Abstract
Haematopoietic stem cells (HSCs) maintain balanced self-renewal and differentiation, but how these functions are precisely regulated is not fully understood. N6-methyladenosine (m6A) messenger RNA methylation has emerged as an important mode of epitranscriptional gene expression regulation affecting many biological processes. We show that deletion of the m6A methyltransferase Mettl3 from the adult haematopoietic system led to an accumulation of HSCs in the bone marrow and a marked reduction of reconstitution potential due to a blockage of HSC differentiation. Interestingly, deleting Mettl3 from myeloid cells using Lysm-cre did not impact myeloid cell number or function. RNA sequencing revealed 2,073 genes with significant m6A modifications in HSCs. Myc was identified as a direct target of m6A in HSCs. Mettl3-deficient HSCs failed to upregulate MYC expression following stimulation to differentiate and enforced expression of Myc rescued differentiation defects of Mettl3-deficient HSCs. Our results reveal a key role of m6A in governing HSC differentiation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Sequencing data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under accession code GSE123527. Source data have been provided as Supplementary Table 3. All other data supporting the findings of this study are available from the corresponding authors on reasonable request.
References
Rossi, L. et al. Less is more: unveiling the functional core of hematopoietic stem cells through knockout mice. Cell Stem Cell 11, 302–317 (2012).
Morrison, S. J. & Scadden, D. T. The bone marrow niche for haematopoietic stem cells. Nature 505, 327–334 (2014).
Desrosiers, R., Friderici, K. & Rottman, F. Identification of methylated nucleosides in messenger RNA from Novikoff hepatoma cells. Proc. Natl Acad. Sci. USA 71, 3971–3975 (1974).
Meyer, K. D. & Jaffrey, S. R. The dynamic epitranscriptome: N 6-methyladenosine and gene expression control. Nat. Rev. Mol. Cell Biol. 15, 313–326 (2014).
Meyer, K. D. & Jaffrey, S. R. Rethinking m6A readers, writers, and erasers. Ann. Rev. Cell Dev. Biol. 33, 319–342 (2017).
Geula, S. et al. Stem cells. m6A mRNA methylation facilitates resolution of naive pluripotency toward differentiation. Science 347, 1002–1006 (2015).
Sledz, P. & Jinek, M. Structural insights into the molecular mechanism of the m6A writer complex. eLife 5, e18434 (2016).
Wang, P., Doxtader, K. A. & Nam, Y. Structural basis for cooperative function of Mettl3 and Mettl14 methyltransferases. Mol. Cell 63, 306–317 (2016).
Wang, X. et al. Structural basis of N 6-adenosine methylation by the METTL3–METTL14 complex. Nature 534, 575–578 (2016).
Xiang, Y. et al. RNA m6A methylation regulates the ultraviolet-induced DNA damage response. Nature 543, 573–576 (2017).
Agarwala, S. D., Blitzblau, H. G., Hochwagen, A. & Fink, G. R. RNA methylation by the MIS complex regulates a cell fate decision in yeast. PLoS Genet. 8, e1002732 (2012).
Fustin, J. M. et al. RNA-methylation-dependent RNA processing controls the speed of the circadian clock. Cell 155, 793–806 (2013).
Chen, T. et al. m6A RNA methylation is regulated by microRNAs and promotes reprogramming to pluripotency. Cell Stem Cell 16, 289–301 (2015).
Batista, P. J. et al. m6A RNA modification controls cell fate transition in mammalian embryonic stem cells. Cell Stem Cell 15, 707–719 (2014).
Yoon, K. J. et al. Temporal control of mammalian cortical neurogenesis by m6A methylation. Cell 171, 877–889 (2017).
Wang, Y. et al. N 6-Methyladenosine RNA modification regulates embryonic neural stem cell self-renewal through histone modifications. Nat. Neurosci. 21, 195–206 (2018).
Zhang, C. et al. m6A modulates haematopoietic stem and progenitor cell specification. Nature 549, 273–276 (2017).
Vu, L. P. et al. The N 6-methyladenosine (m6A)-forming enzyme METTL3 controls myeloid differentiation of normal hematopoietic and leukemia cells. Nat. Med. 23, 1369–1376 (2017).
Weng, H. et al. METTL14 inhibits hematopoietic stem/progenitor differentiation and promotes leukemogenesis via mRNA m6A modification. Cell Stem Cell 22, 191–205 (2017).
Kuhn, R., Schwenk, F., Aguet, M. & Rajewsky, K. Inducible gene targeting in mice. Science 269, 1427–1429 (1995).
