Abstract
Tissue morphogenesis is driven by mechanical forces that elicit changes in cell size, shape and motion. The extent by which forces deform tissues critically depends on the rheological properties of the recipient tissue. Yet, whether and how dynamic changes in tissue rheology affect tissue morphogenesis and how they are regulated within the developing organism remain unclear. Here, we show that blastoderm spreading at the onset of zebrafish morphogenesis relies on a rapid, pronounced and spatially patterned tissue fluidization. Blastoderm fluidization is temporally controlled by mitotic cell rounding-dependent cell–cell contact disassembly during the last rounds of cell cleavages. Moreover, fluidization is spatially restricted to the central blastoderm by local activation of non-canonical Wnt signalling within the blastoderm margin, increasing cell cohesion and thereby counteracting the effect of mitotic rounding on contact disassembly. Overall, our results identify a fluidity transition mediated by loss of cell cohesion as a critical regulator of embryo morphogenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Source data for Figs. 1c–f, 2c,d, 3b–e, 4b–d, 5b,c,e, 6b–f and 7b,d,f–h and Supplementary Figs. 1b–g, 2a,b,d, 3b,c, 4, 5a–d,f, 6b–e,h,i and 7a,d–f,h,i have been provided as Supplementary Table 3. All other data supporting the findings of this study are available from the corresponding author upon request.
References
Heisenberg, C. P. & Bellaïche, Y. Forces in tissue morphogenesis and patterning. Cell 153, 948–962 (2013).
Petridou, N. I., Spiró, Z. & Heisenberg, C. P. Multiscale force sensing in development. Nat. Cell Biol. 19, 581–588 (2017).
Gonzalez-Rodriguez, D., Guevorkian, K., Douezan, S. & Brochard-Wyart, F. Soft matter models of developing tissues and tumors. Science 338, 910–917 (2012).
Forgacs, G., Foty, R. A., Shafrir, Y. & Steinberg, M. S. Viscoelastic properties of living embryonic tissues: a quantitative study. Biophys. J. 74, 2227–2234 (1998).
Lawton, A. K. et al. Regulated tissue fluidity steers zebrafish body elongation. Development 140, 573–582 (2013).
Mongera, A. et al. A fluid-to-solid jamming transition underlies vertebrate body axis elongation. Nature 561, 401–405 (2018).
Serwane, F. et al. In vivo quantification of spatially varying mechanical properties in developing tissues. Nat. Methods 14, 181–186 (2017).
Moore, S. W., Keller, R. E. & Koehl, M. A. The dorsal involuting marginal zone stiffens anisotropically during its convergent extension in the gastrula of Xenopus laevis. Development 121, 3131–3140 (1995).
Shook, D. R., Kasprowicz, E. M., Davidson, L. A. & Keller, R. Large, long range tensile forces drive convergence during Xenopus blastopore closure and body axis elongation. eLife 7, e26944 (2018).
David, R. et al. Tissue cohesion and the mechanics of cell rearrangement. Development 141, 3672–3682 (2014).
Ranft, J. et al. Fluidization of tissues by cell division and apoptosis. Proc. Natl Acad. Sci. USA 107, 20863–20868 (2010).
Stirbat, T. V. et al. Fine tuning of tissues' viscosity and surface tension through contractility suggests a new role for α-catenin. PLoS ONE 8, e52554 (2013).
Bi, D., Lopez, J. H., Schwarz, J. M. & Manning, M. L. A density-independent rigidity transition in biological tissues. Nat. Phys. 11, 1074–1079 (2015).
Bi, D., Yang, X., Marchetti, M. C. & Manning, M. L. Motility-driven glass and jamming transitions in biological tissues. Phys. Rev. X 6, 1–13 (2016).
Angelini, T. E. et al. Glass-like dynamics of collective cell migration. Proc. Natl Acad. Sci. USA 108, 4714–4719 (2011).
