Abstract
Haematopoietic stem cells (HSCs) are maintained by bone marrow niches in vivo1,2, but the ability of niche cells to maintain HSCs ex vivo is markedly diminished. Expression of niche factors by Nestin-GFP+ mesenchymal-derived stromal cells (MSCs) is downregulated upon culture, suggesting that transcriptional rewiring may contribute to this reduced HSC maintenance potential. Using an RNA sequencing screen, we identified five genes encoding transcription factors (Klf7, Ostf1, Xbp1, Irf3 and Irf7) that restored HSC niche function in cultured bone marrow-derived MSCs. These revitalized MSCs (rMSCs) exhibited enhanced synthesis of HSC niche factors while retaining their mesenchymal differentiation capacity. In contrast to HSCs co-cultured with control MSCs, HSCs expanded with rMSCs showed higher repopulation capacity and protected lethally irradiated recipient mice. Competitive reconstitution assays revealed an approximately sevenfold expansion of functional HSCs by rMSCs. rMSCs prevented the accumulation of DNA damage in cultured HSCs, a hallmark of ageing and replication stress. Analysis of the reprogramming mechanisms uncovered a role for myocyte enhancer factor 2c (Mef2c) in the revitalization of MSCs. These results provide insight into the transcriptional regulation of the niche with implications for stem cell-based therapies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
RNA-seq and ATAC-seq data have been deposited in the Gene Expression Omnibus under the accession number GSE112233. Source data for Figs. 1–4 and Supplementary Fig. 1–4 have been provided as Supplementary Table 6. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
Code availability
All codes used in this study are available from the corresponding author on reasonable request
References
Pinho, S. & Frenette, P. S. Haematopoietic stem cell activity and interactions with the niche. Nat. Rev. Mol. Cell Biol. https://doi.org/10.1038/s41580-019-0103-9(2019).
Morrison, S. J. & Scadden, D. T. The bone marrow niche for haematopoietic stem cells. Nature 505, 327–334 (2014).
Asada, N. et al. Differential cytokine contributions of perivascular haematopoietic stem cell niches. Nat. Cell Biol. 19, 214–223 (2017).
Khan, J. A. et al. Fetal liver hematopoietic stem cell niches associate with portal vessels. Science 351, 176–180 (2016).
Mendez-Ferrer, S. et al. Mesenchymal and haematopoietic stem cells form a unique bone marrow niche. Nature 466, 829–834 (2010).
Butler, J. M. et al. Endothelial cells are essential for the self-renewal and repopulation of Notch-dependent hematopoietic stem cells. Cell Stem Cell 6, 251–264 (2010).
Isern, J. et al. Self-renewing human bone marrow mesenspheres promote hematopoietic stem cell expansion. Cell Rep. 3, 1714–1724 (2013).
Pinho, S. et al. PDGFRα and CD51 mark human Nestin+ sphere-forming mesenchymal stem cells capable of hematopoietic progenitor cell expansion. J. Exp. Med. 210, 1351–1367 (2013).
Otto, F. et al. Cbfa1, a candidate gene for cleidocranial dysplasia syndrome, is essential for osteoblast differentiation and bone development. Cell 89, 765–771 (1997).
Oguro, H. et al. Differential impact of Ink4a and Arf on hematopoietic stem cells and their bone marrow microenvironment in Bmi1-deficient mice. J. Exp. Med. 203, 2247–2253 (2006).
Omatsu, Y., Seike, M., Sugiyama, T., Kume, T. & Nagasawa, T. Foxc1 is a critical regulator of haematopoietic stem/progenitor cell niche formation. Nature 508, 536–540 (2014).
Kunisaki, Y. et al. Arteriolar niches maintain haematopoietic stem cell quiescence. Nature 502, 637–643 (2013).
Ding, L., Saunders, T. L., Enikolopov, G. & Morrison, S. J. Endothelial and perivascular cells maintain haematopoietic stem cells. Nature 481, 457–462 (2012).
Wei, Q. & Frenette, P. S. Niches for hematopoietic stem cells and their progeny. Immunity 48, 632–648 (2018).
Itoh, K. et al. Reproducible establishment of hemopoietic supportive stromal cell lines from murine bone marrow. Exp. Hematol. 17, 145–153 (1989).
Flach, J. et al. Replication stress is a potent driver of functional decline in ageing haematopoietic stem cells. Nature 512, 198–202 (2014).
Beerman, I., Seita, J., Inlay, M. A., Weissman, I. L. & Rossi, D. J. Quiescent hematopoietic stem cells accumulate DNA damage during aging that is repaired upon entry into cell cycle. Cell Stem Cell 15, 37–50 (2014).
