Abstract
Vertebrate tissues exhibit mechanical homeostasis, showing stable stiffness and tension over time and recovery after changes in mechanical stress. However, the regulatory pathways that mediate these effects are unknown. A comprehensive identification of Argonaute 2-associated microRNAs and mRNAs in endothelial cells identified a network of 122 microRNA families that target 73 mRNAs encoding cytoskeletal, contractile, adhesive and extracellular matrix (CAM) proteins. The level of these microRNAs increased in cells plated on stiff versus soft substrates, consistent with homeostasis, and suppressed targets via microRNA recognition elements within the 3′ untranslated regions of CAM mRNAs. Inhibition of DROSHA or Argonaute 2, or disruption of microRNA recognition elements within individual target mRNAs, such as connective tissue growth factor, induced hyper-adhesive, hyper-contractile phenotypes in endothelial and fibroblast cells in vitro, and increased tissue stiffness, contractility and extracellular matrix deposition in the zebrafish fin fold in vivo. Thus, a network of microRNAs buffers CAM expression to mediate tissue mechanical homeostasis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
RNA-seq and small RNA-seq data for endothelial cells that support the findings of this study have been deposited in the GEO under accession codes GSE99686 and GSE11021. Small RNA-seq data for HDF have been deposited in the GEO under accession code GSE123008. HITS-CLIP data have been deposited in the Sequence Read Archive (SRA) under accession code PRJNA507245. Proteomics data have been uploaded to the PRIDE depository (PXD011882) and results are summarized in Supplementary Table 5. Source data for all figures and supplementary figures have been provided as Supplementary Table 8.
References
Cyron, C. J. & Humphrey, J. D. Growth and remodeling of load-bearing biological soft tissues. Meccanica 52, 645–664 (2017).
Gilbert, P. M. & Weaver, V. M. Cellular adaptation to biomechanical stress across length scales in tissue homeostasis and disease. Semin. Cell Dev. Biol. 67, 141–152 (2017).
Humphrey, J. D., Dufresne, E. R. & Schwartz, M. A. Mechanotransduction and extracellular matrix homeostasis. Nat. Rev. Mol. Cell Biol. 15, 802–812 (2014).
Humphrey, J. D. Vascular adaptation and mechanical homeostasis at tissue, cellular, and sub-cellular levels. Cell Biochem. Biophys. 50, 53–78 (2008).
Seki, E. & Brenner, D. A. Recent advancement of molecular mechanisms of liver fibrosis. J. Hepatobiliary Pancreat. Sci. 22, 512–518 (2015).
Huang, S. & Ingber, D. E. Cell tension, matrix mechanics, and cancer development. Cancer Cell 8, 175–176 (2005).
Sun, Z., Guo, S. S. & Fassler, R. Integrin-mediated mechanotransduction. J. Cell Biol. 215, 445–456 (2016).
Pasquinelli, A. E. MicroRNAs and their targets: recognition, regulation and an emerging reciprocal relationship. Nat. Rev. Genet. 13, 271–282 (2012).
Hafner, M. et al. Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell 141, 129–141 (2010).
Herranz, H. & Cohen, S. M. MicroRNAs and gene regulatory networks: managing the impact of noise in biological systems. Genes Dev. 24, 1339–1344 (2010).
Tsang, J., Zhu, J. & van Oudenaarden, A. MicroRNA-mediated feedback and feedforward loops are recurrent network motifs in mammals. Mol. Cell 26, 753–767 (2007).
Kasper, D. M. et al. MicroRNAs establish uniform traits during the architecture of vertebrate embryos. Dev. Cell 40, 552–565.e5 (2017).
Chi, S. W., Zang, J. B., Mele, A. & Darnell, R. B. Argonaute HITS-CLIP decodes microRNA–mRNA interaction maps. Nature 460, 479–486 (2009).
