Abstract
In the unicellular eukaryote Saccharomyces cerevisiae, Cln3–cyclin-dependent kinase activity enables Start, the irreversible commitment to the cell division cycle. However, the concentration of Cln3 has been paradoxically considered to remain constant during G1, due to the presumed scaling of its production rate with cell size dynamics. Measuring metabolic and biosynthetic activity during cell cycle progression in single cells, we found that cells exhibit pulses in their protein production rate. Rather than scaling with cell size dynamics, these pulses follow the intrinsic metabolic dynamics, peaking around Start. Using a viral-based bicistronic construct and targeted proteomics to measure Cln3 at the single-cell and population levels, we show that the differential scaling between protein production and cell size leads to a temporal increase in Cln3 concentration, and passage through Start. This differential scaling causes Start in both daughter and mother cells across growth conditions. Thus, uncoupling between two fundamental physiological parameters drives cell cycle commitment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Source data are available online for Figs. 1–6 and Extented Data Figs. 1–8. The mass spectrometry proteomics data have been deposited in the ProteomeXchange Consortium PRIDE81 partner repository with the dataset identifier PXD015327. All other data are available from the authors on reasonable request.
Code availability
At https://github.com/molecular-systems-biology/Litsios-et-al-2019, we provide one CSV file containing raw microscopy data, together with the respective MATLAB file as an example of our data-processing pipeline (smoothing, rate estimation and so on). These data were used in the construction of Fig. 5c and Extended Data Fig. 4h. The custom-made Python script used for analysis of the confocal images is also provided. All other MATLAB scripts used for processing are available from the authors on reasonable request.
References
Johnson, A. & Skotheim, J. M. Start and the restriction point. Curr. Opin. Cell Biol. 25, 717–723 (2013).
Nash, R., Tokiwa, G., Anand, S., Erickson, K. & Futcher, A. B. The WHI1 + gene of Saccharomyces cerevisiae tethers cell division to cell size and is a cyclin homolog. EMBO J. 7, 4335–4346 (1988).
Cross, F. R. DAF1, a mutant gene affecting size control, pheromone arrest, and cell cycle kinetics of Saccharomyces cerevisiae. Mol. Cell. Biol. 8, 4675–4684 (1988).
Tyers, M., Tokiwa, G. & Futcher, B. Comparison of the Saccharomyces cerevisiae G1 cyclins: Cln3 may be an upstream activator of Cln1, Cln2 and other cyclins. EMBO J. 12, 1955–1968 (1993).
Tyers, M., Tokiwa, G., Nash, R. & Futcher, B. The Cln3–Cdc28 kinase complex of S. cerevisiae is regulated by proteolysis and phosphorylation. EMBO J. 11, 1773–1784 (1992).
De Bruin, R. A. M., McDonald, W. H., Kalashnikova, T. I., Yates, J. & Wittenberg, C. Cln3 activates G1-specific transcription via phosphorylation of the SBF bound repressor Whi5. Cell 117, 887–898 (2004).
Costanzo, M. et al. CDK activity antagonizes Whi5, an inhibitor of G1/S transcription in yeast. Cell 117, 899–913 (2004).
Wang, H., Carey, L. B., Cai, Y., Wijnen, H. & Futcher, B. Recruitment of Cln3 cyclin to promoters controls cell cycle entry via histone deacetylase and other targets. PLoS Biol. 7, e1000189 (2009).
Skotheim, J. M., Di Talia, S., Siggia, E. D. & Cross, F. R. Positive feedback of G1 cyclins ensures coherent cell cycle entry. Nature 454, 291–296 (2008).
McInerny, C. J., Partridge, J. F., Mikesell, G. E., Creemer, D. P. & Breeden, L. L. A novel Mcm1-dependent element in the SWI4, CLN3, CDC6, and CDC47 promoters activates M/G1-specific transcription. Genes Dev. 11, 1277–1288 (1997).
Zapata, J. et al. PP2ARts1 is a master regulator of pathways that control cell size. J. Cell Biol. 204, 359–376 (2014).