Sadati, M., Taheri Qazvini, N., Krishnan, R., Park, C. Y. & Fredberg, J. J. Collective migration and cell jamming. Differentiation 86, 121–125 (2013).
Bruce, A. E. E. Zebrafish epiboly: spreading thin over the yolk. Dev. Dyn. 245, 244–258 (2016).
Lepage, S. E. & Bruce, A. E. E. Zebrafish epiboly: mechanics and mechanisms. Int. J. Dev. Biol. 54, 1213–1228 (2010).
Morita, H. et al. The physical basis of coordinated tissue spreading in zebrafish gastrulation. Dev. Cell 40, 354–366.e4 (2017).
Davidson, L. A. Embryo mechanics: balancing force production with elastic resistance during morphogenesis. Curr. Top. Dev. Biol. 95, 215–241 (2011).
Guevorkian, K., Colbert, M. J., Durth, M., Dufour, S. & Brochard-Wyart, F. Aspiration of biological viscoelastic drops. Phys. Rev. Lett. 104, 1–4 (2010).
Guevorkian, K. & Maître, J. L. Micropipette aspiration: a unique tool for exploring cell and tissue mechanics in vivo. Methods Cell Biol. 139, 187–201 (2017).
Fukazawa, C. et al. poky/chuk/ikk1 is required for differentiation of the zebrafish embryonic epidermis. Dev. Biol. 346, 272–283 (2010).
Song, S. et al. Pou5f1-dependent EGF expression controls E-cadherin endocytosis, cell adhesion, and zebrafish epiboly movements. Dev. Cell 24, 486–507 (2013).
Shimizu, T. et al. E-cadherin is required for gastrulation cell movements in zebrafish. Mech. Dev. 122, 747–763 (2005).
Maitre, J.-L. et al. Adhesion functions in cell sorting by mechanically coupling the cortices of adhering cells. Science 338, 253–256 (2012).
Mendieta-Serrano, M. A., Schnabel, D., Lomelí, H. & Salas-Vidal, E. Cell proliferation patterns in early zebrafish development. Anat. Rec. 296, 759–773 (2013).
Kimmel, C. B., Ballard, W. W., Kimmel, S. R., Ullmann, B. & Schilling, T. F. Stages of embryonic development of the zebrafish. Dev. Dyn. 203, 253–310 (1995).
Sorce, B. et al. Mitotic cells contract actomyosin cortex and generate pressure to round against or escape epithelial confinement. Nat. Commun. 6, 8872 (2015).
Stewart, M. P. et al. Hydrostatic pressure and the actomyosin cortex drive mitotic cell rounding. Nature 469, 226–231 (2011).
Théry, M. & Bornens, M. Get round and stiff for mitosis. HFSP J. 2, 65–71 (2008).
Rosenblatt, J. Mitosis: moesin and the importance of being round. Curr. Biol. 18, 1–6 (2015).
Carreno, S. et al. Moesin and its activating kinase Slik are required for cortical stability and microtubule organization in mitotic cells. J. Cell Biol. 180, 739–746 (2008).
Tachibana, K., Haghparast, S. M. A. & Miyake, J. Inhibition of cell adhesion by phosphorylated ezrin/radixin/moesin. Cell Adhes. Migr. 9, 502–512 (2015).
Makita, R., Mizuno, T., Koshida, S., Kuroiwa, A. & Takeda, H. Zebrafish wnt11: pattern and regulation of the expression by the yolk cell and no tail activity. Mech. Dev. 71, 165–176 (1998).
Witzel, S., Zimyanin, V., Carreira-Barbosa, F., Tada, M. & Heisenberg, C. P. Wnt11 controls cell contact persistence by local accumulation of Frizzled 7 at the plasma membrane. J. Cell Biol. 175, 791–802 (2006).
Heisenberg, C.-P. et al. Silberblick/Wnt11 mediates convergent extension movements during zebrafish gastrulation. Nature 405, 76–81 (2000).
