dynamics of genome-wide DNA methylation reprogramming in mouse primordial germ cells. Mol. Cell 48, 849–862 (2012).
Hill, P. W. S. et al. Epigenetic reprogramming enables the transition from primordial germ cell to gonocyte. Nature 555, 392–396 (2018).
Hargan-Calvopina, J. et al. Stage-specific demethylation in primordial germ cells safeguards against precocious differentiation. Dev. Cell 39, 75–86 (2016).
Hajkova, P. et al. Epigenetic reprogramming in mouse primordial germ cells. Mech. Dev. 117, 15–23 (2002).
Malki, S., van der Heijden, G. W., O’Donnell, K. A., Martin, S. L. & Bortvin, A. A role for retrotransposon LINE-1 in fetal oocyte attrition in mice. Dev. Cell 29, 521–533 (2014).
Reznik, B. et al. Heterogeneity of transposon expression and activation of the repressive network in human fetal germ cells. Development 146, dev.171157 (2019).
Brockway, N. L. et al. Multicolour lineage tracing using in vivo time-lapse imaging reveals coordinated death of clonally related cells in the developing vertebrate brain. Dev. Biol. 453, 130–140 (2019).
Watanabe, K. et al. Cripto-1 is a cell surface marker for a tumorigenic, undifferentiated subpopulation in human embryonal carcinoma cells. Stem Cells 28, 1303–1314 (2010).
Spiller, C. M., Bowles, J. & Koopman, P. Nodal/Cripto signaling in fetal male germ cell development: implications for testicular germ cell tumors. Int. J. Dev. Biol. 57, 211–219 (2013).
Dawson, E. P. et al. Delayed male germ cell sex-specification permits transition into embryonal carcinoma cells with features of primed pluripotency. Development 145, dev156612 (2018).
Aravin, A. A. et al. A piRNA pathway primed by individual transposons is linked to de novo DNA methylation in mice. Mol. Cell 31, 785–799 (2008).
Hansson K. et al. RipleyGUI: software for analyzing spatial patterns in 3D cell distributions. Front. Neuroinform. 7, 5 (2013).
Susaki, E. A. et al. Whole-brain imaging with single-cell resolution using chemical cocktails and computational analysis. Cell 157, 726–739 (2014).
Jin, Y. et al. TEtranscripts: a package for including transposable elements in differential expression analysis of RNA-seq datasets. Bioinformatics 31, 3593–3599 (2015).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Subramanaian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Acknowledgements
Mice were provided by I.L. Weissman, M. Matzuk, and M. Bartolomei. ORF1p antibody was provided by A. Wilkin, A. Bortvin, and A. Wiblin. J. Sneddon, M. Conti, T. Nystul, A.W. De Tomaso, L. Byrnes, D. Wong, R.G. Jaszczak, S. Cincotta, and B. Reznik provided feedback. J. Daza, G. Zhang, A. Villanueva, D. Palomera, I. Vainchtein, C. Bispo, R. Prasasva, and B. Dreskin assisted with experiments. Funding included an NSF Predoctoral Fellowship to D.H.N. and NIH 1DP2OD007420, R01GM122902, R01ES028212, and Cancer Research Coordinating Committee grants to D.J.L.
Author information
Authors and Affiliations
Contributions
D.H.N. and D.J.L. conceived of the study, analysed the data and wrote the manuscript. D.H.N., B.S., S.P., S.M. and G.H. conducted experiments and analysed results. All authors contributed comments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Apoptotic clusters are randomly distributed throughout fetal testes.