Thorburn, R. R. et al. Aneuploid yeast strains exhibit defects in cell growth and passage through START. Mol. Biol. Cell 24, 1274–1289 (2013).
Schmoller, K. M., Turner, J. J., Kõivomägi, M. & Skotheim, J. M. Dilution of the cell cycle inhibitor Whi5 controls budding-yeast cell size. Nature 526, 268–272 (2015).
Jorgensen, P. & Tyers, M. How cells coordinate growth and division. Curr. Biol. 14, R1014–R1027 (2004).
Polymenis, M. & Schmidt, E. V. Coupling of cell division to cell growth by translational control of the G1 cyclin CLN3 in yeast. Genes Dev. 11, 2522–2531 (1997).
Schmoller, K. M. & Skotheim, J. M. The biosynthetic basis of cell size control. Trends Cell Biol. 25, 793–802 (2015).
Elliott, S. G. & McLaughlin, C. S. Rate of macromolecular synthesis through the cell cycle of the yeast Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 75, 4384–4388 (1978).
Di Talia, S., Skotheim, J. M., Bean, J. M., Siggia, E. D. & Cross, F. R. The effects of molecular noise and size control on variability in the budding yeast cell cycle. Nature 448, 947–951 (2007).
Cookson, N. A., Cookson, S. W., Tsimring, L. S. & Hasty, J. Cell cycle-dependent variations in protein concentration. Nucleic Acids Res. 38, 2676–2681 (2010).
Soifer, I., Robert, L. & Amir, A. Single-cell analysis of growth in budding yeast and bacteria reveals a common size regulation strategy. Curr. Biol. 26, 356–361 (2016).
Bryan, A. K., Engler, A., Gulati, A. & Manalis, S. R. Continuous and long-term volume measurements with a commercial Coulter counter. PLoS ONE 7, e29866 (2012).
Goranov, A. I. et al. The rate of cell growth is governed by cell cycle stage. Genes Dev. 23, 1408–1422 (2009).
Ferrezuelo, F. et al. The critical size is set at a single-cell level by growth rate to attain homeostasis and adaptation. Nat. Commun. 3, 1012 (2012).
Vergés, E., Colomina, N., Garí, E., Gallego, C. & Aldea, M. Cyclin Cln3 is retained at the ER and released by the J chaperone Ydj1 in late G1 to trigger cell cycle entry. Mol. Cell 26, 649–662 (2007).
Yahya, G., Parisi, E., Flores, A., Gallego, C. & Aldea, M. A Whi7-anchored loop controls the G1 Cdk–cyclin complex at Start. Mol. Cell 53, 115–126 (2014).
Dorsey, S. et al. G1/S transcription factor copy number is a growth-dependent determinant of cell cycle commitment in yeast. Cell Syst. 6, 539–554.e11 (2018).
Blank, H. M., Callahan, M., Pistikopoulos, I. P. E., Polymenis, A. O. & Polymenis, M. Scaling of G1 duration with population doubling time by a cyclin in Saccharomyces cerevisiae. Genetics 210, 895–906 (2018).
Tu, B. P., Kudlicki, A., Rowicka, M. & McKnight, S. L. Logic of the yeast metabolic cycle: temporal compartmentalization of cellular processes. Science 310, 1152–1158 (2005).
Papagiannakis, A., Niebel, B., Wit, E. C. & Heinemann, M. Autonomous metabolic oscillations robustly gate the early and late cell cycle. Mol. Cell 65, 285–295 (2017).
Slavov, N., Macinskas, J., Caudy, A. & Botstein, D. Metabolic cycling without cell division cycling in respiring yeast. Proc. Natl Acad. Sci. USA 108, 19090–19095 (2011).
Burnetti, A. J., Aydin, M. & Buchler, N. E. Cell cycle Start is coupled to entry into the yeast metabolic cycle across diverse strains and growth rates. Mol. Biol. Cell 27, 64–74 (2016).
Cai, L. & Tu, B. P. Driving the cell cycle through metabolism. Annu. Rev. Cell Dev. Biol. 28, 59–87 (2012).
