Abstract
Mechanistic target of rapamycin (mTOR) kinase functions in two multiprotein complexes: lysosomal mTOR complex 1 (mTORC1) and mTORC2 at the plasma membrane. mTORC1 modulates the cell response to growth factors and nutrients by increasing protein synthesis and cell growth, and repressing the autophagy–lysosomal pathway1,2,3,4; however, dysfunction in mTORC1 is implicated in various diseases3,5,6. mTORC1 activity is regulated by phosphoinositide lipids7,8,9,10. Class I phosphatidylinositol-3-kinase (PI3K)-mediated production of phosphatidylinositol-3,4,5-trisphosphate6,11 at the plasma membrane stimulates mTORC1 signalling, while local synthesis of phosphatidylinositol-3,4-bisphosphate by starvation-induced recruitment of class II PI3K-β (PI3KC2-β) to lysosomes represses mTORC1 activity12. How the localization and activity of PI3KC2-β are regulated by mitogens is unknown. We demonstrate that protein kinase N (PKN) facilitates mTORC1 signalling by repressing PI3KC2-β-mediated phosphatidylinositol-3,4-bisphosphate synthesis downstream of mTORC2. Active PKN2 phosphorylates PI3KC2-β to trigger PI3KC2-β complex formation with inhibitory 14-3-3 proteins. Conversely, loss of PKN2 or inactivation of its target phosphorylation site in PI3KC2-β represses nutrient signalling via mTORC1. These results uncover a mechanism that couples mTORC2-dependent activation of PKN2 to the regulation of mTORC1-mediated nutrient signalling by local lipid signals.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author on request. Mass spectrometry data have been deposited in ProteomeXchange. The primary accession code for screening data used for identifying nutrient-dependent interaction partners and phosphorylation of PI3KC2-β is PXD014533, the accession code for data used for identifying small Rab GTPases interacting with PI3KC2-β is PXD014530. Raw data for results from screening kinase activity towards T279 of PI3KC2-β are provided in Supplementary Table 2. Source data for Figs. 1–5, Supplementary Figs. 2, 4 and 5 are provided in Supplementary Table 4.
Change history
12 July 2021
In the version of this Article originally published, the file for Supplementary Table 3 was corrupted. The file has been replaced.
References
Jewell, J. L. & Guan, K. L. Nutrient signaling to mTOR and cell growth. Trends Biochem. Sci. 38, 233–242 (2013).
Kim, J., Kundu, M., Viollet, B. & Guan, K. L. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat. Cell Biol. 13, 132–141 (2011).
Laplante, M. & Sabatini, D. M. mTOR signaling in growth control and disease. Cell 149, 274–293 (2012).
Settembre, C. & Ballabio, A. Lysosomal adaptation: how the lysosome responds to external cues. Cold Spring Harb. Perspect. Biol. 6, a016907 (2014).
Shimobayashi, M. & Hall, M. N. Making new contacts: the mTOR network in metabolism and signalling crosstalk. Nat. Rev. Mol. Cell Biol. 15, 155–162 (2014).
Vanhaesebroeck, B., Stephens, L. & Hawkins, P. PI3K signalling: the path to discovery and understanding. Nat. Rev. Mol. Cell Biol. 13, 195–203 (2012).
Balla, T. Phosphoinositides: tiny lipids with giant impact on cell regulation. Physiol. Rev. 93, 1019–1137 (2013).
De Matteis, M. A. & Godi, A. PI-loting membrane traffic. Nat. Cell Biol. 6, 487–492 (2004).
Di Paolo, G. & De Camilli, P. Phosphoinositides in cell regulation and membrane dynamics. Nature 443, 651–657 (2006).
Marat, A. L. & Haucke, V. Phosphatidylinositol 3-phosphates-at the interface between cell signalling and membrane traffic. EMBO J. 35, 561–579 (2016).
Wymann, M. P. & Schneiter, R. Lipid signalling in disease. Nat. Rev. Mol. Cell Biol. 9, 162–176 (2008).
Marat, A. L. et al. mTORC1 activity repression by late endosomal phosphatidylinositol 3,4-bisphosphate. Science 356, 968–972 (2017).
Alliouachene, S. et al. Inactivation of the class II PI3K-C2β potentiates insulin signaling and sensitivity. Cell Rep. 13, 1881–1894 (2015).
Campa, C. C., Franco, I. & Hirsch, E. PI3K-C2α: one enzyme for two products coupling vesicle trafficking and signal transduction. FEBS Lett. 589, 1552–1558 (2015).