a, Percent apoptotic cleaved-PARP+ germ cells in whole-mount testes on a mixed genetic background during the apoptotic wave. n=4 animals. Data presented as mean values ± SD. Dashed line denotes reported average apoptotic index in developing fetal tissues18. b, Percent cleaved-PARP+ cells on apoptosis-sensitive C57/Bl6 background20 at peak apoptosis (E13.5). c, Distribution of apoptotic clusters measured for spatial bias. 3D coordinates of each cluster’s centroid were evaluated by Ripley’s K function. These were compared against simulated distributions for an equivalent number of points. Representative plots depict each simulated distribution type (right). Cluster coordinates from n=5 animals.**: p=0, †: p=0.10622, between-treatments sum of squares. d, E13.5 testis section stained for Sertoli cells (AMH), germ cells (TRA98), and apoptosis (cleaved-PARP). Apoptotic cords are defined as cords containing any apoptotic germ cells (red outline) in contrast to non-apoptotic cords (white outline). Scale bar = 20µm. e, Ratio of total germ cells to Sertoli cells in apoptotic and non-apoptotic cords, p=0.584, two-tailed Student’s t-test. Data points represent individual cords pooled from sections from 4 animals. n =15 non-apoptotic cords and n = 11 apoptotic cords. Data presented as mean values ± SD. f, Average numbers of Sertoli cells in direct contact with each germ cell in apoptotic and non-apoptotic cords. Within apoptotic cords, apoptotic (PARP+) and non-apoptotic (PARP-) germ cells were separately considered. p=0.737154, one-way ANOVA. Data points represent individual germ cells (n) analysed in sections from 4 animals. Total cell counts per category are detailed in source data. Data presented as mean values ± SD. g, Quantification of apoptotic germ cells in E13.5 Tex14 mutants. p=0.4491, two-tailed Student’s t-test. n=4 animals per genotype. Data presented as mean values ± SD. h, Apoptotic cluster size by cell number in Tex14 E13.5. p=0.0073, two-tailed Student’s t-test. Data points (n) are available in source data and represent individual apoptotic clusters analysed from 2 animals (Tex14+/+) and 3 animals (Tex14-/-). Data presented as mean values ± SD.
Extended Data Fig. 2 Clonal boundary determination for fragmented apoptotic clusters.
a, Detection of clonal apoptosis in Confetti-labelled germ cells at E13.5. Apoptotic germ cells identified by cleaved-PARP staining (white outlines). Scale bar = 20um. 5 animals. Inset: Hoechst staining. b, Confetti -labelled germ cells imaged in whole-mount and segmented using the Imaris Cell module displayed fragmentation by E13.5, in accordance with published findings21. We identified three-dimensional coordinates for centroids (grey dots) of each cell (represented by yellow spheres) within a clone to measure the centroid-to-centroid distances between nearest-neighbour germ cells belonging to different fragments. In depicted clone, \(\overline {BC}\) spans nearest-neighbour cells between Fragments 1 and 2 while \(\overline {DE}\) spans nearest-neighbour cells from Fragments 3 and 2. We took the largest of these nearest-neighbour distances from all fragments pairs to represent the maximum potential dispersion between fragments of a single clone. Scale bar = 5um.
Extended Data Fig. 3 Clonal apoptotic clusters are maintained in the absence of intercellular bridges.
a, Cleaved-PARP staining for apoptosis in whole-mount E13.5 Tex14-/- testes on a R26R Rainbow clonally labelled background reveals mCherry apoptotic clone (outlined in yellow). Scale bar = 10µm. Rotated view of same clones confirming that constituent cells of mCherry clone are disconnected. Nearby mCerulean clone is negative for apoptosis. b, Extended view of entire whole-mount E13.5 testis. Scale bar = 100µm.
Extended Data Fig. 4 Single-cell-RNAseq analysis of E13.5 male germ cells.
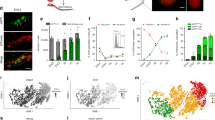
a, Average expression of p53-activated genes by germ cell subpopulation. Expression from n=2,556 single cells. b, Subpopulation size in cell number and as a percentage of total germ cells analysed for scRNA-seq at E13.5. c, Average pro-survival gene Nanos3 expression by germ cell subpopulation. Expression from n=2,556 single cells. d, Average expression of prosurvival genes Kit, adhesion factor Epcam, and PGC identity gene Ifitm3 by germ cell subpopulation. Expression from n=2,556 single cells.
Extended Data Fig. 5 LEFTY1/2+ cells are increased in p53-/- testes by E13.5 compared to p53+/-.
a, Immunofluorescence detection of LEFTY1/2 in sections from E13.5 p53+/- and p53-/- testes. Scale bar = 50µm. 2 animals per genotype. b, Quantification of percent LEFTY1/2+ germ cells from E13.5 p53+/- and p53-/- testes. n=2 animals per genotype.