Ulrich, F. et al. Slb/Wnt11 controls hypoblast cell migration and morphogenesis at the onset of zebrafish gastrulation. Development 130, 5375–5384 (2003).
Dzamba, B. J., Jakab, K. R., Marsden, M., Schwartz, M. A. & DeSimone, D. W. Cadherin adhesion, tissue tension, and noncanonical Wnt signaling regulate fibronectin matrix organization. Dev. Cell 16, 421–432 (2009).
Brodland, G. W. et al. CellFIT: a cellular force-inference toolkit using curvilinear cell boundaries. PLoS ONE 9, e99116 (2014).
Firmino, J., Rocancourt, D., Saadaoui, M., Moreau, C. & Gros, J. Cell division drives epithelial cell rearrangements during gastrulation in chick. Dev. Cell 36, 249–261 (2016).
Meilhac, S. M. et al. Active cell movements coupled to positional induction are involved in lineage segregation in the mouse blastocyst. Dev. Biol. 331, 210–221 (2009).
Xenopoulos, P., Kang, M. & Hadjantonakis, A.-K. Cell lineage allocation within the inner cell mass of the mouse blastocyst. Results Probl. Cell Differ. 55, 185–202 (2012).
Ulrich, F. et al. Wnt11 functions in gastrulation by controlling cell cohesion through Rab5c and E-cadherin. Dev. Cell 9, 555–564 (2005).
Schoetz, E.-M., Lanio, M., Talbot, J. A. & Manning, M. L. Glassy dynamics in three-dimensional embryonic tissues. J. R. Soc. Interface 10, 20130726 (2013).
Detrich, W. H. I., Westerfield, M. & Zon, L. I. (eds) The Zebrafish: Biology (Academic Press, Cambridge, 1999).
Behrndt, M. et al. Forces driving epithelial spreading in zebrafish gastrulation. Science 338, 257–260 (2012).
Quesada-Hernández, E. et al. Stereotypical cell division orientation controls neural rod midline formation in zebrafish. Curr. Biol. 20, 1966–1972 (2010).
Wagner, D. S., Dosch, R., Mintzer, K. A., Wiemelt, A. P. & Mullins, M. C. Maternal control of development at the midblastula transition and beyond: mutants from the zebrafish II. Dev. Cell 6, 781–790 (2004).
Kimmel, C. B. & Law, R. D. Cell lineage of zebrafish blastomeres. I. Cleavage pattern and cytoplasmic bridges between cells. Dev. Biol. 108, 78–85 (1985).
Kimmel, R. A. & Meyer, D. Molecular regulation of pancreas development in zebrafish. Methods Cell Biol. 100, 261–280 (2010).
Iioka, H., Ueno, N. & Kinoshita, N. Essential role of MARCKS in cortical actin dynamics during gastrulation movements. J. Cell Biol. 164, 169–174 (2004).
Arboleda-Estudillo, Y. et al. Movement directionality in collective migration of germ layer progenitors. Curr. Biol. 20, 161–169 (2010).
Keller, P. J., Schmidt, A. D., Wittbrodt, J. & Stelzer, E. H. K. Reconstruction of zebrafish early embryonic development by scanned light sheet microscopy. Science 322, 1065–1069 (2008).
Ruprecht, V. et al. Cortical contractility triggers a stochastic switch to fast amoeboid cell motility. Cell 160, 673–685 (2015).
Jopling, C. & den Hertog, J. Fyn/Yes and non-canonical Wnt signalling converge on RhoA in vertebrate gastrulation cell movements. EMBO Rep. 6, 426–431 (2005).
Krens, S. F. G. et al. Interstitial fluid osmolarity modulates the action of differential tissue surface tension in progenitor cell segregation during gastrulation. Development 144, 1798–1806 (2017).
Brodland, G. W. et al. Video force microscopy reveals the mechanics of ventral furrow invagination in Drosophila. Proc. Natl Acad. Sci. USA 107, 22111–22116 (2010).