Gwinn, D. M. et al. AMPK phosphorylation of raptor mediates a metabolic checkpoint. Mol. Cell 30, 214–226 (2008).
Collins, B. C. et al. Quantifying protein interaction dynamics by SWATH mass spectrometry: application to the 14-3-3 system. Nat. Methods 10, 1246–1253 (2013).
Yaffe, M. B. et al. The structural basis for 14-3-3:phosphopeptide binding specificity. Cell91, 961–971 (1997).
Fruman, D. A. et al. The PI3K pathway in human disease. Cell 170, 605–635 (2017).
Lemmon, M. A. & Schlessinger, J. Cell signaling by receptor tyrosine kinases. Cell 141, 1117–1134 (2010).
Falk, M. D. et al. Enzyme kinetics and distinct modulation of the protein kinase N family of kinases by lipid activators and small molecule inhibitors. Biosci. Rep. 34, e00097 (2014).
Menon, S. et al. Spatial control of the TSC complex integrates insulin and nutrient regulation of mTORC1 at the lysosome. Cell 156, 771–785 (2014).
Behnia, R. & Munro, S. Organelle identity and the signposts for membrane traffic. Nature 438, 597–604 (2005).
Pacold, M. E. et al. Crystal structure and functional analysis of Ras binding to its effector phosphoinositide 3-kinase gamma. Cell 103, 931–943 (2000).
Dibble, C. C. et al. TBC1D7 is a third subunit of the TSC1–TSC2 complex upstream of mTORC1. Mol. Cell 47, 535–546 (2012).
Arencibia, J. M., Pastor-Flores, D., Bauer, A. F., Schulze, J. O. & Biondi, R. M. AGC protein kinases: from structural mechanism of regulation to allosteric drug development for the treatment of human diseases. Biochim. Biophys. Acta 1834, 1302–1321 (2013).
Gaubitz, C., Prouteau, M., Kusmider, B. & Loewith, R. TORC2 structure and function. Trends Biochem. Sci. 41, 532–545 (2016).
Lim, W. G. et al. The C-terminus of PRK2/PKNγ is required for optimal activation by RhoA in a GTP-dependent manner. Arch. Biochem. Biophys. 479, 170–178 (2008).
Schmidt, A., Durgan, J., Magalhaes, A. & Hall, A. Rho GTPases regulate PRK2/PKN2 to control entry into mitosis and exit from cytokinesis. EMBO J. 26, 1624–1636 (2007).
Flynn, P., Mellor, H., Casamassima, A. & Parker, P. J. Rho GTPase control of protein kinase C-related protein kinase activation by 3-phosphoinositide-dependent protein kinase. J. Biol. Chem. 275, 11064–11070 (2000).
Danno, S. et al. PKN2 is essential for mouse embryonic development and proliferation of mouse fibroblasts. Genes Cells 22, 220–236 (2017).
Quetier, I. et al. Knockout of the PKN family of Rho effector kinases reveals a non-redundant role for PKN2 in developmental mesoderm expansion. Cell Rep. 14, 440–448 (2016).
Lachmann, S. et al. Regulatory domain selectivity in the cell-type specific PKN-dependence of cell migration. PLoS ONE 6, e21732 (2011).
Yang, Z., Zhao, X., Xu, J., Shang, W. & Tong, C. A novel fluorescent reporter detects plastic remodeling of mitochondria-ER contact sites. J. Cell Sci. 131, jcs208686 (2018).
Cheng, Y. et al. PKN2 in colon cancer cells inhibits M2 phenotype polarization of tumor-associated macrophages via regulating DUSP6–Erk1/2 pathway. Mol. Cancer 17, 13 (2018).
Smythe, E. & Ayscough, K. R. The Ark1/Prk1 family of protein kinases: regulators of endocytosis and the actin skeleton. EMBO Rep. 4, 246–251 (2003).
Wallroth, A. & Haucke, V. Phosphoinositide conversion in endocytosis and the endolysosomal system. J. Biol. Chem. 293, 1526–1535 (2018).
Riggi, M. et al. Decrease in plasma membrane tension triggers PtdIns(4,5)P2 phase separation to inactivate TORC2. Nat. Cell Biol. 20, 1043–1051 (2018).
Mansueto, G. et al. Transcription factor EB controls metabolic flexibility during exercise. Cell Metab. 25, 182–196 (2017).