Rube, C. E. et al. Accumulation of DNA damage in hematopoietic stem and progenitor cells during human aging. PLoS One 6, e17487 (2011).
Rossi, D. J. et al. Deficiencies in DNA damage repair limit the function of haematopoietic stem cells with age. Nature 447, 725–729 (2007).
Walter, D. et al. Exit from dormancy provokes DNA-damage-induced attrition in haematopoietic stem cells. Nature 520, 549–552 (2015).
Yamazaki, S. et al. Nonmyelinating Schwann cells maintain hematopoietic stem cell hibernation in the bone marrow niche. Cell 147, 1146–1158 (2011).
Torisawa, Y. S. et al. Bone marrow-on-a-chip replicates hematopoietic niche physiology in vitro. Nat. Methods 11, 663–669 (2014).
Dhami, S. P., Kappala, S. S., Thompson, A. & Szegezdi, E. Three-dimensional ex vivo co-culture models of the leukaemic bone marrow niche for functional drug testing. Drug Discov. Today 21, 1464–1471 (2016).
Naldini, L. Gene therapy returns to centre stage. Nature 526, 351–360 (2015).
Kawamura, Y., Tanaka, Y., Kawamori, R. & Maeda, S. Overexpression of Kruppel-like factor 7 regulates adipocytokine gene expressions in human adipocytes and inhibits glucose-induced insulin secretion in pancreatic beta-cell line. Mol. Endocrinol. 20, 844–856 (2006).
Xu, G. et al. Expression of XBP1s in bone marrow stromal cells is critical for myeloma cell growth and osteoclast formation. Blood 119, 4205–4214 (2012).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinformatics 14, 128 (2013).
Pierce, H. et al. Cholinergic signals from the CNS regulate G-CSF-mediated HSC mobilization from bone marrow via a glucocorticoid signaling relay. Cell Stem Cell 20, 648–658.e4 (2017).
Bruns, I. et al. Megakaryocytes regulate hematopoietic stem cell quiescence through CXCL4 secretion. Nat. Med. 20, 1315–1320 (2014).
Lucas, D. et al. Chemotherapy-induced bone marrow nerve injury impairs hematopoietic regeneration. Nat. Med. 19, 695–703 (2013).
Wang, Z. & Ma’ayan, A. An open RNA-seq data analysis pipeline tutorial with an example of reprocessing data from a recent Zika virus study. F1000Res. 5, 1574 (2016).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Clark, N. et al. The characteristic direction: a geometrical approach to identify differentially expressed genes. BMC Bioinformatics 15, 79 (2014).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21–29 (2015).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
Zhang, Y. et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. 9, R137 (2008).
Li, Q., Brown, J. B., Huang, H. & Bickel, P. J. Measuring reproducibility of high-throughput experiments. Ann. Appl. Stat. 5, 1752–1779 (2011).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Acknowledgements
We thank C. Prophete and P. Ciero for technical assistance, L. Tesfa for assistance with cell sorting and S. Maqbool for RNA-seq. We thank L. Ding and S. J. Morrison for providing the Scf-GFP knock-in mice. We thank C.-J. Chang and E. E. Bouhassira for the FUWC-GW and pHIV-dTmt vectors. We thank T. Taniguchi (University of Tokyo, Japan) for the pCAGGS/Irf3 and pCAGGS/Irf7 vectors and S. Maeda (University of Kagoshima, Japan) for the pEF/Runx2 vectors. We thank J. F. Reidhaar-Olson and J. Shan for supplying overexpression and knockdown vectors. This work was supported by R01 grants from the US National Institutes of Health (NIH) (DK056638, HL069438, DK116312 and DK112976 to P.S.F., and U54HL127624 and U24CA224260 to A.M.). We are also grateful to the New York State Department of Health (NYSTEM Program) for shared facility (C029154) and research support (N13G-262) and the Leukemia and Lymphoma Society’s Translational Research Program. F.N. was supported by the Postdoctoral Fellowship for Research Abroad from the Japan Society for the Promotion of Science (JSPS). D.K.B. is partially supported by a NIH Training Grant (T32 GM007288). M.M. is a New York Stem Cell Foundation (NYSCF) Druckenmiller fellow. A.H.Z. was supported by a NIH Training Grant (T32 NS007098) and by a National Cancer Institute (NCI) predoctoral MD/PhD fellowship (F30 CA203446). P.E.B. is supported by a postdoctoral fellowship from Fonds de recherche du Québec-Santé (FRQS).