Rodriguez-Fraticelli, A. E. et al. Clonal analysis of lineage fate in native haematopoiesis. Nature 553, 212–216 (2018).
Carrelha, J. et al. Hierarchically related lineage-restricted fates of multipotent haematopoietic stem cells. Nature 554, 106–111 (2018).
Ding, L., Saunders, T. L., Enikolopov, G. & Morrison, S. J. Endothelial and perivascular cells maintain haematopoietic stem cells. Nature 481, 457–462 (2012).
Ding, L. & Morrison, S. J. Haematopoietic stem cells and early lymphoid progenitors occupy distinct bone marrow niches. Nature 495, 231–235 (2013).
de Luca, C. et al. Complete rescue of obesity, diabetes, and infertility in db/db mice by neuron-specific LEPR-B transgenes. J. Clin. Invest. 115, 3484–3493 (2005).
Clausen, B. E., Burkhardt, C., Reith, W., Renkawitz, R. & Forster, I. Conditional gene targeting in macrophages and granulocytes using LysMcre mice. Transgenic Res. 8, 265–277 (1999).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Forsberg, E. C. et al. Molecular signatures of quiescent, mobilized and leukemia-initiating hematopoietic stem cells. PLoS One 5, e8785 (2010).
He, S., Kim, I., Lim, M. S. & Morrison, S. J. Sox17 expression confers self-renewal potential and fetal stem cell characteristics upon adult hematopoietic progenitors. Genes Dev. 25, 1613–1627 (2011).
Ivanova, N. B. et al. A stem cell molecular signature. Science 298, 601–604 (2002).
Reavie, L. et al. Regulation of hematopoietic stem cell differentiation by a single ubiquitin ligase-substrate complex. Nat. Immunol. 11, 207–215 (2010).
Ehninger, A. et al. Posttranscriptional regulation of c-Myc expression in adult murine HSCs during homeostasis and interferon-alpha-induced stress response. Blood 123, 3909–3913 (2014).
Wilson, A. et al. c-Myc controls the balance between hematopoietic stem cell self-renewal and differentiation. Genes Dev. 18, 2747–2763 (2004).
Zeller, K. I., Jegga, A. G., Aronow, B. J., O’Donnell, K. A. & Dang, C. V. An integrated database of genes responsive to the Myc oncogenic transcription factor: identification of direct genomic targets. Genome Biol. 4, R69 (2003).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Huang, C. Y., Bredemeyer, A. L., Walker, L. M., Bassing, C. H. & Sleckman, B. P. Dynamic regulation of c-Myc proto-oncogene expression during lymphocyte development revealed by a GFP-c-Myc knock-in mouse. Eur. J. Immunol. 38, 342–349 (2008).
Yue, Y., Liu, J. & He, C. RNA N 6-methyladenosine methylation in post-transcriptional gene expression regulation. Genes Dev. 29, 1343–1355 (2015).
Yao, Q. J. et al. Mettl3–Mettl14 methyltransferase complex regulates the quiescence of adult hematopoietic stem cells. Cell Res. 28, 952–954 (2018).
Barbieri, I. et al. Promoter-bound METTL3 maintains myeloid leukaemia by m6A-dependent translation control. Nature 552, 126–131 (2017).
Rodriguez, C. I. et al. High-efficiency deleter mice show that FLPe is an alternative to Cre-loxP. Nat. Genet. 25, 139–140 (2000).
Yan, Q. et al. Systematic discovery of regulated and conserved alternative exons in the mammalian brain reveals NMD modulating chromatin regulators. Proc. Natl Acad. Sci. USA 112, 3445–3450 (2015).