Extended Data Fig. 6 E12.5 and E13.5 germ cells remain transcriptionally distinct in merged analysis.
a, Merged dataset with E12.5 and E13.5 germ cells after transcriptional clustering. E13.5 AP-germ cell identities coloured while E12.5 boundary is denoted by dashed black line. t-SNE represents 5,106 cells. b, t-SNE plots showing expression of apoptotic genes (Bad, Bax), male differentiation genes (Nanos2, Mael), and genes associated with a pre-male differentiation population (Rhox6, Lefty2). Cell shading is derived from single-cell expression values normalized to global expression for all genes and log-transformed via Seurat.
Extended Data Fig. 7 Model for coordination of clonal apoptosis with male differentiation.
Clonal variation in differentiation potential results in divergent clonal fates: germ cells normally activate male differentiation programs marked by Nanos2 but a subset of clones aberrantly maintain an undifferentiated state characterized by expression of pro-apoptotic genes that ultimately results in clonal death.
Extended Data Fig. 8 LEFTY expression is clonally heterogeneous.
LEFTY expression in E13.5 Confetti testis, LEFTY-high cells (open arrowheads) and LEFTY-low cells (white arrowheads) with clone boundaries indicated by dashed outlines. Scale bar = 20µm.
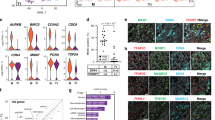
Extended Data Fig. 9 Population-specific expression of epigenetically regulated genes including LINE1 during male differentiation.
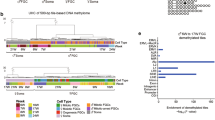
a, Whole-genome bisulfite sequencing comparison of sorted AP and MD populations for relative methylation percentage. Data are presented as median values with first and third quartiles. Whiskers represent minimum and maximums. Sequencing was performed on 2 separate runs of n=2,000 and n=6,000 cells each for both populations. b, ORF1p expression during the apoptotic wave. ORF1p is expressed in small clusters of germ cells at E13.5 (white arrowhead) and progresses toward homogenous expression by E15.5. Scale bar = 100µm. c, ORF1p and cleaved PARP staining in E13.5 wild-type sections. Scale bar = 25µm. d, ORF1p-negative germ cell clusters in E15.5 Bax-/- compared to Bax+/-. Arrowheads denote ORF1p-negative cells. Insets: Higher-magnification image of ORF1p-negative cluster compared to ORF1p-positive neighbouring germ cells. Scale bar = 50µm.
Extended Data Fig. 10 Differentially expressed repeat elements identify distinct subpopulations of E13.5 germ cells.
a, Abridged table of repeat element markers for subpopulations defined by scRNA-seq clustering based on expression of non-repetitive elements. p value calculated by two-sided Wilcoxon rank-sum test. Expression from n=2,043 cells. Full table of markers available in source data. b, Expression of MD marker Nanos2, AP marker Rhox6, and LINE1 members across clusters identified by expression of non-repetitive elements. Predominant expression of Nanos2 in cluster 4 is characteristic of MD population. Predominant expression of AP marker Rhox6 in cluster 5 is characteristic of AP population. Violin plots display cell densities as shape and individual cells as dots. The number of cells (n) for each cluster are detailed in source data. c, Abridged table of repeat element markers for subpopulations defined by scRNA-seq based only on expression of repetitive elements. P value calculated by two-sided Wilcoxon rank-sum test. Expression from n=2,043 cells. Full table of markers available in source data. d, Expression of MD marker Nanos2, AP marker Rhox6, and LINE1 members across clusters identified only by expression of repeat elements. Violin plots display cell densities as shape and individual cells as dots. The number of cells (n) for each cluster are detailed in source data.