Shi, L. & Tu, B. P. Acetyl-CoA induces transcription of the key G1 cyclin CLN3 to promote entry into the cell division cycle in Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 110, 7318–7323 (2013).
Unger, M. W. & Hartwell, L. H. Control of cell division in Saccharomyces cerevisiae by methionyl-tRNA. Proc. Natl Acad. Sci. USA 73, 1664–1668 (1976).
Özsezen, S. et al. (2019). Inference of the high-level interaction topology between the metabolic and cell cycle oscillators from single-cell dynamics. Cell Syst. https://doi.org/10.1016/j.cels.2019.09.003 (2019).
Lee, S. S., Avalos Vizcarra, I., Huberts, D. H. E. W., Lee, L. P. & Heinemann, M. Whole lifespan microscopic observation of budding yeast aging through a microfluidic dissection platform. Proc. Natl Acad. Sci. USA 109, 4916–4920 (2012).
Huberts, D. H. E. W. et al. Construction and use of a microfluidic dissection platform for long-term imaging of cellular processes in budding yeast. Nat. Protoc. 8, 1019–1027 (2013).
Elbing, K. et al. Role of hexose transport in control of glycolytic flux in Saccharomyces cerevisiae. Appl. Environ. Microbiol. 70, 5323–5330 (2004).
Yoshioka, K. et al. A novel fluorescent derivative of glucose applicable to the assessment of glucose uptake activity of Escherichia coli. Biochim. Biophys. Acta 1289, 5–9 (1996).
Snoep, J. L., Mrwebi, M., Schuurmans, J. M., Rohwer, J. M. & Teixeira de Mattos, M. J. Control of specific growth rate in Saccharomyces cerevisiae. Microbiology 155, 1699–1707 (2009).
Youk, H. & van Oudenaarden, A. Growth landscape formed by perception and import of glucose in yeast. Nature 462, 875–879 (2009).
Otterstedt, K. et al. Switching the mode of metabolism in the yeast Saccharomyces cerevisiae. EMBO Rep. 5, 532–537 (2004).
Novak, S., Zechner-Krpan, V. & Marić, V. Regulation of maltose transport and metabolism in Saccharomyces cerevisiae. Food Technol. Biotech. 42, 213–218 (2004).
Blount, B. A., Weenink, T., Vasylechko, S. & Ellis, T. Rational diversification of a promoter providing fine-tuned expression and orthogonal regulation for synthetic biology. PLoS ONE 7, e33279 (2012).
Gustavsson, A.-K. et al. Sustained glycolytic oscillations in individual isolated yeast cells. FEBS J. 279, 2837–2847 (2012).
Aon, M. A. et al. Dynamic regulation of yeast glycolytic oscillations by mitochondrial functions. J. Cell Sci. 99, 325–334 (1991).
Kang, H. T. & Hwang, E. S. 2-Deoxyglucose: an anticancer and antiviral therapeutic, but not any more a low glucose mimetic. Life Sci. 78, 1392–1399 (2006).
De Felipe, P. et al. E unum pluribus: multiple proteins from a self-processing polyprotein. Trends Biotechnol. 24, 68–75 (2006).
Nishimura, K., Fukagawa, T., Takisawa, H., Kakimoto, T. & Kanemaki, M. An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nat. Methods 6, 917–922 (2009).
Papagiannakis, A., de Jonge, J. J., Zhang, Z. & Heinemann, M. Quantitative characterization of the auxin-inducible degron: a guide for dynamic protein depletion in single yeast cells. Sci. Rep. 7, 4704 (2017).
Leupold, S. et al. Saccharomyces cerevisiae goes through distinct metabolic phases during its replicative lifespan. eLife 8, e41046 (2019).
Lew, D. J. & Reed, S. I. Morphogenesis in the yeast cell cycle: regulation by Cdc28 and cyclins. J. Cell Biol. 120, 1305–1320 (1993).
Bryan, A. K., Goranov, A., Amon, A. & Manalis, S. R. Measurement of mass, density, and volume during the cell cycle of yeast. Proc. Natl Acad. Sci. USA 107, 999–1004 (2010).