Byfield, F. J., Reen, R. K., Shentu, T. P., Levitan, I. & Gooch, K. J. Endothelial actin and cell stiffness is modulated by substrate stiffness in 2D and 3D. J. Biomech. 42, 1114–1119 (2009).
Grimson, A. et al. MicroRNA targeting specificity in mammals: determinants beyond seed pairing. Mol. Cell 27, 91–105 (2007).
Saphirstein, R. J. & Morgan, K. G. The contribution of vascular smooth muscle to aortic stiffness across length scales. Microcirculation 21, 201–207 (2014).
Mullokandov, G. et al. High-throughput assessment of microRNA activity and function using microRNA sensor and decoy libraries. Nat. Methods 9, 840–846 (2012).
Kamata, M., Liang, M., Liu, S., Nagaoka, Y. & Chen, I. S. Live cell monitoring of hiPSC generation and differentiation using differential expression of endogenous microRNAs. PLoS ONE 5, e11834 (2010).
Discher, D. E., Janmey, P. & Wang, Y. L. Tissue cells feel and respond to the stiffness of their substrate. Science 310, 1139–1143 (2005).
Kim, Y. K., Kim, B. & Kim, V. N. Re-evaluation of the roles of DROSHA, Export in 5, and DICER in microRNA biogenesis. Proc. Natl Acad. Sci. USA 113, E1881–E1889 (2016).
Dupont, S. et al. Role of YAP/TAZ in mechanotransduction. Nature 474, 179–183 (2011).
Kumar, A. et al. Talin tension sensor reveals novel features of focal adhesion force transmission and mechanosensitivity. J. Cell Biol. 213, 371–383 (2016).
Bassett, A. R. et al. Understanding functional miRNA–target interactions in vivo by site-specific genome engineering. Nat. Commun. 5, 4640 (2014).
Shi-Wen, X., Leask, A. & Abraham, D. Regulation and function of connective tissue growth factor/CCN2 in tissue repair, scarring and fibrosis. Cytokine Growth Factor Rev. 19, 133–144 (2008).
Kawakami, A., Fukazawa, T. & Takeda, H. Early fin primordia of zebrafish larvae regenerate by a similar growth control mechanism with adult regeneration. Dev. Dyn. 231, 693–699 (2004).
Mathew, L. K. et al. Comparative expression profiling reveals an essential role for raldh2 in epimorphic regeneration. J. Biol. Chem. 284, 33642–33653 (2009).
Mateus, R. et al. In vivo cell and tissue dynamics underlying zebrafish fin fold regeneration. PLoS ONE 7, e51766 (2012).
Cifuentes, D. et al. A novel miRNA processing pathway independent of Dicer requires Argonaute2 catalytic activity. Science 328, 1694–1698 (2010).
Amelio, I. et al. miR-24 triggers epidermal differentiation by controlling actin adhesion and cell migration. J. Cell Biol. 199, 347–363 (2012).
Nechiporuk, A. & Keating, M. T. A proliferation gradient between proximal and msxb-expressing distal blastema directs zebrafish fin regeneration. Development 129, 2607–2617 (2002).
Hasegawa, T. et al. Transient inflammatory response mediated by interleukin-1β is required for proper regeneration in zebrafish fin fold. eLife 6, e22716 (2017).
Mateus, R. et al. Control of tissue growth by Yap relies on cell density and F-actin in zebrafish fin regeneration. Development 142, 2752–2763 (2015).
Mori, M. et al. Hippo signaling regulates microprocessor and links cell-density-dependent miRNA biogenesis to cancer. Cell 156, 893–906 (2014).
Chaulk, S. G., Lattanzi, V. J., Hiemer, S. E., Fahlman, R. P. & Varelas, X. The Hippo pathway effectors TAZ/YAP regulate dicer expression and microRNA biogenesis through Let-7. J. Biol. Chem. 289, 1886–1891 (2014).