Thisse, C., Thisse, B., Halpern, M. E. & Postlethwait, J. H. Goosecoid expression in neurectoderm and mesendoderm is disrupted in zebrafish cyclops gastrulas. Dev. Biol. 164, 420–429 (1994).
Acknowledgements
We thank O. Campas and members of the Heisenberg laboratory for technical advice and discussions, and the Bioimaging and zebrafish facilities of the IST Austria for continuous support. This work was supported by a postdoctoral fellowship from EMBO Long-term fellowships to N.I.P. (ALTF-534 2016) and an ERC Advanced Grant (MECSPEC) to C.-P.H. S.G. and G.S. were supported by The Francis Crick Institute, which receives its core funding from Cancer Research UK (FC001317), the UK Medical Research Council (FC001317) and the Wellcome Trust (FC001317).
Author information
Authors and Affiliations
Contributions
N.I.P. and C.-P.H. designed the research. N.I.P. performed the experiments and analysed the experimental data. G.S. and S.G. developed the theoretical doming models. S.G. performed the doming simulations. E.H. analysed the cell cohesion and tissue rigidity data. N.I.P. and C.-P.H. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Deep cell protrusive activity and tissue viscosity measurements, micropipette aspiration controls, and simulations of tissue thinning with different tissue viscosities.
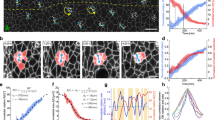
a, High-resolution images of deep cells within the blastoderm (dashed grey box in embryo schematic) in a transgenic embryo expressing GFP-Utrophin to visualize F-actin at consecutive time points during doming. Double-arrowed white line indicates radial axis of the blastoderm. Active radial intercalation was evaluated by the presence of polarized actin accumulation along the radial axis of deep cells (white arrowheads). b, Plot of the relative distribution of F-actin intensity (left axis) along the radial (I rad) versus lateral (I lat) axis of deep cells (outlined in the cell schematic in (a); 3 min intervals, n = 10 cells for each time point) and percentage of dividing cells (right axis; 30 min interval) during doming. The grey shaded box represents the time period of doming initiation. c, Representative deformation curves of deep cell tissues in the aspiration experiments for central and marginal blastoderm and for central blastoderm explants. The shaded lines show the time point of pressure release. d, Examples of viscosity measurements from the plot of the length of the aspirated tissue over time (black line indicates the slopes of the linear part of the curve, where tissue flow is constant and the rate of aspiration and retraction are obtained). ∆P, aspiration pressure (200 Pa); Pc, critical pressure. e, Bar plots of blastoderm viscosity calculated from aspiration experiments of the indicated stages and tissues using different pipette size and associated applied pressure (25 μm – 250 Pa, 35 μm – 200 Pa, 45 μm – 150 Pa), (25 μm: n = 24 embryos for centre, n = 18 embryos for margin; 35 μm: n = 25 embryos for centre, n = 33 embryos for margin; 45 μm: n = 34 embryos for centre, n = 26 embryos for margin). f, Plot of viscosity as a function of micropipette size from the data in (e); linear regression analysis on the right reveals no significant linear relationship between viscosity and micropipette size (R2, Pearson coefficient of determination). g, Bar plot of the TST of the central and marginal aspirated tissues for the indicated stages, measured from the micropipette experiments. The absence of recoil in the central tissue is due to the very low TST of this tissue when aspirated by the pipette (centre high stage, n = 10 embryos; centre onset stage, n = 30 embryos; margin onset stage, n = 22 embryos). h, Exemplary bright-field/fluorescence images of EVL and deep cells during and after deep cell aspiration. Cells are labelled by GFP-Utrophin to mark Actin (green). White arrowheads indicate wound response in the EVL tissue after the micropipette is removed from the embryo. i, Schematic representation of the simplified model describing the flow of the deep cell tissue with different viscosities when subjected to an external force by the EVL driving its elongation (black filled rectangle). The system is represented by 3 rectangles with different viscosities in the central versus marginal rectangles. The EVL is assumed to exert a force at the left and right boundaries of the deep cell tissue, maintaining a constant velocity at the margin. j, k, Time evolution of the rectangle widths (j) and heights (k), normalized with respect to the initial widths and heights. Green and red thick lines correspond to marginal and central regions when central viscosity is smaller than marginal viscosity; green and red dashed line corresponds to marginal and central regions when central viscosity is equal to marginal viscosity. Viscosity values are taken from the Supplementary Table 1 and the velocity of the margin is assumed to be 1.1 µm/min. Images are representative of 3 (a), 83 (c-d, centre), 81 (c, margin) or 10 embryos (h), 37 explants (c, explants), and 3 (e), 4 (b) or 9 (e) biological replicates (different embryos in b, h; independent embryo batches in e). Data are mean ± s.e.m; Two-tailed Mann-Whitney test (e), Kruskal-Wallis test (g). Scale bar: (a) 20 µm; (h) 50 µm.