Wu, J., Anczukow, O., Krainer, A. R., Zhang, M. Q. & Zhang, C. OLego: fast and sensitive mapping of spliced mRNA-Seq reads using small seeds. Nucleic Acids Res. 41, 5149–5163 (2013).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Acknowledgements
L.D. was supported by the Rita Allen Foundation and the National Heart, Lung and Blood Institute (grant no. R01HL132074). H.L. was supported by the Columbia Medical Scientist Training Program and the National Heart, Lung and Blood Institute (grant no. F30HL142196-01). The work by S.B., Y.Q. and C.Z. is in part supported by grants from the National Institute of Health (grant nos R01NS89676, R01GM124486 and R03HG009528). S.B. is supported by a Columbia Precision Medicine Research Fellowship. J.H.H. is supported by a generous gift from Ilana and Pascal Mantoux, and research grants from the European Research Council, Kimmel Stem Cell Research Center, Flight Attendant Medical Research Council (FAMRI), Israel Science Foundation, Israel Cancer Research Fund and New York Stem Cell Foundation. J.H.H. is a New York Stem Cell Foundation–Robertson Investigator. We thank M. Kissner at the Columbia Stem Cell Initiative, S. Ho at the Columbia Center for Translational Immunology and A. Figueroa at the Department of Microbiology and Immunology for assistance with flow cytometry. We thank I. Ivanov at the Department of Microbiology and Immunology for sharing the Lysm-cre mice and H. Snoeck for sharing the Mx1-cre mice. We thank C. Schindler, S. Ghosh and A. Lepelley at the Department of Microbiology and Immunology for helping with the macrophage assays. This research was funded in part through the NIH/NCI Cancer Center support grant no. P30CA013696.
Author information
Authors and Affiliations
Contributions
H.L., J.L. and L.D. performed all of the experiments. S.B., Y.Q. and C.Z. performed the bioinformatics analysis on all sequencing data. S.G. and J.H.H. generated and validated the Mettl3fl strain and assisted with gene expression analysis and the development of the project. H.L. and L.D. designed the experiments, interpreted the results and wrote the manuscript with input from J.H.H. and C.Z. L.D. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Fig. 1 Mettl3 is efficiently deleted from HSCs in Mx1-cre; Mettl3fl/fl mice after Cre induction.
(a) qPCR analysis of Mettl3 mRNA expression in wild-type bone marrow populations. (n=7 for WBM, n=6 for LSK, n=10 for HSC, n=6 for MPP, n=6 for LMPP, n=6 for CLP, n=4 for MEP, n=4 for CMP, n=3 for GMP, n=10 for Gr1+Mac1+). (b) Schematic of Cre-mediated recombination of Mettl3fl allele (left) and Mx1-cre; Mettl3fl/fl pIpC treatment schedule (right). (c) Representative genotyping results of colonies formed by single sorted HSCs after pIpC treatment. Experiment was repeated with biologically independent samples three times with similar results. Unprocessed gel scans are in Supplementary Fig. 7. (d) qPCR analysis of Mettl3 mRNA expression in HSCs from pIpC-treated control and Mx1-cre; Mettl3fl/fl mice (n=4 for control, n=5 for Mx1-cre; Mettl3fl/fl). All samples are from biologically independent animals. Values are shown as individual points with mean ± s.d. P values were determined by unpaired two-sided Student’s t-test.
Supplementary Fig. 2 Mx1-cre; Mettl3fl/fl mice have defective haematopoiesis.
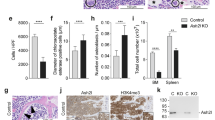
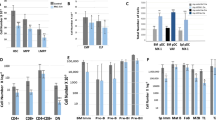
(a) Red blood cell count of pIpC-treated control and Mx1-cre; Mettl3fl/fl mice (n=7 for control (10–14d), n=7 for Mx1-cre; Mettl3fl/fl (10–14d), n=4 for control (2–3m), n=4 for Mx1-cre; Mettl3fl/fl (2-3m), n=3 for control (4m), n=4 for Mx1-cre; Mettl3fl/fl (4m)). (b) White blood cell count differential for control and Mx1-cre; Mettl3fl/fl mice 4 months after pIpC treatment (n=3 for control, n=4 for Mx1-cre; Mettl3fl/fl). (c) Frequency of cell populations in the spleens of Mx1-cre; Mettl3fl/fl mice at least 14 days after pIpC treatment with prominent splenomegaly compared to controls (n=5 for control, n=5 for