Davis, B. N., Hilyard, A. C., Lagna, G. & Hata, A. SMAD proteins control DROSHA-mediated microRNA maturation. Nature 454, 56–61 (2008).
Felix, M. A. & Wagner, A. Robustness and evolution: concepts, insights and challenges from a developmental model system. Heredity (Edinb.) 100, 132–140 (2008).
Mouw, J. K. et al. Tissue mechanics modulate microRNA-dependent PTEN expression to regulate malignant progression. Nat. Med. 20, 360–367 (2014).
Liu, G. et al. miR-21 mediates fibrogenic activation of pulmonary fibroblasts and lung fibrosis. J. Exp. Med. 207, 1589–1597 (2010).
Cushing, L. et al. miR-29 is a major regulator of genes associated with pulmonary fibrosis. Am. J. Respir. Cell Mol. Biol. 45, 287–294 (2011).
Herrera, J. et al. Dicer1 deficiency in the idiopathic pulmonary fibrosis fibroblastic focus promotes fibrosis by suppressing microRNA biogenesis. Am. J. Respir. Crit. Care Med. 198, 486–496 (2018).
Parker, M. W. et al. Fibrotic extracellular matrix activates a profibrotic positive feedback loop. J. Clin. Invest. 124, 1622–1635 (2014).
Pandit, K. V. & Milosevic, J. MicroRNA regulatory networks in idiopathic pulmonary fibrosis. Biochem. Cell Biol. 93, 129–137 (2015).
Wynn, T. A. & Ramalingam, T. R. Mechanisms of fibrosis: therapeutic translation for fibrotic disease. Nat. Med. 18, 1028–1040 (2012).
Liu, J. et al. Argonaute2 is the catalytic engine of mammalian RNAi. Science 305, 1437–1441 (2004).
McCall, M. N. et al. MicroRNA profiling of diverse endothelial cell types. BMC Med. Genomics 4, 78 (2011).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Kozomara, A. & Griffiths-Jones, S. miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 42, D68–D73 (2014).
Agarwal, V., Bell, G. W., Nam, J. W. & Bartel, D. P. Predicting effective microRNA target sites in mammalian mRNAs. eLife 4, e05005 (2015).
Aranguren, X. L. et al. Unraveling a novel transcription factor code determining the human arterial-specific endothelial cell signature. Blood 122, 3982–3992 (2013).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Bodenhofer, U., Bonatesta, E., Horejs-Kainrath, C. & Hochreiter, S. msa: an R package for multiple sequence alignment. Bioinformatics 31, 3997–3999 (2015).
Elosegui-Artola, A. et al. Mechanical regulation of a molecular clutch defines force transmission and transduction in response to matrix rigidity. Nat. Cell Biol. 18, 540–548 (2016).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
McCarthy, D. J., Chen, Y. & Smyth, G. K. Differential expression analysis of multifactor RNA-seq experiments with respect to biological variation. Nucleic Acids Res. 40, 4288–4297 (2012).
Goeminne, L. J., Gevaert, K. & Clement, L. Peptide-level robust ridge regression improves estimation, sensitivity, and specificity in data-dependent quantitative label-free shotgun proteomics. Mol. Cell. Proteomics 15, 657–668 (2016).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Smyth, G. K. Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, Article3 (2004).
Berginski, M. E. & Gomez, S. M. The Focal Adhesion Analysis Server: a web tool for analyzing focal adhesion dynamics. F1000Res. 2, 68 (2013).
Gutierrez, E. & Groisman, A. Measurements of elastic moduli of silicone gel substrates with a microfluidic device. PLoS ONE 6, e25534 (2011).
Han, S. J., Oak, Y., Groisman, A. & Danuser, G. Traction microscopy to identify force modulation in subresolution adhesions. Nat. Methods 12, 653–656 (2015).
Kapacee, Z. et al. Tension is required for fibripositor formation. Matrix Biol. 27, 371–375 (2008).