Ruby, M. A., Riedl, I., Massart, J., Ahlin, M. & Zierath, J. R. Protein kinase N2 regulates AMP kinase signaling and insulin responsiveness of glucose metabolism in skeletal muscle. Am. J. Physiol. Endocrinol. Metab. 313, E483–E491 (2017).
Ebner, M., Sinkovics, B., Szczygiel, M., Ribeiro, D. W. & Yudushkin, I. Localization of mTORC2 activity inside cells. J. Cell Biol. 216, 343–353 (2017).
Acknowledgements
We thank H. Stephanowitz, S. Zillmann and D. Löwe for technical assistance, M. Hessenberger and W.-T. Lo for help with biochemical experiments, and Y. Yudushkin (University of Vienna, Vienna, Austria) for the mSin1-GFP plasmid. This work was supported by a fellowship from the Sonnenfeld foundation to P.A.K. and a grant from the German Research Foundation to V.H. (TRR186/A08).
Author information
Authors and Affiliations
Contributions
A.W. and P.A.K. performed all of the biochemical and cell biological experiments. A.L.M. initiated the study. E.K. conducted the proteomic analyses. V.H. designed the study and wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Serum growth factor-induced phosphorylation of PI3KC2β at T279 induces dimerisation and complex formation with inhibitory 14-3-3 proteins.
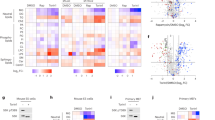
a, Proteins found as interaction partners of PI3KC2β by SILAC-based quantitative mass spectrometry of material immunoprecipiated from serum-fed vs. serum-starved HEK293T cells endogenously expressing eGFP-PI3KC2β. b, HA-14-3-3β, HA-14-3-3γ or HA-14-3-3τ were overexpressed in HEK293T endogenously expressing eGFP-PI3KC2β. Cells were serum-starved for 2h (+) or fed with 10% FBS (-), lysed, and subjected to affinity capture by GFP-nanotrap magnetic beads or magnetic beads as a control. Bound proteins were detected by immunoblotting. SM, 5% of total lysate. c, HEK293T endogenously expressing eGFP-PI3KC2β were starved in serum-free medium overnight and subsequently incubated for 30 min with either serum-free medium (SF), fresh DMEM with 10% FBS (steady state) or serum-free medium supplemented with 50ng/mL of IGF (SF+IGF), lysed and experimentall handled as in b. Experiment was performed 3 times. d, PI3KC2β-derived phosphopeptides detected by LC-MS/MS. e, Alignment of human class II PI3K isoforms. Binding sites within PI3KC2β for Raptor (red) and the 14-3-3 binding motif (green box) are marked in the sequence. f, Conservation of the 14-3-3 binding motif in PI3KC2β among vertebrate species. g, eGFP, eGFP-PI3KC2α or eGFP-PI3KC2β expressed in HEK293T cells were experimentally handled as in b. Experiment was performed 3 times. h, exemplary Ponceau-staining (corresponding to g) visualizing the bands of eGFP, eGFP-PI3KC2α or eGFP-PI3KC2β being immunoprecipitated using GFP. i, PI3KC2β dimerisation in cells. HEK293T cells endogenously expressing eGFP-PI3KC2β were transfected with a plasmid encoding HA-tagged PI3KC2β, lysed and experimentalyy handled as in b. j, eGFP-PI3KC2β was purified from HEK293T and subsequently incubated with PKN2, ATP and recombinant 14-3-3β protein (PI3KC2β + 14-3-3) or PBS (PI3KC2β). PI3KC2β was cut from the beads using PreScission protease and subjected to crosslinking with glutaraldehyde. PI3KC2β was detected by immunoblot with specific antibodies. Monomer and dimer bands were identified by appropriate size markers.
Supplementary Figure 2 PKN2 phosphorylates PI3KC2β at T279 to induce its cytoplasmic sequestration with 14-3-3 proteins and facilitate mTORC1 signaling.