Author information
Authors and Affiliations
Contributions
F.N. designed the study, performed most of the experiments and analysed the data. D.K.B. performed the Mef2c and Klf7 overexpression experiments, helped with the co-culture experiments and analysed the data. Q.W. advised on experimental design and helped with making the list of candidate genes from the RNA-seq data. S.P. advised on experimental design and helped with the implantation experiments of hydroxyapatite scaffolds. M.M. advised on experimental design and helped with HSC imaging. A.H.Z. advised on experimental design. M.S. and J.M.G. analysed the ATAC-seq data. C.D.C. helped with vector cloning and virus production. Z.W. and A.M. analysed the RNA-seq data. P.E.B. and C.X. helped with improving the culture system. P.S.F. supervised the study. F.N. and P.S.F. interpreted the data and wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
F.N. and P.S.F. are co-authors on the U.S. provisional patent application (No 62/517,271) titled “Methods for expanding hematopoietic stem cells using revitalized mesenchymal stem cells”.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Fig. 1 Combination of four genes (OXII) or five genes (KOXII) increases niche activity in MSCs.
(a-c) The expression of BM niche genes (Cxcl12, Vcam1 and Angpt1) was assessed in each clone by real-time qPCR analysis. Statistical significance is showing comparison of the expression in each clone with that of empty vector-transduced control MSCs. Expression level was normalized with Actb and the mean expression level in control MSC was defined as 1. n = 3 biologically independent samples for each clone. Vcam1: F6, P = 0.03; J9, P = 0.006; K10, P = 0.002; L11, P = 0.007 (d) Isolated MSCs from Scf-GFP mice BM were cultured for 21 days and transduced with mix of lentivirus containing 4 genes (Ostf1, Xbp1, Irf3, and Irf7; OXII) or 4 genes minus the indicated factor, or 4 genes plus Klf7 (KOXII). On day 7 post transduction, re-emerged Scf-GFP+ cells were sorted and cell count was assessed after 14 days of culture. n = 3 biologically independent samples for each group. KOXII vs. OXII, P = 0.02 (e-i) The expression of the indicated virally integrated factors (Klf7, Ostf1, Xbp1, Irf3 and Irf7) was assessed by real-time qPCR analysis in control MSCs and 5 genes (Klf7, Ostf1, Xbp1, Irf3 and Irf7)-transduced rMSCs (KOXII). Expression level was normalized with Actb level and the mean expression level in control MSC was defined as 1. n = 3 biologically independent samples for Control, n = 3 biologically independent samples for KOXII in each gene. (j and k) Expression of bone marrow niche genes (Vcam1 and Angpt1) was assessed by real-time qPCR in control MSC, 4 genes (Ostf1, Xbp1, Irf3, and Irf7)-transduced MSC clone (OXII), and 5 genes (Klf7, Ostf1, Xbp1, Irf3 and Irf7)-transduced MSC clone (KOXII). Expression level was normalized with Actb level and the mean expression level in control MSC was defined as 1. n = 4 Control, n = 9 OXII, n = 12 KOXII for Vcam1 and n = 4 Control, n = 9 OXII, n = 15 KOXII for Angpt1. (all n represent biologically independent samples) (l and m) Expression of bone marrow niche genes (Scf and Cxcl12) was assessed by real-time qPCR in control MSC, Klf7-transduced MSC, and 5 gene (Klf7, Ostf1, Xbp1, Irf3, Irf7)-transduced MSC. Expression level was normalized to Actb and the mean expression level in control MSC was defined as 1. n = 2 biologically independent samples for Control, Klf7, and KOXII. Error bars, mean ± s.e.m. in (a-c and e-k), mean ± s.d. in (d, l, and m). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, n.s. (not significant); One-way ANOVA followed by Tukey’s multiple comparison test (a-d, j–m), two-tailed unpaired Student’s t test (e-i).
Supplementary Fig. 2 Characterization of revitalized KOXII MSCs.