Parisi, E., Yahya, G., Flores, A. & Aldea, M. Cdc48/p97 segregase is modulated by cyclin-dependent kinase to determine cyclin fate during G1 progression. EMBO J. 37, e98724 (2018).
Futcher, B. Metabolic cycle, cell cycle, and the finishing kick to Start. Genome Biol. 7, 107 (2006).
Wullschleger, S., Loewith, R. & Hall, M. N. TOR signaling in growth and metabolism. Cell 124, 471–484 (2006).
Lindqvist, L. M., Tandoc, K., Topisirovic, I. & Furic, L. Cross-talk between protein synthesis, energy metabolism and autophagy in cancer. Curr. Opin. Genet. Dev. 48, 104–111 (2018).
Moore, S. A. Kinetic evidence for a critical rate of protein synthesis in the Saccharomyces cerevisiae yeast cell cycle. J. Biol. Chem. 263, 9674–9681 (1988).
Edgington, N. P. & Futcher, B. Relationship between the function and the location of G1 cyclins in S. cerevisiae. J. Cell Sci. 114, 4599–4611 (2001).
Jorgensen, P. et al. The size of the nucleus increases as yeast cells grow. Mol. Biol. Cell 18, 3523–3532 (2007).
Webster, M., Witkin, K. L. & Cohen-Fix, O. Sizing up the nucleus: nuclear shape, size and nuclear-envelope assembly. J. Cell Sci. 122, 1477–1486 (2009).
Wach, A. PCR-synthesis of marker cassettes with long flanking homology regions for gene disruptions in S. cerevisiae. Yeast 12, 259–265 (1996).
Verduyn, C., Postma, E., Scheffers, W. A. & Van Dijken, J. P. Effect of benzoic acid on metabolic fluxes in yeasts: a continuous-culture study on the regulation of respiration and alcoholic fermentation. Yeast 8, 501–517 (1992).
Shaner, N. C., Steinbach, P. A. & Tsien, R. Y. A guide to choosing fluorescent proteins. Nat. Methods 2, 905–909 (2005).
Shaner, N. C. et al. A bright monomeric green fluorescent protein derived from Branchiostoma lanceolatum. Nat. Methods 10, 407–409 (2013).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Waters, J. C. Accuracy and precision in quantitative fluorescence microscopy. J. Cell Biol. 185, 1135–1148 (2009).
Zopf, C. J., Quinn, K., Zeidman, J. & Maheshri, N. Cell-cycle dependence of transcription dominates noise in gene expression. PLoS Comput. Biol. 9, e1003161 (2013).
Doncic, A., Falleur-Fettig, M. & Skotheim, J. M. Distinct interactions select and maintain a specific cell fate. Mol. Cell 43, 528–539 (2011).
Rasmussen, C. E. & Williams, C. K. I. Gaussian Processes for Machine Learning (MIT Press, 2006).
Swain, P. S. et al. Inferring time derivatives including cell growth rates using Gaussian processes. Nat. Commun. 7, 13766 (2016).
MacKay, D. J. C. Information Theory, Inference, and Learning Algorithms (Cambridge Univ. Press, 2003).
Rasmussen, C. E. & Nickisch, H. Gaussian processes for machine learning (GPML) toolbox. J. Mach. Learn. Res. 11, 3011–3015 (2010).
Khmelinskii, A. et al. Tandem fluorescent protein timers for in vivo analysis of protein dynamics. Nat. Biotechnol. 30, 708–714 (2012).
Wang, X., Errede, B. & Elston, T. C. Mathematical analysis and quantification of fluorescent proteins as transcriptional reporters. Biophys. J. 94, 2017–2026 (2008).
Anderlei, T., Zang, W., Papaspyrou, M. & Büchs, J. Online respiration activity measurement (OTR, CTR, RQ) in shake flasks. Biochem. Eng. J. 17, 187–194 (2004).
Rosebrock, A. P. Synchronization of budding yeast by centrifugal elutriation. Cold Spring Harb. Protoc. 2017, pdb.prot088732 (2017).