Supplementary Figure 2 Promoting deep cell cohesion suppresses blastoderm tissue fluidization at the onset of doming.
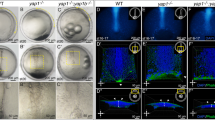
a, Exemplary confocal images of a transplanted E-cadherin coated bead marked by Alexa Fluor 546 rabbit IgG in magenta and attached to surrounding deep cells within the central blastoderm of a WT embryo at dome stage. White arrowhead indicates F-actin accumulation marked by Lifeact-GFP (red) in deep cells surrounding the bead. Interstitial fluid is labelled by dextran (blue). Bar plot on the right shows interstitial fluid intensity/accumulation between deep cells in areas near (magenta dashed box) and further away (grey dashed box) from the bead (distant areas n = 19; close areas n = 16). b, Bright-field/fluorescence images of exemplary deep cell aspirations in the blastoderm centre of WT control and transplanted embryos containing E-Cadherin coated beads (magenta) in the centre at the onset of doming. Black arrowheads indicate how far the cells have flown into the aspiration pipette under the same pressure. White arrowhead points at transplanted bead within the aspiration pipette. Bar plot on the right shows blastoderm viscosity calculated from the aspiration experiments for the indicated conditions (control, n = 4 embryos; E-cadherin bead transplants, n = 4 embryos). c, Schematic diagram and exemplary single plane bright-field/fluorescence images of WT embryos containing E-cadherin coated beads on one side of the central blastoderm at the onset and end of doming. White arrowhead indicates defective doming in the transplanted area. d, Schematic diagram outlining the quantification of BYI deformation at the side of the transplanted bead (magenta arrow) versus the non-transplanted side (grey arrows). Plot on the right shows BYI displacement in the transplanted versus non-transplanted side of the central blastoderm as a function of time during doming (n = 2 transplants). The onset of doming is indicated with grey dashed line. Images are representative of 4 embryos (c), and 2 (d) or 3 (a, b) biological replicates (different embryos in a; independent embryo batches in b, d). Data are mean ± s.e.m; Mann-Whitney test (a, b). Scale bars: (a) 50 µm; (b, c) 100 µm.
Supplementary Figure 3 Interstitial fluid accumulations are dispensable for deep cell tissue fluidization and contact dynamics.