Mx1-cre; Mettl3fl/fl, except LSK, n=4 for control, n=4 for Mx1-cre; Mettl3fl/fl). (d) Representative flow cytometric plots of Lin- gated and LSK gated bone marrow cells stained for the indicated cell surface markers. Numbers represent the population frequencies among single live bone marrow cells. (e) Bone marrow HSCs per hindlimb (n=7 for control (10-14d), n=6 for Mx1-cre; Mettl3fl/fl (10-14d), n=5 for control (2-3m), n=6 for Mx1-cre; Mettl3fl/fl (2-3m), n=4 for control (4m), n=4 for Mx1-cre; Mettl3fl/fl (4m)). (f) Bone marrow CD150-CD48-LSK MPP, CD150-CD48+LSK and CD150+CD48+LSK progenitor frequencies (MPP, n=6 for control (10-14d), n=6 for Mx1-cre; Mettl3fl/fl (10-14d), n=6 for control (2-3m), n=7 for Mx1-cre; Mettl3fl/fl (2-3m), n=4 for control (4m), n=4 for Mx1-cre; Mettl3fl/fl (4m); CD150-CD48+LSK and CD150+CD48+LSK, n=5 for control (10-14d), n=5 for Mx1-cre; Mettl3fl/fl (10-14d), n=5 for control (2-3m), n=6 for Mx1-cre; Mettl3fl/fl (2-3m), n=4 for control (4m), n=4 for Mx1-cre; Mettl3fl/fl (4m)). (g) Frequencies of bone marrow progenitor populations. CLP, common lymphoid progenitor; CMP, common myeloid progenitors; GMP, granulocyte–monocyte progenitors; MEP, megakaryocyte–erythroid progenitors; LMPP, lymphoid-primed multipotent progenitors (n=4 for control (10-14d), n=4 for Mx1-cre; Mettl3fl/fl (10-14d), n=5 for control (2-3m), n=6 for Mx1-cre; Mettl3fl/fl (2-3m), n=4 for control (4m), n=4 for Mx1-cre; Mettl3fl/fl (4m)). (h) 2’-7’-dichlorofluorescein diacetate (DCFDA) staining (ROS levels) by flow cytometry in HSCs 2-3 months after pIpC treatment (n=3 for control, n=3 for Mx1-cre; Mettl3fl/fl). (i) γH2AX staining by flow cytometry in HSCs 2-3 months after pIpC treatment (n=3 for control, n=3 for Mx1-cre; Mettl3fl/fl). All samples were from biologically independent animals. Values are shown as individual points with mean ± s.d. In (h) and (i), normalized geometric mean fluorescence intensity (MFI ± s.d.) values are shown. P values were determined by unpaired two-sided Student’s t-test.
Supplementary Fig. 3 HSCs preferentially require Mettl3 for differentiation in vitro compared with other haematopoietic progenitors.
(a) The proportion of large colonies (estimated >5,000 cells) out of all colonies formed from HSCs or LSK cells (n=6 for control, n=6 for Mx1-cre; Mettl3fl/fl). (b) Representative images of colonies from control and Mx1-cre; Mettl3fl/fl LSK cells. Experiment was repeated four times from biologically independent animals with similar results. Scale bars are 400um. (c) Representative genotyping image of colonies formed by single HSCs grouped by size. Experiment was repeated four times from biologically independent animals with similar results. Unprocessed gel scans are in Supplementary Fig. 7. (d) Quantification of the frequency of Gr1+ and Mac1+ cells by flow cytometry on colony cells from splenic HSCs in Mx1-cre; Mettl3fl/fl mice at least 14 days after pIpC treatment with prominent splenomegaly compared to controls (n=5 for control, n=5 for Mx1-cre; Mettl3fl/fl). (e) Number of colonies formed from control and Mx1-cre; Mettl3fl/fl cells of the indicated populations 10 days after pIpC treatment. 200 LSK cells, 50 MPP cells, 24 CD150-CD48+LSK and 24 CD150+CD48+LSK cells were plated per experiment. (For LSK and MPP: n=4 for control, n=4 for Mx1-cre; Mettl3fl/fl; for CD150-CD48+LSK and CD150+CD48+LSK: n=3 for control, n=3 for Mx1-cre; Mettl3fl/fl). (f) Quantification of the frequency of Gr1+ and Mac1+ cells by flow cytometry on colony cells from the indicated cell populations from mice 10 days after pIpC treatment. (MPP: n=7 for control, n=7 for Mx1-cre; Mettl3fl/fl; CD150-CD48+LSK: n=10 for control, n=8 for Mx1-cre; Mettl3fl/fl; CD150+CD48+LSK: n=7 for control, n=6 for Mx1-cre; Mettl3fl/fl;). All samples are from biologically independent animals. Values are shown as individual points with mean ± s.d. P values were determined by unpaired two-sided Student’s t-test.
Supplementary Fig. 4 HSCs, but not mature myeloid cells, require Mettl3 for function in vivo.