Peterson, A. C., Russell, J. D., Bailey, D. J., Westphall, M. S. & Coon, J. J. Parallel reaction monitoring for high resolution and high mass accuracy quantitative, targeted proteomics. Mol. Cell. Proteomics 11, 1475–1488 (2012).
Ahrné, E. et al. Evaluation and improvement of quantification accuracy in isobaric mass tag-based protein quantification experiments. J. Proteome Res. 15, 2537–2547 (2016).
Del Vecchio, D. & Murray, R. M. Biomolecular Feedback Systems (Princeton Univ. Press, 2015).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 47, D442–D450 (2019).
Acknowledgements
The authors thank B. Tu and A. D. Ortega for critical comments on an early version of the manuscript, Z. Zhang for advice on microscopy, C. Åberg for discussions on model-based analysis of microscopy data, and the Ida van der Klei laboratory for provision of the pSNA10 plasmid. Financial support was provided by the EU ITN project ISOLATE (grant agreement 289995).
Author information
Authors and Affiliations
Contributions
A.L. and M.H. conceived the study. A.L., M.H. and A.M.-A. designed the study. A.L. constructed the strains, performed the experiments and analysed the data. D.H.E.W.H. performed the preliminary experiments and contributed conceptually. H.M.T. participated in strain construction and culture sampling for targeted proteomics. A.M.-A. performed the smoothing and derivative estimation for the single-cell time-lapse data. P.G. performed and analysed the verification experiments with confocal microscopy. A.S. and K.B. performed the targeted proteomics and analysed the data. A.P. participated in strain construction and metabolite measurements during batch cultivation, and performed the preliminary data analysis. M.R. performed the elutriation, and participated in culture sampling for targeted proteomics and respective data analysis. J.H. prepared the protein samples for mass spectrometry. G.H. performed the model-based analysis of the metabolite data for estimation of cellular physiology. M.E. participated in strain construction. A.L. and M.H. wrote the manuscript with input from A.M.-A. M.H. and A.M.-A. supervised the study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Cell cycle arrest in low flux conditions.
(a) Total sfGFP content of non-dividing cells (n = 10 cells) over time. sfGFP expressed via TEF1 promoter. (b) Left: Whi5 localizes primarily in the nucleus during G1 and shuttles in the cytoplasm during the Start transition69. Example merged phase-contrast and fluorescent images showing Whi5-mGFP localization in non-dividing and dividing TM6* (middle) and VW100-tet-Hxt1 (right) cells. Experiments repeated independently 3 times with similar results. (c) Glucose uptake rate assayed via 2-NBDG uptake, versus G1 duration in individual TM6* cells (n = 86 cells, Spearman r: -0.3784, p-value: 0.0003). (d) Percentage of G1-arrested cells as a function of glucose concentration in the microfluidics device. The number of cells analysed per condition is indicated in the parentheses. Grey line shows the exponential fit. (e) Hxt1 expression in response to tetracycline in the HXT-null strain carrying an Hxt1 copy under the control of a Tet-On promoter. Data from 3 technical replicates per condition are shown (except for VW100-tet-HXT1). (f) Rates of carbon uptake, ethanol and CO2 production, and O2 consumption for TM6* and wild type during growth on glucose and maltose. Centre values present means, and error bars propagated standard errors, as determined in the model-based regression analysis (data from 3 independent experiments used in regression; see Methods) with Maximum Likelihood formulation. (g) Distribution of time of first Start (as indicated by bud emergence) in TM6* cells (n = 956 cells from 4 experiments). (h) Dynamics of yeGFP production rate in G1-arrested TM6* cells (n = 20 cells) in response to increase in glycolytic flux achieved by switching the feed to 10 gL−1 maltose after ≈40 hours of cultivation on 10 gL−1 glucose. Vertical red line indicates the time when cells started passing Start (as indicated by bud emergence) in response to the nutrient switch. Source data for a and c-h are provided in Source Data Extended Data Fig. 1.
Extended Data Fig. 2 NAD(P)H autofluorescence as a reporter of metabolic dynamics and determination of protein production rate.