a, Exemplary confocal images of deep cells in the blastoderm centre of WT and pky embryos at the onset of doming. Deep cells are marked by membrane-RFP (red), interstitial fluid by dextran (cyan) and nuclei by H2B-GFP (blue). b, Bar plot of cell-cell contact time between central cells normalized to the total duration of the indicated stages in WT and pky mutant embryos (WT centre, n = 90 contacts; pky centre, n = 80 contacts). c, Bright-field images of exemplary deep cell aspirations in the central and marginal blastoderm at the onset of doming in WT and pky embryos either in isotonic (iso) or hypotonic (hypo) medium. Black arrowheads indicate how far the cells have flown into the aspiration pipette under the same pressure. Bar plot on the right shows blastoderm viscosity calculated from the aspiration experiments for the conditions described above (WT centre, n = 10 embryos; WT margin, n = 10 embryos; pky centre iso, n = 14 embryos; pky margin iso, n = 13 embryos; pky centre hypo, n = 8 embryos; pky margin hypo, n = 6 embryos). Images are representative of 3 embryos each (a) and 3 biological replicates (b, c) (different embryos in b; independent embryo batches in c). Data are mean ± s.e.m; Two-tailed Mann-Whitney test (b), Kruskal-Wallis test (c). Scale bars: (a) 20 µm; (c) 100 µm; (e) 30 µm.
Supplementary Figure 4 Blastoderm central cells intrinsically loose cell-cell contacts.
Exemplary fluorescence images of explanted central and marginal deep cells from transgenic embryos expressing Lifeact-GFP to mark Actin and pseudo-coloured in red (central) and green (marginal), respectively, at high stage and the onset of doming. Plot on the right shows cell-cell contact length normalized to cell diameter before, during and after doming (centre, n = 10 cells each time point; margin, n = 10 cells each time point). Note, that explanted deep cells often bleb at the contact-free surfaces (white arrowheads). The grey dashed line indicates the onset of doming. Images are representative of 7 cell cultures each and 2 independent biological replicates (independent embryo batches). Data are mean ± s.e.m. Scale bar: 30 µm.
Supplementary Figure 5 Mitotic deep cell rounding-induced contact disassembly temporally controls blastoderm fluidization.
a, Bar plot of the aspect ratio of interphase and/or metaphase deep cells from high stage to the onset of doming for the conditions outlined in Fig. 5a (WT centre S-phase, n = 91 cells; WT centre M-phase, n = 90 cells; WT margin S-phase, n = 90 cells; WT margin M-phase, n = 90 cells; HUA centre, n = 91 cells; DN Ezrin centre S-phase, n = 87 cells; DN Ezrin centre M-phase, n = 89 cells). b, Bar plot of cell-cell contact length reduction for the conditions in Fig. 5a (Cycle 11: WT centre n = 24 contacts; WT margin n = 14 contacts; DN Ezrin centre n = 20 contacts; HUA-treated and transplanted centre n = 10 contacts; Cycle 12: WT centre n = 11 contacts; WT margin n = 14 contacts; DN Ezrin centre n = 20 contacts; HUA-treated and transplanted centre n = 5 contacts). Contact length reduction, D, is expressed as D = Lbm - Lm; where Lbm, contact length 9 min before metaphase and Lm, contact length at metaphase (data were normalized to Lbm for each cycle). c, Bar plot of contact length change for the conditions in (b). Contact length change (R) is expressed as R = Lam - Lbm; where Lam, contact length 9 min after metaphase and Lbm, contact length at 9 min before metaphase (data were normalized to Lbm for each cycle). d, Bright-field images of exemplary deep cell aspirations in the blastoderm centre of WT control and HUA-treated embryos at the onset of doming. Bar plot below shows blastoderm viscosity calculated from the aspiration experiments for the indicated conditions (WT, n = 5 embryos; HUA, n = 14 embryos). e, Exemplary confocal images of HUA-treated deep cells transplanted into the blastoderm centre of a WT embryo at the onset of doming and at 50% epiboly stage. Host plasma membrane is labelled by membrane-RFP (red), host nuclei by H2B-GFP (green) and interstitial fluid is marked by dextran (cyan). Donor plasma membrane is labelled by membrane-GFP (green) and donor nuclei by H2A-mCherry (red). White dashed outlines demarcate the transplanted area. f, Bright-field/fluorescence images of exemplary deep cell aspirations in the blastoderm centre of either control homotypic transplanted embryos or embryos transplanted with DN Ezrin expressing central deep cells at the onset of doming. Bar plot below shows blastoderm viscosity calculated from the aspiration experiments for the indicated conditions (WT, n = 9 transplants; DN Ezrin, n = 5 transplants). Black arrowheads in (d, f) indicate how far the cells have flown into the aspiration pipette under the same pressure. Images are representative of 3 transplants (e), and 2 (f) or 3 (a-d) biological replicates (different embryos in a-c; independent embryo batches in d, f). Data are mean ± s.e.m; Kruskal-Wallis test (a-c); Two-tailed Mann-Whitney test (d, f). Scale bars: (d, f) 100 µm; (e) 30 µm.