Narayanan, A. et al. In vivo mutagenesis of miRNA gene families using a scalable multiplexed CRISPR/Cas9 nuclease system. Sci. Rep. 6, 32386 (2016).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Le Guyader, D. et al. Origins and unconventional behavior of neutrophils in developing zebrafish. Blood 111, 132–141 (2008).
Sneddon, I. N. The relation between load and penetration in the axisymmetric boussinesq problem for a punch of arbitrary profile. Int. J. Eng. Sci. 3, 47–57 (1965).
Uren, P. J. et al. Site identification in high-throughput RNA–protein interaction data. Bioinformatics 28, 3013–3020 (2012).
Acknowledgements
We thank M. Cavanaugh for help with zebrafish husbandry. We thank the Wellcome Trust Centre for Cell-Matrix Research for technical support with tissue culture assays. We thank M. Trombly and A. Andersen, Life Science Editors, for proofreading the manuscript, and J. Humphrey and V. Greco for critical reading. We thank B. Coon for assistance during the preparation of the CRISPR–Cas9 experiment. We thank J. Lu’s laboratory for providing miRNA reporter lentiviral plasmid. We thank A. Giraldez’s laboratory (Genetics Department, Yale University) for providing the zebrafish ago2 mutant. J.S. was funded by a Biotechnology and Biological Sciences Research Council (BBSRC) David Phillips Fellowship (BB/L024551/1). D.M.K. was supported by F32HL132475, U54DK106857 and 1K99HL141687. V.M. was partially supported by a studentship from the Sir Richard Stapley Educational Trust. Mass spectrometry was carried out at the Wellcome Trust Centre for Cell-Matrix Research (203128/Z/16/Z) Proteomics Core Facility. This work was supported by the USPHS grant RO1 GM47214 to M.S. and RO1 HL130246 and AHA 17GRNT33460426 to S.N.
Author information
Authors and Affiliations
Contributions
S.N. and M.S. conceived the project. A.M. and T.D. performed the experiments, analysed the data and A.M. developed the computational data analysis. W.A. performed and analysed the zebrafish fin-fold regeneration experiments. L.C.B. performed and analysed the experiments in Supplementary Figs. 2b,c, 3b–d and 6. S.J.A. performed the cell culture experiments in Supplementary Fig. 3a. N.B. and C.J. performed the 3D cell culture experiments and analysed the data. D.L., M.Gu, J.Z. and M.Gerstein developed the sequence data processing and analysed (mapping and peak calling) the AGO2-HITS-CLIP experiment. V.M. and J.S. performed the mass spectrometry proteomic analysis. D.M.K. made the miRNA and RNA-seq libraries. A.M., T.D., M.S. and S.N. designed the experiments. M.S. and S.N. wrote the manuscript. All authors edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 AGO2-HITS-CLIP.
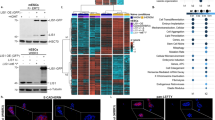
(a) Top left, SDS-PAGE of RNA-AGO2 or control IPs with nonspecific IgG were transferred to nitrocellulose and exposed to X-Ray film to reveal P32-RNAs bound to AGO2 (dotted square). P32 RNAs were isolated (dotted line) from nitrocellulose membranes and cDNAs generated with specific primers containing an Illumina barcode for high throughput sequencing (Methods). AGO2-mRNA complexes appear as ~200 nt bands (70 nt AGO-mRNA + 120 nt Illumina primer), while AGO2-miRNAs are ~150 nt (30 nt miRNA+120 nt Illumina primer). DNA MW markers are indicated in the left lane of the gel. Bottom left, Integrative Genomics Viewer display of HITS-CLIP reads. Right, computational pipeline to identify reads from miRNAs and mRNAs in AGO2 IPs. Three replicates for each cell type (HUAEC and HUVEC) were processed as described in Fig.1a. 68 million (M) total reads were analyzed. Upon removal of Illumina adapters and duplicates from PCR-based library preparation, reads were mapped to the human genome (UCSC hg19), resulting in 1M unique reads for each cell type. Reads were mapped to miRBase to identify miRNAs and processed using Piranha software to identify significant AGO2-binding sites (peaks) (methods). (b) Left, Bar graph of the Log(FDR) for the significant enriched Gene Ontology terms resulting from the HITS-CLIP assay and identified by DAVID software. Right, Bar graph of the Log(FDR) for the significant enriched Gene Ontology terms resulting from the microarray analysis of endothelial cells in culture versus freshly isolated.