(a, left panel) PDGFRα+ CD51+ cell frequency in cultured MSCs transduced with empty vector (control) or 5 genes (Klf7, Ostf1, Xbp1, Irf3 and Irf7; KOXII)-transduced revitalized MSCs (rMSC). (a, right panel) FACS analysis plot of representative sample from (a, left panel). n = 4 biologically independent samples for Control and rMSC. (b) MSC marker (CD44, Sca1, CD105, LepR and CD140b) expression in control MSC or rMSC. n = 3 biologically independent samples for Control and rMSC in each marker. (c) Self-renewing sphere-forming capacity was assessed by plating control MSCs or rMSC at clonal densities. (d) Phase-contrast images of self-renewing spheres in (c). n = 4 biologically independent samples for Control and rMSC. (e-g, upper panel) Multilineage differentiation capacity of control MSCs and KOXII-transduced rMSCs. Fully differentiated phenotypes of control and rMSCs were tested by Oil Red O (adipogenic; e), Alizarin Red S (osteogenic; f), and Alcian Blue (chondrogenic; g) stainings. (e-g, lower panel) Differentiation kinetic of control MSCs and KOXII-transduced rMSCs was evaluated by real-time qPCR for the expression of adipogenic specific gene (Pparg) at days 0, 14, osteogenic specific gene (Sp7) at days 0, 20, or chondrogenic specific gene (Sox9) at days 0, 21. Expression level was normalized with Actb and the mean expression level of day 0 control MSC was defined as 1. n = 3 biologically independent samples for each group in Pparg, Sp7 and Sox9. Scale bars: (d-f) 50 μm; (g) 1 mm. Error bars, mean ± s.d. in (a-c), mean ± s.e.m. in (e-g). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001; two-sided unpaired Student’s t test (a–c), One-way ANOVA followed by Tukey’s multiple comparison test (e-g).
Supplementary Fig. 3 rMSCs expand HSCs and maintain their genomic integrity.
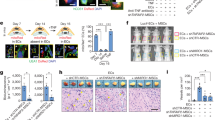
(a) Lin− BM cells were co-cultured with the MS-5 stromal cell line, empty vector-transduced MSC (control), KOXII-transduced rMSC, or cell-free media, with TPO (10 ng ml−1) and in the presence or absence of SCF (20 ng ml−1). Quantification of LSK (Lin− Sca1+ cKit+) and Lin− cell numbers was assessed by FACS analysis 6 days post co-culture. n = 3 biologically independent samples for MS-5, Control, and rMSC in LSK and Lin−. LSK: rMSC+SCF vs. control+SCF, P = 0.02 (b) Lin-depleted bone marrow cells were cultured for 6 days with fresh StemSpan or StemSpan conditioned over rMSC for 24 hr and then filtered (Conditioned). In both cases, TPO (10 ng ml−1) and SCF (20 ng ml−1) were added immediately before culture of lineage-depleted bone marrow cells. n = 3 independent samples for all groups. (c) Survival curve of lethally irradiated recipient mice (CD45.2) transplanted with freshly isolated 40,000 BM mononuclear cells (BMNCs) from C57BL/6 mice (CD45.1), or transplanted with whole CD45+ cells derived from co-culture of 40,000 C57BL/6 BMNCs (CD45.1) with control MSCs, KOXII-transduced rMSCs, or without stromal cells, by non-competitive transplantation. n = 8 mice for each group. rMSC vs. control, P = 0.03; vs. stroma(-), P = 0.045; vs. fresh, P = 0.045 (d) Competitive repopulating unit (CRU) assay using limiting numbers of fresh whole BMNCs, or whole BMNCs co-cultured with rMSCs. Freshly isolated whole BMNCs were immediately used for BM transplantation, or co-cultured with rMSCs in the serum-free medium supplemented with SCF (20 ng ml−1) and TPO (10 ng ml−1) for 6 days, and then a fraction of the cultured cells corresponding to the indicated number (1, 2.5, 5, 12.5 and 25×103) of initial BM cells was transplanted. Percent chimerism of donor cells in the recipient peripheral blood was determined at 16 weeks after transplantation. Mice with chimerism > 1% in all three lineages (myeloid, B and T cells) were considered successfully engrafted and the others were defined as negative. The frequency of HSCs was calculated using L-Calc software (STEMCELL Technologies). The number of BM cells injected, the proportion of engrafted mice, and estimated frequency of functional HSCs are summarized in the Table. (e) Quantification of the DNA damage markers gH2AX in HSCs co-cultured with empty vector-transduced MSC (control), KOXII-transduced rMSC, or without stromal cells in the serum-free medium supplemented with SCF (20 ng ml−1) and TPO (10 ng ml−1) for 6 days. gH2AX foci were counted in each HSC and scored. n calculated as mean of total 150 HSCs in Stroma−, 150 HSCs in Control, 150 HSCs in rMSC setting scored from n = 3 biologically independent samples for each group. (f) CRU assay using limiting numbers of CD34+ cells from human cord blood, or human cord blood CD34+ cells co-cultured with murine rMSCs. Freshly isolated CD34+ cells from human cord blood were immediately used for BM transplantation into NOD-scid Il2rg−/− (NSG) immunocompromised mice, or co-cultured with rMSCs for 6 days, and then a fraction of the cultured cells corresponding to the indicated number (0.2, 1, 3 and 5 × 103) of initial cord blood cells was transplanted into NSG mice. Percent chimerism of donor cells in the recipient peripheral blood was determined at 16 weeks after transplantation. Mice with donor chimerism > 1% were considered successfully engrafted and the others were scored as negative. n = 5 mice for each group. Error bars, mean ± s.d. in (a, b, and e). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, n.s. (not significant); One-way ANOVA followed by Tukey’s multiple comparison test (a). Two-sided log-rank analysis was used for the Kaplan-Meier survival curves in (c). Two-sided unpaired Student’s t test (b and e).