(a) Dynamics of NAD(P)H autofluorescence in wild type cells (n = 30 cells) in response to an increase in glycolytic flux achieved via a switch from glucose-low to glucose-rich media. Values of each single-cell NAD(P)H trajectory were normalized by division with the mean NAD(P)H value of the whole trajectory. (b) Dynamics of NAD(P)H autofluorescence aligned for the moment of birth in wild type daughter cells (n = 53 cells) in steady nutrient conditions. Histogram shows the distribution of the timing of Start in the same cells, as reported by the exit of Whi5-mCherry from the nucleus. (c) Example of smoothing the total sfGFP time series of a single cell during G1 using Gaussian process regression. Blue squares: measurement data; Red curve: posterior mean function; Grey band: 95% posterior confidence region. (d) The posterior Gaussian processes (from (c)) can be used to analytically derive an estimate of the rate of sfGFP production. This is again a Gaussian process, whose posterior distribution can be analytically obtained (see Methods). Red line: posterior mean of the derivative; Grey band: 95% posterior confidence region. (e) Comparison of cellular-autofluorescence-corrected and non-corrected total sfGFP measurements (n = 50 cells). For correction, the average total cellular autofluorescence of WT cells (n = 25 cells) at the GFP channel was smoothed, and was then subtracted from the average smoothed total sfGFP fluorescence. Data are normalized to the fluorescence value at the moment of bud appearance. Note that control cells undergo also unperturbed growth, and thus, corrected signals include correction for cell-size-dependent changes in cellular autofluorescence. Source data for Extended Data Fig. 2 are provided in Source Data Extended Data Fig. 2.
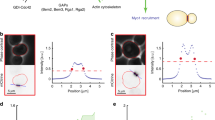
Extended Data Fig. 3 Confirmation of differential scaling with confocal microscopy and different analyses.
(a) Rate of sfGFP production versus cell size in daughter cells during G1, estimated by confocal microscopy (n = 27 cells), and by (b) widefield microscopy but determining total sfGFP on the basis of the integrated fluorescence over the whole cell area (n = 43 cells). (c) Coefficient of variation of Whi5-sfGFP fluorescence as a measure of Whi5 localization in a single cell, in comparison to the ratio between nuclear to whole-cell mean Whi5-sfGFP fluorescence (apart from Whi5-sfGFP, this cell expressed Hta2-mCherry for defining the nuclear area, which was used for the determination of the nuclear mean Whi5-sfGFP intensity). (d) Dynamics of sfGFP production rate and Whi5-mCherry localization (determined as shown in (c)) during G1 for the single cell in Fig. 3f, displayed in absolute time. As can be observed, Whi5 exhibits only a transient, partial exit from the nucleus during the first pulse in protein production, and exits completely the nucleus during the second pulse, suggesting that in the case of more than one pulses in protein production, cells attempt, but fail to pass Start during the first pulse likely due to insufficient phosphorylation of Whi5. In all cases sfGFP expression was driven by the TEF1 promoter, and cells were grown on 20 gL−1 glucose. Source data for Extended Data Fig. 3 are provided in Source Data Extended Data Fig. 3.
Extended Data Fig. 4 Differential scaling between Cln3 production rate and cell size dynamics.