Supplementary Figure 2 Validation of AGO2 CRISPR/Cas9 targeting in HUVECs.
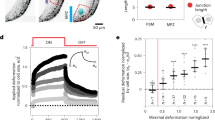
(a) Quantification of elastic modulus of PDMS gels by compression testing on an instron 5848 (bar = mean, n = 2 independent gels per condition plotted as dots). Gels were compressed with a cylindrical indenter to 10% strain at 0.1%/s and allowed to stress-relax for 150 seconds, modulus was measured at equilibrium. Quantification of Fibronectin staining intensity on the two PDMS gels with representative images showing uniform Fibronectin deposition on the gel surface (mean ± SEM, n = 14 fields of view per stiffness, ns = not significant, p = 0.2167, unpaired two-sided t-test, scale bar = 10 µm). (b) Schematics showing the experimental approach to generate AGO2 CRISPR/Cas9 mutant cells in primary human and mouse cells. (c) Left, Representative western blot of three independent replicates showing reduced AGO2 (~92 kDa) levels in HUVECs with a lentivirus co-expressing Cas9 and AGO2 gRNAs, or a no-target gRNA (control, has no homology to any known mammalian gene). The bar graphs show quantification of the proteins normalized to GAPDH (mean ± SEM; n = 3 biologically independent experiments, **** p < 0.0001, unpaired two-sided t-test). Right, qRT-PCR analysis of endothelial specific miRNAs in HUVECs infected with Cas9 and gRNA targeting AGO2 or no-target gRNA (control) at 7 days post infection (dpi). Results are shown normalized by one control for each microRNA (mean ± SEM; n = 3 biologically independent experiments). (d) Left, Schematics of CAM Sensor-Seq strategy. Sensor-seq lentivirus library was titrated to allow single vector copy integration and expression of physiological amounts of each GFP/mCherry transcript per cell. HUVECs were seeded on 3 and 30 kPa PDMS substrates for 48 h and analyzed as described. Right, bar plot representing distribution obtained upon FACS analysis of 10,000 wild-type or AGO2 gRNA mutant HUVECs infected with CAM MRE Sensors and treated as indicated (mean ± SEM; n = 9 biologically independent experiments for control 3 kPa, control 30 kPa; n = 8 biologically independent experiments miR125 and empty sensor; n=3 biologically independent experiments for AGO 3 kPa and AGO 30 kPa, p values indicated on plot, two-way ANOVA with Fisher’s LSD). mCherry/GFP ratio of CAM MRE Sensors increase in AGO2 mutant ECs, as the cell distribution is shifted significantly toward the Empty-Sensor control bin in both 3 and 30 kPa stiffness conditions. Source data can be found in Supplementary Table 8. Unprocessed blots for 2c can be found in Supplementary Figure 8.
Supplementary Figure 3 Validation of AGO2 or DROSHA loss of function in HUVECs.