Supplementary Figure 6 Wnt11/Fz7-signalling within the blastoderm margin spatially restricts blastoderm fluidization.
a, Exemplary confocal images of deep cell-cell contacts (white arrowheads) during mitotic rounding for WT, slb and MZ fz7a/b marginal cells labelled by membrane-RFP pseudo-coloured in green, yellow and magenta, respectively. Nuclei are labelled by H2B-GFP (blue) and interstitial fluid by dextran (cyan). b, Deep cell-cell contact length normalized to cell diameter plotted for the following conditions: 9 min before metaphase (tbm), at metaphase (tm) and 9 min after metaphase (tam) for cycles 11th to 13th for WT, slb and MZ fz7a/b marginal deep cells (WT n = 16 contacts; slb n = 12 contacts; MZ fz7a/b n = 15 contacts). Grey dashed lines indicate the metaphase time points for each cell cycle. Cyan shaded box shows the period of contact disassembly and orange shaded box the period of contact re-assembly. c, Bar plot of cell-cell contact length reduction for WT, slb and MZ fz7a/b marginal deep cells (cycle 11: WT n = 14 contacts; slb n = 10 contacts; MZ fz7a/b n = 14 contacts; cycle 12: WT centre n = 14 contacts; slb n = 7 contacts; MZ fz7a/b n = 7 contacts). Contact length reduction (D) is expressed as D = Lbm - Lm; where Lbm, contact length 9 min before metaphase and Lm, contact length at metaphase (data were normalized to Lbm for each cycle). d, Bar plot of cell-cell contact length change for the conditions in (c). Contact length change (R) is expressed as: R = Lam - Lbm; where Lam, contact length 9 min after metaphase and Lbm, contact length at 9 min before metaphase (data were normalized to Lbm for each cycle). e, Bright-field images of exemplary deep cell aspirations in the blastoderm margin of WT and MZ fz7a/b mutant embryos at the onset of doming. Black arrowheads indicate how far the cells have flown into the aspiration pipette. Bar plot on the right shows marginal blastoderm viscosity calculated from the aspiration experiments for WT and MZ fz7a/b mutant embryos at the onset of doming (WT, n = 11 embryos; MZ fz7a/b, n = 13 embryos). f, Exemplary single plane fluorescence/bright-field images of WT embryos containing transplanted WT central, WT marginal and MZ fz7a/b marginal cells in their blastoderm centre. Transplanted cells are fluorescently marked by H2B-GFP pseudo-coloured in red, green and magenta, respectively. g, Exemplary confocal images of donor and host deep cells within the blastoderm centre for the conditions shown in (f). Transplanted donor cells are marked by membrane-GFP (WT centre, red; WT margin, green; MZ fz7a/b margin, magenta), host cells by membrane-RFP (grey), nuclei by H2B-GFP (magenta) and interstitial fluid by dextran (blue). h, BYI displacement as a function of time during doming for the different conditions shown in (f) (WT centre to centre, n = 7 transplants; WT margin to centre, n = 4 transplants; MZ fz7a/b margin to centre, n = 5 transplants). i, Bar plot of the normalized cell-cell contact time in the conditions shown in (f) (WT central cells n = 100; WT marginal cells n = 360; MZ fz7a/b marginal cells n = 200). The onset of doming is indicated by grey dashed line. Images are representative of 3 embryos each (a), at least 5 transplants each (f, g), and 3 (b-e, h) or 5 (i) biological replicates (different embryos in b-d, i; independent embryo batches in e, h). Data are mean ± s.e.m; Kruskal-Wallis test (c, d, i); Two-tailed Mann-Whitney test (e). Scale bars: (a) 20 µm; (e, f) 100 µm; (g) 20 µm.