(a) Likely due to the rapid Cln3 degradation, Cln3-sfGFP fusions do not generate detectable fluorescent signals, as can be seen in (b) (merged phase-contrast and fluorescent images of Cln3-sfGFP wild type cells mixed with wild type Hta2-mRFP1 cells as control for cellular autofluorescence). Experiment performed once with cells in multiple imaging positions, with each position yielding similar results. (c) Mean cell sfGFP in wild type (n = 43) and A-315T/CLN3 (n = 66) cells expressing the CLN3-2A-sfGFP fusion during growth on poor carbon source (20 gL−1 lactate) (Mann Whitney test p-value <0.0001). Horizontal lines denote the median. (d) Relative change in Cln3 levels across different growth rates, measured via the Cln3-2A-sfGFP fusion in single cells (this study), or via immunoblots in chemostat cultures (data from27). In both cases Cln3 levels are normalized against the highest Cln3 value. (e) Example of smoothing the total sfGFP originating from the Cln3-2A-sfGFP fusion for a single cell during G1 using Gaussian processes and (f) respective analytically derived estimate of Cln3 production rate (similarly to Extended Data Fig. 2c, d). (g) Comparison of cellular-autofluorescence-corrected and non-corrected total sfGFP measurements from the Cln3-2A-sfGFP construct (n = 38 cells). For correction, the average total cellular autofluorescence of WT cells (n = 35 cells) at the GFP channel was smoothed, and then subtracted from the average smoothed total sfGFP fluorescence. Data are normalized to fluorescence value at moment of bud appearance. Note that control cells undergo unperturbed growth, and thus, corrected signals include also correction for cell-size-dependent changes in cellular autofluorescence. (h) Rate of Cln3 production versus cell size in daughter cells for normalized G1 duration (n = 41 cells). (i) Same as (h) but for cells aligned for the moment of bud appearance. Unless otherwise indicated, cells were grown on 20 gL−1 glucose. Source data for c-i are provided in Source Data Extended Data Fig. 4.
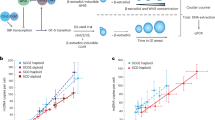
Extended Data Fig. 5 The increase in Cln3 concentration determines the timing of Start.
(a) Comparison of cellular-autofluorescence-corrected and non-corrected mean Whi5-sfGFP (n = 50 cells) and Whi5-mCherry (n = 50 cells) concentrations. For correction, the mean cellular autofluorescence (integrated intensity divided by cell volume) of WT cells at the GFP (n = 20 cells) or RFP (n = 20 cells) channels was subtracted from the mean Whi5-sfGFP or Whi5-mCherry based estimated Whi5 concentration respectively. Data are normalized to the value at the moment of bud appearance. Note that control cells undergo also unperturbed growth, and thus, corrected signals include correction for cell-size-dependent changes in cellular autofluorescence. (b) Whi5 concentration dynamics normalized to t = 0 in small G1 cells isolated by centrifugal elutriation and released to YPD. Whi5 concentration was calculated by measuring cell size changes, in parallel with Whi5 abundance via targeted proteomics, in the same samples as in Fig. 4e, f. The continuous line denotes the smoothing spline. Error bars show propagated SEM (n = 4 independent biological replicates). (c) Addition of higher NAA concentration leads to even further delay of Start. Duration of pre-Start G1 before (n = 49 and 24 cells) and after (n = 30 and 19 cells) addition of 2 mM NAA in OsTIR Cln3-AID and OsTIR Cln3 (control) cells. Indicated p-value from Mann Whitney test. Horizontal lines denote the median. (d) sfGFP production rate (driven by TEF1 promoter) in OsTIR Cln3-AID (n = 19 cells) and OsTIR Cln3 (control) (n = 17) cells treated with NAA (1 mM) before the second budding. Data are shown in normalized time between the two subsequent buddings, aligned for the moment of last bud appearance before addition of NAA (t = 0), and first bud appearance after addition of NAA (t = 100). Source data for Extended Data Fig. 5 are provided in Source Data Extended Data Fig. 5.
Extended Data Fig. 6 There is differential scaling between Cln3 production rate and cell size across different growth conditions.
(a) Whi5 concentration in wild type daughter cells for galactose (n = 50 and 46 cells for WF-1 and WF-2, respectively) and lactate (n = 50 cells for both WF-1 and WF-2) conditions, normalized for concentration at birth and aligned for the moment of Start. Dashed lines: WF-1 (mean cell fluorescence). Solid lines: WF-2 (integrated fluorescence over whole cell area divided by cell volume). (b) Change in cell size and Whi5-mCherry concentration (integrated fluorescence over whole cell area divided by cell volume) between cytokinesis and Start in mother cells on galactose (n = 42 cells) and (c) lactate (n = 42 cells). The vertical lines denote the respective population average. (d) Dynamics of sfGFP production rate and rate of NAD(P)H change in a single wild type cell at steady galactose (20 gL−1) or (e) lactate (20 gL−1) environment. (f) Cln3 production rate and respective cell size dynamics in wild type daughter cells aligned for the moment of bud appearance, on galactose (n = 36 cells) or (g) lactate (n = 43 cells) conditions. Source data for Extended Data Fig. 6 are provided in Source Data Extended Data Fig. 6.