(a) Left, Cell spread area using fibronectin coated polyacrylamide gels over a wider range of stiffness display results consistent with results from PDMS gel experiments (mean ± SEM, NT n = 43, 52, 78, 79, 89, 59 cells from soft to stiff, AGO2 n = 52, 52, 73, 63, 72, 54 cells from soft to stiff, * p = 0.0250, **** p < 0.0001, two-way ANOVA with Sidak’s multiple comparisons test). Right: Box plot representing HUVEC cell area for cells infected with AGO2 gRNA and non-targeting control seeded on fibronectin coated 30 kPa PDMS gels (box plots indicate minimum, maximum, median and quartiles, dots show single cell values, Control n = 440, AGO2 n = 289, **** p < 0.0001, unpaired two-sided t-test). (b) Top, representative immunofluorescence images of HUVECs after infection with pLentiCRISPR virus directed at AGO2 or a non-targeting control seeded on fibronectin coated 3 kPa PDMS gels (scale bar = 50 µm). Bottom: Quantification of AGO2 expression in n = 100 Control and n = 112 AGO2 cells. Intensity of the staining in each cell is normalized for the average intensity of all AGO2 cells and transformed in %. Cell area based on F-ACTIN staining or nuclear to cytoplasmic ratio of YAP/TAZ are plotted as individual value per cell. Highlighted in red cells infected with gRNA targeting AGO2, in black with no-target gRNA and in blue with gRNA targeting AGO2 expressing ~50% expression of AGO2 (~92 kDa) compared to control cells. (c) Western blot and quantification of DROSHA (~150 kDa) after knockdown with two distinct shRNAs delivered via lentivirus (5 days post infection, pSMART Dharmacon), normalized to GAPDH. Source data can be found in Supplementary Table 8. Unprocessed blots for 3c can be found in Supplementary Figure 8.
Supplementary Figure 4 Mutagenesis and targeting of CAM MREs.
(a) MiSeq 2x250 analysis of PCR amplicons derived from genomic DNA of HUVEC mutated with the indicated CAM MRE gRNAs. MREs are highlighted with red boxes, PAMs are highlighted with black boxes. Bar blot represent the % wild-type and mutated CAM 3’UTR sequences. Only mutations with a frequency > of 2% are represented. MSA = Multiple Sequence Alignment; nt = nucleotide. Numbers indicate the mutations as nt inserted (+) or deleted (Δ). (b) Left, Western blot of the CTGF protein (~37 kDa) normalized to β-ACTIN at 4 days post transfection. Right, box plots representing the quantification of mechanical parameters as in Fig. 4c of HUVECs transfected with miRTP (miRNA target protector) directed at CTGF 3’UTR for 4 days. miRTP is a single-stranded, modified RNA oligonucleotide that blocks a miRNA interaction with an individual MRE. Negative control target protector (QIAGEN) has no homology to any known mammalian gene. Box plots indicate minimum, maximum, median and quartiles. Cell spread area (miRTP Neg n = 109 cells, miRTP CTGF n = 103 cells, **p < 0.004, unpaired two-sided t-test), YAP nucelar/cytosol ratio (miRTP Neg n = 98 cells, miRTP CTGF n = 101 cells, **** p < 0.0001, unpaired two-sided t-test), and total force per cell (miRTP Neg n = 13 cells, miRTP CTGF n = 15 cells, * p = 0.0398, unpaired two-sided t-test). Source data can be found in Supplementary Table 8. Unprocessed blots for 4b can be found in Supplementary Figure 8.
Supplementary Figure 5 Analysis of miRNAs and AGO2 loss of function in fibroblast.