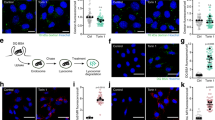
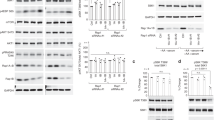
a, PKN induces PI3KC2β-14-3-3 interaction. Immunoblot analysis of HEK293T endogenously expressing eGFP-PI3KC2β treated with the indicated inhibitors for 2h, lysed, and affinity captured by GFP-nanotrap. SM, 5% of lysate. b, PKN1 phosphorylates eGFP-PI3KC2β to induce 14-3-3 complex formation. eGFP-PI3KC2β purified from HEK293T was incubated with PKN1 or PKCβ and recombinant GST-14-3-3β, isolated on GFP-nanotrap and analyzed by immunoblotting. c, PKN2 expression facilitates complex formation of 14-3-3 proteins with wild-type (WT) but not PI3KC2βT279A. HEK293T expressing WT or eGFP-PI3KC2βT279A with or without mCherry-PKN2 were analyzed as in a. d, HA-tagged kinases were expressed in HEK293T eGFP-PI3KC2β knock-in cells. Cells were lysed and analyzed as in a. e, WT or T279A mutant eGFP-PI3KC2β expressed in HEK293T. Cells were serum starved for 2h and refed with full medium for 30min, lysed and analyzed as in a. f, Overexpression of PKN2 represses PI3KC2β complex formation with Raptor. Experiment performed as in d. g-h, PKN2 overexpression leads to higher mTORC1 activity only in cells cultured in serum containing media. g, Immunoblot of HEK293T expressing mCherry or mCherry-PKN2 (steady state), lysed and analyzed by immunoblotting. Right, quantification of representative data shown left. Ratio of phospho-S6K Thr389/total S6K. Mean ± SEM (n=3 independent experiments). One-sample two-tailed T test, hypothetical mean of 1, *p=0.0333. h, Immunoblot of HEK293T overexpressing mCherry or mCherry-PKN2, fed or serum starved 3h, lysed and analyzed by immunoblotting. i, Depletion of PKN1/2 impairs PI3KC2β-14-3-3 interaction while inducing Raptor-association. HEK293T eGFP-PI3KC2β knock-in cells were treated with indicated siRNAs, lysed and analyzed as in a. j, PKN1/2 loss impairs mTORC1 signaling. PKN1/2 WT or KO eGFP-PI3KC2β knock-in HEK293T cells were kept in serum-containing media, serum-starved, or serum-starved and refed with full media for 30 min. Cell lysates were analyzed by immunoblotting. k, Loss of PKN2 reduces cell size. HeLa cells were treated with indicated siRNAs, fixed and stained for the LyLE marker LBPA (white). DAPI (blue), cell nuclei. Scale bar, 10 µm. Footprint area, (scrambled n=90 cells, siPKN2 n=90 cells). Values in μm2. Unpaired two-tailed T-test **, p=0.006.
Supplementary Figure 3 PKN2 and Rab7 regulate PI3KC2β localization independent of mTOR and TSC.
a, Representative confocal images of HeLa cells that were treated with indicated siRNAs and stained for endogenous mTOR and LAMP1. Scale bar, 10 µm. b, Representative confocal images of HeLa cells that were treated with indicated siRNAs and stained for endogenous TSC2 and LAMP1. Scale bar, 10 µm. c, HEK293T endogenously expressing eGFP-PI3KC2β were either treated with indicated siRNAs, lysed, subjected to affinity capture by GFP-nanotrap or control beads (ctr-IP) and analyzed by immunoblotting as indicated. SM, 5% of the total lysate. d, Multiple sequence alignment of PI3KC2β and PI3Kγ. The red box highlights D1168 in PI3KC2β that was mutated to K for mass spectrometry-based screening of interacting small GTPases. e, Interacting Rab GTPases identified by LC-MS/MS. f, HEK293T endogenously expressing eGFP-PI3KC2β were serum-starved for 2h or fed with 10% FBS, lysed, and experimentally treated as in c. SM, 5% of the total lysate. g, Representative confocal images of HeLa cells endogenously expressing eGFP-PI3KC2β treated with indicated siRNAs and stained for GFP. Scale bar, 10 µm. Experiment was performed 1 time. h, Representative confocal images of HeLa cells endogenously expressing eGFP-PI3KC2β that were either serum-starved for 2h or re-fed with 10% FBS and stained for endogenous Rab7 and GFP. Scale bar, 10 µm.