Supplementary information
Supplementary Tables 1 and 2
Supplementary Table 1. Transcriptional markers that distinguish each germ-cell state compared with all others were identified among differentially expressed genes in an E13.5 germ-cell population using the two-tailed Wilcoxon rank-sum test-based FindMarkers operation in Seurat, which provides a P value that was further adjusted using Bonferroni’s correction using all genes in the dataset. Supplementary Table 2. Comparison of the markers for MD germ cells with GRRs and genes upregulated in Dnmt1-cKO male germ cells42. Comparison of markers for AP germ cells with genes upregulated in Tet1-KO41 male germ cells and genes downregulated in Dnmt1 cKO. Adjusted P values for the MD and AP markers were generated using the Wilcoxon rank-sum test with Bonferroni’s correction. Statistics for markers from Dnmt1 cKO and Tet1-KO were obtained from referenced studies.
41556_2020_603_MOESM3_ESM.mp4
Supplementary Video 1. Whole-mount visualization of apoptotic clusters of PGCs at E13.5. Whole E13.5 testis with cell nuclei identified by Hoechst and PGCs by TRA98. Apoptosis was identified with cPARP and rotated views highlight the spatial distribution of apoptotic PGCs. The magnified views show a representative apoptotic germ cell cluster.
41556_2020_603_MOESM4_ESM.mp4
Supplementary Video 2. PGC proliferation is clonal. E13.5 PGCs were clonally labelled using Rainbow and cells in the late G2/M phase were marked by phospho-histone H3. The magnified view shows two separate groups of phospho-histone H3+ cells that are clonal (mOrange clone and a GFP− clone).
41556_2020_603_MOESM5_ESM.mov
Supplementary Video 3. Multicolour clone labelling reveals shared fluorophores within germ-cell cysts. Three-dimensional perspective of whole E13.5 Rainbow; Tex14+/+; Pou5f1-Cre-ER testis with a bicoloured clone (mCerulean and mCherry) outlined as a surface, followed by individual channels confirming bicoloured labelling.
41556_2020_603_MOESM6_ESM.mp4
Supplementary Video 4. Tex14-mutant testes differ in clone structure and do not form bicoloured clones. Three-dimensional perspective of whole E13.5 Rainbow; Tex14−/−; Pou5f1-Cre-ER testis with strictly single-coloured clones as a consequence of the absence of intercellular bridges.
Source data
Source Data Fig. 1
Statistical source data for Fig. 1b
Source Data Fig. 2
Statistical source data for Fig. 2b
Source Data Fig. 3
Statistical source data for Fig. 3d,e
Source Data Fig. 4
Statistical source data for Fig. 4b,d
Source Data Fig. 6
Statistical source data for Fig. 6a–c
Source Data Fig. 7
Statistical source data for Fig. 7c–e
Source Data Fig. 8
Statistical source data for Fig. 8b,d,e
Source Data Extended Data Fig. 1
Statistical source data for Extended Data Fig. 1c,e–h
Source Data Extended Data Fig. 4
Statistical source data for Extended Data Fig. 4a,c,d
Source Data Extended Data Fig. 5
Statistical source data for Extended Data Fig. 5b
Source Data Extended Data Fig. 9
Statistical source data for Extended Data Fig. 9a
Source Data Extended Data Fig. 10
Statistical source data for Extended Data Fig. 10a–d
Rights and permissions
About this article
Cite this article
Nguyen, D.H., Soygur, B., Peng, SP. et al. Apoptosis in the fetal testis eliminates developmentally defective germ cell clones. Nat Cell Biol 22, 1423–1435 (2020). https://doi.org/10.1038/s41556-020-00603-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-00603-8
This article is cited by
-
MAX controls meiotic entry in sexually undifferentiated germ cells
Scientific Reports (2024)
-
Apoptotic cell death in disease—Current understanding of the NCCD 2023
Cell Death & Differentiation (2023)
-
Single-cell RNA sequencing of mitotic-arrested prospermatogonia with DAZL::GFP chickens and revealing unique epigenetic reprogramming of chickens
Journal of Animal Science and Biotechnology (2022)
-
SLC38A4 functions as a tumour suppressor in hepatocellular carcinoma through modulating Wnt/β-catenin/MYC/HMGCS2 axis
British Journal of Cancer (2021)
-
Natural selection at the cellular level: insights from male germ cell differentiation
Cell Death & Differentiation (2021)