(a) DEG for miRNAs at 30 kPa vs. 3 kPa (left) and table with the results (right top). Scatterplots show miRNA in HUVEC and HDF. miRNAs are plotted following a decreasing DEG in both graphs. The rug plot shows the localization of the miRNA targeting CAM-MRE (CAM targeting miRNAs). Each miRNA in the EC rug plot is connected with the same miRNA in HDF plot. miRNAs with similar behavior in both conditions (overexpression or down-regulation at 30 vs 3 kPa in EC and HDF) are 58%, compared to 51% of total miRNAs with same behavior (p-Value = 0.1946). Within the miRNA group with similar behavior, 72% are up regulated on 30 vs. 3 kPa (against a total trend of 56% [Z-score pValue = 0.006432]). Bottom right, Venn diagram representing the overlap between CAM targeting miRNAs with a cut off of |log2(FC)| > 0.5 in EC (Blue circle) and HDF (Yellow circle). With a proportion of 31% upregulated CAM targeting miRNAs [17/54 in green sector] against a proportion of 10% downregulated [3/31 in red sector] the Z score proportion test reveals a significant enrichment of common upregulated miRNAs (two tail Z-score test pValue = 0.04385). (b) Left, western blot of 3 experiments showing AGO2 in HDFs infected with Cas9 and AGO2 gRNAs or a non-targeting gRNA (control) at 7 dpi with quantification of AGO2 normalized to GAPDH (n = 3 experiments, **** p < 0.0001, unpaired two-sided t-test). Right, box plot representing HDF cell area for cells infected with AGO2 gRNA and non-targeting control seeded on fibronectin-coated 30 kPa PDMS gels (box plot indicates minimum, maximum, median and quartiles, dots show single cell analysis, Control n = 311 cells AGO2 n = 264 cells, **p = 0.0009, unpaired two-sided t-test). (c) Representative blot of Ago2 levels in mouse skin fibroblast for 3D culture infected with gRNAs targeting Ago2 and no-target gRNA. Blots were obtained with Odyssey system from Licor to quantify near-IR fluorescence emitted by the secondary antibodies. Bar plots indicate densitometry measurements of Ago2 protein normalized for β-Actin at 7 dpi (mean ± SEM, Control n = 4, AGO2 n = 3, *p = 0.0261). Source data can be found in Supplementary Table 8. Unprocessed blots for 5b can be found in Supplementary Figure 8.
Supplementary Figure 6 Reduced levels of Ago2 in mz ago2 -/- zebrafish model.
(a) Left, Confocal lateral view of whole mount zebrafish embryos at 48 hpf. Wild-Type and ago2 maternal zygotic homozygous mutant (mz ago2-/-) were stained with the Ab-panAGO-2A8 and secondary only as a control of the total staining background (scale bar = 100 µm). Bar plots indicate average of Ago2 fluorescence intensity normalized for the DAPI staining for each genotype (mean ± SEM, n = 3 replica with 10 embryos / group, * p = 0.0261, unpaired two-sided t-test). Right, RT-qPCR representing miRNAs level in wild-type vs mz ago2 -/- at 72 hpf (mean ± SEM, n = 3 embryos per group, value normalized for one wild-type embro). (b) Left panel, schematics representing the experimental procedure to test defective miRNA-mRNA interaction in the mz ago2-/- fin fold model. In vitro transcribed RNAs encoding for mCherry control and GFP coding sequence upstream of three perfect MREs for miR-24, a miRNA expressed in epidermis29. 25 picograms (pg) of each mRNA was co-injected into wild-type and ago2-/- zebrafish embryos at the one cell stage post fertilization. Right panel, confocal whole mount image of 48 hpf embryos expressing GFP and mCherry (head is to the left, scale bar = 60 µm). GFP and mCherry pixel intensities were quantified and the GFP/mCherry ratio was plotted for each genotype (mean ± SEM, n = 4 embryos / group, *p = 0.0137, unpaired two-sided t-test). (c) and (d) Top, Confocal lateral view of whole mount zebrafish embryos treated as indicated after zebrafish find fold assay (scale bar = 120 µm). White dotted line shows the edge of the wound. Bottom, box plot with minimum, maximum, median and quartiles representing the quantification of each whole mount stained embryos as indicated. Average intensity was calculated for 4 individual fish for PCNA and 4 to 6 fish for TUNEL assay. (Single dots represent single embryos, n.s.= non-significant). Source data can be found in Supplementary Table 8.
Supplementary Figure 7 Fin fold regeneration model in zebrafish wild-type and mutant embryos.