Supplementary Figure 4 Rho and PKN2 facilitate nutrient signaling via mTORC1 by repressing lysosomal PI(3,4)P2 synthesis mediated by PI3KC2β.
a, PI(3,4)P2 accumulates at Raptor containing lysosomes in HeLa depleted of PKN2. Representative confocal image of HeLa cells treated with indicated siRNAs, stained as indicated. Scale bar, 10 µm. Experiment was performed 3 times. b, Hek293T were treated with indicated siRNAs. Cell lysates were subjected to immunoblotting. c, PI3KC2βT279A represses mTORC1-activity more potent than wild type PI3KC2β independent of TSC2. Hela cells overexpressing eGFP-PI3KC2β wild-type or phosphorylation-defectiveT279A mutant PI3KC2β (50min amino acid-starved) in a background of TSC2-knockdown. Cells were analyzed by immunoblotting. Right, quantification of representative data shown on the left. Ratio of phospho-S6K Thr389/total S6K. Mean ± SEM (n=3 independent experiments). One-sample two-tailed T test, hypothetical mean of 1, *p=0.0176. d, PKN2 overexpression inhibits starvation-dependent recruitment of PI3KC2β to Ly/LEs. Representative confocal images of HeLa that were mock-transfected or transfected with mCherry-PKN2 and either refed 2h prior or serum-starved for 2h prior to fixation. Cells were stained as indicated. Scale bar, 10 µm. e, Representative confocal images of HeLa cells that were either mock-transfected or transfected with mCherry-PKN2 and serum starved 2h prior to fixation. Cells were stained as indicated. Scale bar, 10 µm. f, Representative confocal images of HeLa cells that were either mock-transfected or transfected with mCherry-PKN2 and serum-starved 2h prior to fixation. Cells were stained as indicated. Scale bar, 10 µm.
Supplementary Figure 5 Inhibition of Rho signaling impairs PI3KC2β sequestration with inhibitory 14-3-3 proteins.
a, immunoblot of mTORC2 purified from HEK293T cells via mSin1-eGFP (corresponding to F5b). Experiment was performed 3 times. b, Inhibition of Rho signaling impairs PI3KC2β sequestration with 14-3-3 proteins. HEK293T cells endogenously expressing eGFP-PI3KC2β were treated (4h) with the specific Rho inhibitor Rhosin or DMSO as a solvent control. Cells were lysed, and subjected to affinity capture by GFP-nanotrap magnetic beads or magnetic beads as a control. Bound 14-3-3 proteins were detected by immunoblotting. In parallel, cell lysates (SM, 5% of the total lysate) were analyzed with specific antibodies against PKN2 or phosphorylated, active pPKN1/2 (T774/T816). c, Quantification of representative data shown on the left. Ratio of 14-3-3 proteins bound to endogenous eGFP-PI3KC2β in control vs. Rhosin-treated cells. Mean ± SEM (n=3 independent experiments). One-sample two-tailed T test, hypothetical mean of 1, *p=0.02.
Supplementary Figure 6 Unprocessed scans for all blots presented in figures.
Included here are uncropped blots corresponding to the indicated figures.
Supplementary information
Supplementary Information
Supplementary Figures 1–6 and Supplementary Table titles/legends.
Supplementary Table 1
SILAC-based proteomic screen of proteins associated with endogenously expressing eGFP-PI3KC2β 10 immunoprecipitated from serum-fed vs. serum-starved HEK293T cells.
Supplementary Table 2
Medium-throughput screen of 245 mammalian kinases for activity towards the PI3KC2β-derived T279-containing 14- 15 3–3 consensus peptide.
Supplementary Table 3
D1168K-mutant eGFP-PI3KC2β interactome from HEK293T cells.
Supplementary Table 4
Numerical source data for Figures 1,2,3,4,5, S2, S4, and S5.
Supplementary Table 5
Antibody information.
Rights and permissions
About this article
Cite this article
Wallroth, A., Koch, P.A., Marat, A.L. et al. Protein kinase N controls a lysosomal lipid switch to facilitate nutrient signalling via mTORC1. Nat Cell Biol 21, 1093–1101 (2019). https://doi.org/10.1038/s41556-019-0377-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-019-0377-3
This article is cited by
-
Phosphoinositides as membrane organizers
Nature Reviews Molecular Cell Biology (2022)
-
Melatonin induces the rejuvenation of long-term ex vivo expanded periodontal ligament stem cells by modulating the autophagic process
Stem Cell Research & Therapy (2021)
-
Class II phosphatidylinositol 3-kinase α and β isoforms are required for vascular smooth muscle Rho activation, contraction and blood pressure regulation in mice
The Journal of Physiological Sciences (2020)
-
1H, 15N and 13C resonance assignments of the HR1c domain of PRK1, a protein kinase C-related kinase
Biomolecular NMR Assignments (2020)