Extended Data Fig. 7 The differential scaling between Cln3 production rate and cell size causes Start across different growth conditions.
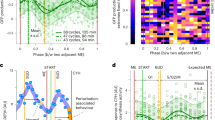
(a) Heatmap showing the dynamics of the Cln3 production rate during G1 in single wild type daughter cells on galactose and (b) lactate. The dark squares indicate the moment of Start in each cell. (c) Heatmap showing the dynamics of the Cln3 production rate in single wild type mother cells on galactose and (d) lactate. Cells are aligned for Start (t = 0) and cytokinesis is indicated in each cell by a dark square. In all cases data are normalized as described in Fig. 5b. Source data for Extended Data Fig. 7 are provided in Source Data Extended Data Fig. 7.
Extended Data Fig. 8 The effect of protein degradation on protein abundance dynamics.
To demonstrate the impact of degradation rate on protein abundance dynamics, we simulated the system described by Eq. (1) for different values of kd. To facilitate comparisons with the experimental data across different kd values, we used a smoothed version of the estimated Cln3 production rate (Fig. 5c) as kp(t) and assumed that the system is at equilibrium at t = 0 (that is \(p\left( 0 \right) = \frac{{k_p\left( 0 \right)}}{{k_d}}\)) for all values of kd except kd = 0 and kd = ∞. Furthermore, for all values of kd≠0 the plotted trajectories were normalized to their initial point by dividing p(t) by p(0). Finally, in the case of kd = 0 (no degradation), p(0) was set to zero. The use of the experimentally determined production rate allows us to assess the impact of different values of kd on time scales that are relevant for protein synthesis in the G1 phase. Altering kd affects the protein half-life (T1/2) since the two quantities are connected by the formula T1/2=ln(2)/kd. The curve corresponding to the asymptotic case T1/2→0 (kd→∞) is just kp(t) itself (normalized to its initial value), while the case T1/2 = ∞ corresponds to the accumulation of a highly stable protein such as GFP. As can be observed, a range of short protein half-lives (such as those reported for Cln3) between these two extremes results in protein abundance profiles that are very similar in shape to kp(t). As T1/2 increases, the peak of the protein abundance gets shifted to the right and the overall variation of the response gets reduced. This happens because the bandwidth of the system decreases with increasing half-life (decreasing degradation rate). In the limit of zero protein half-life, the protein abundance tracks the time integral of kp(t) (cyan line). Source data for Extended Data Fig. 8 are provided in Source Data Extended Data Fig. 8.
Supplementary information
Supplementary Table 1
List of the S. cerevisiae strains used in this study
Supplementary Table 2
List of the primers, and information on how they were used in strain construction
Supplementary Table 3
Detailed widefield microscopy settings
Supplementary Table 4
Peptides and transitions for PRM analysis
Source data
Rights and permissions
About this article
Cite this article
Litsios, A., Huberts, D.H.E.W., Terpstra, H.M. et al. Differential scaling between G1 protein production and cell size dynamics promotes commitment to the cell division cycle in budding yeast. Nat Cell Biol 21, 1382–1392 (2019). https://doi.org/10.1038/s41556-019-0413-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-019-0413-3
This article is cited by
-
Cell cycle-linked vacuolar pH dynamics regulate amino acid homeostasis and cell growth
Nature Metabolism (2023)
-
Temporal segregation of biosynthetic processes is responsible for metabolic oscillations during the budding yeast cell cycle
Nature Metabolism (2023)
-
Novel determinants of cell size homeostasis in the opportunistic yeast Candida albicans
Current Genetics (2023)
-
The G1/S repressor WHI5 is expressed at similar levels throughout the cell cycle
BMC Research Notes (2022)
-
Cell region fingerprints enable highly precise single-cell tracking and lineage reconstruction
Nature Methods (2022)