Supplementary Fig. 4 Depletion of Mef2c alters the expression of niche factors in rMSCs.
(a) Enriched pathways of 626 peaks overlapping between 2 groups of genes with increased accessibilities (open) and genes with decreased accessibilities (closed) in ATAC-seq. Pathway analysis was obtained from Enrichr53,54. n = 3 for each group. P-values determined by one-sided Fisher’s exact test for 230 pathways, with no correction for multiple comparisons. (b) Heat map of expression levels of Mef2c in sorted non-cultured CD45− TER119− CD31− Scf-GFP− cells, non-cultured CD45− TER119− CD31− Scf-GFP+ cells, empty vector-transduced control MSCs, and rMSCs. The values of log-transformed reads per kilobase per million mapped reads (RPKM) obtained from RNA-seq were visualized using GraphPad Prism7. (n = 3 samples per group) (c and d) Expression of Mef2c and bone marrow niche genes (Scf, Cxcl12, Vcam1 and Angpt1) were determined by real-time qPCR in the shCntrl-transduced rMSCs and shMef2c-transduced rMSCs. Expression level was normalized with Actb and the mean expression level in shCntrl-transduced rMSCs was defined as 1. n = 3 biologically independent samples for shCntrl and shMef2c in each gene. Scf, P = 0.01; Cxcl12, P = 0.03; Vcam1, P = 0.004; Angpt1, P = 0.07 (e) Expression of Mef2c was assessed by real-time qPCR in empty vector- and Mef2c-transduced rMSCs. Expression level was normalized to Actb. n = 3 biologically independent samples for each group. (f) Lin− BM cells were co-cultured with empty vector- and Mef2c-transduced rMSCs with TPO (10 ng ml−1) and SCF (20 ng ml−1). Quantification of HSCs was assessed by FACS analysis 6 days post co-culture. n = 3 biologically independent samples for rMSC+mock and rMSC+Mef2c. P = 0.0099. Error bars, mean ± s.e.m. in (c, d, and f) ± s.d. in (e). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001; two-tailed unpaired Student’s t test (c–f).
Supplementary information
Supplementary Information
Supplementary Figs. 1–4, Supplementary Table titles/legends.
Supplementary Table 1
This table contains a list of open chromatin regions in the ATAC-seq analysis of the four cell populations.
Supplementary Table 2
This table contains a list of 626 overlapping peaks between two groups of peaks with increased accessibilities (open) and peaks with decreased accessibilities (closed) in the ATAC-seq analysis of the four cell populations as shown in Fig. 4e.
Supplementary Table 3
This table contains a quality control summary of the RNA-seq analysis.
Supplementary Table 4
This table contains a quality control summary of the ATAC-seq analysis.
Supplementary Table 5
This table contains the sequences of all DNA oligomers used in this study.
Supplementary Table 6
Statistics source data.
Rights and permissions
About this article
Cite this article
Nakahara, F., Borger, D.K., Wei, Q. et al. Engineering a haematopoietic stem cell niche by revitalizing mesenchymal stromal cells. Nat Cell Biol 21, 560–567 (2019). https://doi.org/10.1038/s41556-019-0308-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-019-0308-3
This article is cited by
-
Influential factors for optimizing and strengthening mesenchymal stem cells and hematopoietic stem cells co-culture
Molecular Biology Reports (2024)
-
Culture-expanded mesenchymal stromal cell therapy: does it work in knee osteoarthritis? A pathway to clinical success
Cellular & Molecular Immunology (2023)
-
Ageing and rejuvenation of tissue stem cells and their niches
Nature Reviews Molecular Cell Biology (2023)
-
WHIM Syndrome-linked CXCR4 mutations drive osteoporosis
Nature Communications (2023)
-
The Intercellular Communication Between Mesenchymal Stromal Cells and Hematopoietic Stem Cells Critically Depends on NF-κB Signalling in the Mesenchymal Stromal Cells
Stem Cell Reviews and Reports (2022)