(a) Left, Confocal lateral view of whole mount zebrafish embryos treated as indicated (scale bar = 120 µm). White dotted line shows the edge of the wound. Right, quantification of each whole mount stained embryos as indicated. Pixel matrix of intensity was reconstructed for 50 μm within the wound edge to the fish body. Average intensity was calculated for 4 fish within each group and show as box plot with minimum, maximum, median and quartiles (Single dots represent single embryos, * in wt 2 h vs. ago 2 h = 0.0127; * in ago 2 h vs. ctgf 2 h = 0.0174; n.s.= non-significant; unpaired two-sided t-test) (b) Representative confocal lateral images of YAP immunostaining in the uncut fin fold. A DAPI binary mask was used to generate the Nuclear YAP (Fig. 7c) and Cytosol YAP images. (c) Alignment of human and zebrafish ctgfa 3’UTR sequence containing the conserved MREs. This 3′ UTR sequence was used to generate a wild-type and mutated miRNA sensor vector (as above). An in vitro transcribed mRNAs encoding for a GFP reporter was under the post-transcriptional regulation of ctgfa 3’UTR (wild-type), a mutated version lacking the MREa sequence and co-injected with an mCherry mRNA with no 3UTR regulation as negative control. 75 picograms (pg) of each mRNA was co-injected into Wild-Type and mz ago2-/- zebrafish embryos at the one cell stage post fertilization. GFP and mCherry pixel intensities were quantified and the GFP/mCherry ratio was plotted for each genotype (mean ± SEM, n = 5 embryo / group, *p = 0.0253, unpaired two-sided t-test). (e) Sequence alignment of the ctgfa 3UTR of zebrafish wilt-type and ctgfa MRE embryos injected with Cas9 and the gRNAs which PAM region was highlighted in black. Mutations are represented as nucleotide (nt) inserted (+) or deleted (Δ) and were cloned from individual embryos 24 h post injection. Red boxes represent the MREs targeted. Representative agarose gel for T7 Endonuclease I assay for Wild-Type and ctgfa MRE mutant zebrafish. Source data can be found in Supplementary Table 8. Unprocessed blots for 7b and d can be found in Supplementary Figure 8.
Supplementary Figure 8 Unprocessed scans of blots and gels.
This figure contains the unprocessed scans of blots and gels shown in the figures. Each blot or gel is labeled with the lanes, antibody used, and the corresponding figure.
Supplementary information
Supplementary Information
Supplementary Figures 1–8 and legends for Supplementary Tables 1–9.
Supplementary Table 1
Ago2-peak localization.
Supplementary Table 2
Ago2-miRNAs are clustered in 155 families.
Supplementary Table 3
miRNA-CAM–MRE interaction network.
Supplementary Table 4
CAM–MRE sequences.
Supplementary Table 5
CAM–MRE sequences.
Supplementary Table 6
List of oligos.
Supplementary Table 7
miRNA differentially expressed between EC and HDF.
Supplementary Table 8
Compiled figure source data.
Supplementary Table 9
Polyacrylamide gels of varying stiffness.
Rights and permissions
About this article
Cite this article
Moro, A., Driscoll, T.P., Boraas, L.C. et al. MicroRNA-dependent regulation of biomechanical genes establishes tissue stiffness homeostasis. Nat Cell Biol 21, 348–358 (2019). https://doi.org/10.1038/s41556-019-0272-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-019-0272-y
This article is cited by
-
The impact of substrate stiffness on morphological, transcriptional and functional aspects in RPE
Scientific Reports (2024)
-
LIS1 RNA-binding orchestrates the mechanosensitive properties of embryonic stem cells in AGO2-dependent and independent ways
Nature Communications (2023)
-
AGO2 silences mobile transposons in the nucleus of quiescent cells
Nature Structural & Molecular Biology (2023)
-
Mechanical regulation of chromatin and transcription
Nature Reviews Genetics (2022)
-
Hippo-Yap/Taz signalling in zebrafish regeneration
npj Regenerative Medicine (2022)