(a) Chimera levels of bone marrow HSCs 16 weeks after transplantation from Fig. 3a (n=7 for control, n=6 for Mx1-cre; Mettl3fl/fl ; samples were from independent recipients of two independent donor pairs from two experiments). (b) Representative genotyping images of Mx1-cre; Mettl3fl/fl donor-derived cells 20 weeks after transplantation from Fig. 3a showing incomplete deletion of Mettl3. Experiment was repeated twice with similar results. Unprocessed gel scans are in Supplementary Fig. 7. (c) Chimera levels of bone marrow HSCs 16 weeks after transplantation from Fig. 3b (n=9 for control, n=9 for Mx1-cre; Mettl3fl/fl ; samples were from independent recipients of two independent donor pairs from two experiments). (d) Representative genotyping images of Mx1-cre; Mettl3fl/fl donor-derived cells 12 weeks after pIpC treatment from Fig. 3c showing incomplete deletion of Mettl3. Experiment was repeated twice with similar results. Unprocessed gel scans are in Supplementary Fig. 7. (e) 1 million donor whole bone marrow cells from untreated Rosa26-CreER; Mettl3fl/fl or control mice were non-competitively transplanted into lethally irradiated recipient mice. Recipients with peripheral blood chimerism of >90% were treated with tamoxifen (TAM) 6 weeks after the transplantation. The frequency of donor HSCs 8 weeks after TAM is shown (n=4 for control, n=3 for Rosa26-CreER; Mettl3fl/fl ; samples were from independent recipients of two independent donor pairs from two experiments). (f) Schematic of experimental design (top). 500,000 donor whole bone marrow cells from untreated Rosa26-CreER; Mettl3fl/fl or control mice with 500,000 competitor cells were transplanted into lethally irradiated recipient mice. Recipients were treated with tamoxifen (TAM) 6 weeks after the transplantation. Multi-lineage peripheral blood chimera levels are shown as the percentage of original chimera levels up to 16 weeks after TAM treatment (n=12 for control, n=11 for Rosa26-CreER; Mettl3fl/fl ; samples were from independent recipients of three independent donor pairs from three experiments). (g) qPCR analysis of Mettl3 mRNA expression in cultured bone marrow macrophages (n=3 for control, n=3 for LysM-cre; Mettl3fl/fl). (h) Red blood cell and platelet peripheral blood counts (n=6 for control, n=6 for LysM-cre; Mettl3fl/fl). (i) Spleen cellularity (n=5 for control, n=5 for LysM-cre; Mettl3fl/fl). (j) Representative images of macrophages derived from control or LysM-cre; Mettl3fl/fl bone marrow. Experiment was repeated three times with similar results. Scale bars are 100um. All samples are from biologically independent animals. Values are shown as individual points with mean ± s.d. P values were determined by unpaired two-sided Student’s t-test.
Supplementary Fig. 5 meRIP-seq analysis identifies the direct mRNA targets of m6A in HSCs.
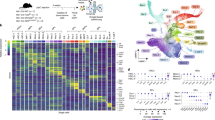
(a) Schematic of meRIP-seq workflow for low numbers of cells (such as HSCs). (b) Representative bioanalyzer graphs of meRIP-seq elution sample libraries after tagmentation generated using anti-m6A-antibody and IgG for immunoprecipitation. Experiment was repeated three times with similar results. (c) Volcano plot of meRIP-seq transcript expression differences between m6A-tagged and unbound fractions in Mx1-cre; Mettl3fl/fl (Mettl3Δ/Δ) HSCs 10 days after Cre induction. The x-axis specifies the log2 fold-changes (FC) and the y-axis specifies the log10 false discovery rate (FDR). Dashed vertical and horizontal lines indicate the filtering criteria (log2(FC)>1 or <-1 and FDR<0.05). Orange dots represent transcripts showing statistically significant differences between m6A-tagged and unbound factions, with select genes labelled. Data were from three biologically independent animals. The statistics was controlled for multiple testing by false discovery rate. (d) Comparison of m6A enrichment in wild-type (WT) HSCs vs Mettl3Δ/Δ HSCs. The x-axis specifies the log2 fold-change (FC) expression difference between m6A-tagged and unbound fractions in WT HSCs. The y-axis specifies the log2FC expression difference between m6A-tagged and unbound fractions in Mx1-cre; Mettl3fl/fl (Mettl3Δ/Δ) HSCs 10 days after Cre induction. In both axes, positive sums represent enrichment in the m6A fraction. All dots shown are transcripts found to be methylated in WT HSCs (log2(FC)>1 and FDR<0.05). Blue dots and select labelled genes represent transcripts found to have significantly different m6A enrichment in WT vs. Mettl3Δ/Δ (p<0.05). The dotted line represents equal m6A enrichment in WT vs. Mettl3Δ/Δ - dots below this line have decreased m6A enrichment in Mettl3Δ/Δ compared to WT HSCs. The statistics was controlled for multiple testing by false discovery rate. (e) Gene ontology analysis of biological processes enriched in m6A-tagged transcripts in WT HSCs. (f) meRIP-seq data showing that many components of the m6A pathway are targets of m6A in HSCs and this methylation is dependent on Mettl3. Data were from n=3 biologically independent samples for control and Mettl3Δ/Δ. Values are shown as mean ± s.d.
Supplementary Fig. 6 Loss of Mettl3 in HSCs does not cause major transcriptomic changes and forced expression of Myc rescues the HSC differentiation defects caused by Mettl3 deficiency.
(a) List of differentially expressed genes in control and Mx1-cre; Mettl3fl/fl HSCs 10 days after Cre induction by RNA-seq analysis (FDR<0.05). The statistics was controlled for multiple testing by false discovery rate. (b) Gene set enrichment analysis plots showing that Mettl3-deficient HSCs downregulate HSC signature, as determined by RNA-seq profiling of control and Mx1-cre; Mettl3fl/fl HSCs 10 days after pIpC treatment (n=3 for control and Mettl3Δ/Δ). Analysis was completed on gene list ranked by log10 FDR and fold-change sign. P values were determined by the GSEA algorithm after 1,000 permutations. The statistics was controlled for multiple testing by false discovery rate. (c) Diagram of overlap between 384 upregulated genes in Mettl3-deficient HSCs compared with controls (defined as p<0.05, fold>1.2) identified by RNA-seq and 2,073 methylated targets identified by meRIP-seq in wild-type HSCs. P value for enrichment was determined by hypergeometric probability test. (d) qPCR analysis of Myc expression by control and Mettl3-deficient HSCs 10-14 days after pIpC treatment (n=5 for control, n=6 for Mx1-cre; Mettl3fl/fl) and after overnight cytokine activation (right; n=5 for control, n=5 for Mx1-cre; Mettl3fl/fl). All samples were from biologically independent animals. Values are shown as mean ± s.d. P values were determined by paired two-sided Student’s t-test. (e) Additional gene set enrichment analysis plots of MYC targets, as determined by RNA-seq profiling of control and Mx1-cre; Mettl3fl/fl HSCs 10 days after pIpC treatment (n=3 for control and Mettl3Δ/Δ). Analysis was completed on gene list ranked by log10 FDR and fold-change sign. P values was determined by the GSEA algorithm after 1,000 permutations. The statistics was controlled for multiple testing by false discovery rate. (f) Representative FACS plots showing gating strategy for analysis of virally transduced colony cells derived from sorted HSCs from Mx1-cre; Mettl3fl/fl mice 10 days after pIpC treatment. In (a), (b), (c), (e), all sequencing data were from three independent animals of control and Mettl3Δ/Δ.
Supplementary Fig. 7 Uncropped gel images.
Unprocessed gel scans are provided for all DNA gel images in the study.
Supplementary information
Supplementary Information
Supplementary Figures 1–7, Supplementary Table titles/legends.
Supplementary Table 1
meRIP-seq identifies Mettl3-dependent m6A targets in HSCs.
Supplementary Table 2
RNA-seq identifies transcriptome differences between Mettl3-deficient and wild-type HSCs.
Supplementary Table 3
Statistics source data.
Rights and permissions
About this article
Cite this article
Lee, H., Bao, S., Qian, Y. et al. Stage-specific requirement for Mettl3-dependent m6A mRNA methylation during haematopoietic stem cell differentiation. Nat Cell Biol 21, 700–709 (2019). https://doi.org/10.1038/s41556-019-0318-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-019-0318-1
This article is cited by
-
Profiling the role of m6A effectors in the regulation of pluripotent reprogramming
Human Genomics (2024)
-
Age-related noncanonical TRMT6–TRMT61A signaling impairs hematopoietic stem cells
Nature Aging (2024)
-
A Mettl16/m6A/mybl2b/Igf2bp1 axis ensures cell cycle progression of embryonic hematopoietic stem and progenitor cells
The EMBO Journal (2024)
-
Therapeutic effect and transcriptome-methylome characteristics of METTL3 inhibition in liver hepatocellular carcinoma
Cancer Cell International (2023)
-
METTL3-mediated m6A methylation regulates ovarian cancer progression by recruiting myeloid-derived suppressor cells
Cell & Bioscience (2023)