Supplementary Figure 7 Localized Wnt/Fz7-signalling defines the spatial pattern of deep cell cohesion within the blastoderm and validation of the CellFIT3D method for estimating in vivo TST.
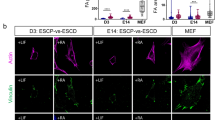
a, Exemplary confocal image of WT marginal cells marked by membrane-GFP (green) and transplanted in the blastoderm centre of a WT host embryo marked by membrane-RFP (grey) at the onset of doming. Interstitial fluid is labelled by dextran (blue). Bar plot on the right of cell-cell contact time at dome stage between WT central cells, WT host central cells and WT donor marginal cells (WT centre, n = 20 contacts; WT host centre, n = 17 contacts; WT margin donor, n = 25 contacts). b, Schematic illustration of WT homotypic (WT marginal cells to WT margin), WT heterotypic (WT central cells to WT margin), Fz7a overexpression (oe) WT heterotypic (Fz7a oe central cells to WT margin) and Fz7a oe slb heterotypic (Fz7a oe central cells to slb margin) deep cell transplantations. c, Exemplary confocal images of transplanted donor and host deep cells for the conditions shown in (b) at the onset of doming. Transplanted donor cells are marked by membrane-GFP pseudo-coloured in green (WT homotypic), red (WT heterotypic) and by Fz7a-Neon pseudo-coloured in cyan (Fz7a oe heterotypic), host cells by membrane-RFP (grey for WT, yellow for slb) and interstitial fluid by dextran (blue). d, Bar plot of normalized cell-cell contact time between transplanted donor cells at dome stage (WT homotypic n = 71; WT heterotypic n = 107; Fz7a oe WT heterotypic n = 44; Fz7a oe slb heterotypic n = 32). e, Bright-field/fluorescent images of exemplary deep cell aspirations in the transplantations described in (b). Black arrowheads indicate how far the cells have flown into the micropipette upon application of the same pressure. Bar plot on the right shows tissue viscosity of the transplanted area at the onset of doming calculated from the aspiration experiments (WT homotypic, n = 10 transplants; WT heterotypic, n = 17 transplants; Fz7a oe WT heterotypic, n = 15 transplants; Fz7a oe slb heterotypic, n = 8 transplants). f, Exemplary single plane fluorescence/bright-field images of the transplanted embryos for WT homotypic (WT marginal cells to WT margin), WT heterotypic (WT central cells to WT margin) and Fz7a oe WT heterotypic (Fz7a oe central cells to WT margin) at the onset and end of doming. Transplanted cells are fluorescently marked by H2B-GFP pseudo-coloured in green (WT marginal cells), red (WT central cells) and cyan (Fz7a oe central cells), respectively. Dashed lines indicate the BYI angle at the margin-to-centre transition zone. Schematic diagram and bar plot on the right shows the BYI angle formed at the transition zone between the marginal and central regions at dome stage (indicated by the dashed lines) for the different transplantations described above (WT homotypic n = 28 transplants; WT heterotypic n = 32 transplants; Fz7a oe WT heterotypic n = 12 transplants). g, Representative CellFIT-3D analysis of TST from fluorescent confocal z-stacks of central and marginal cells in an in vivo (left panel) versus ex vivo (right panel) context. For the in vivo context, marginal donor cells (marked by membrane-GFP in green) were transplanted in the central blastoderm of WT host embryos (host cells were marked by membrane-RFP in red and the surrounding interstitial fluid by dextran in blue). For the ex vivo context, central (red) and marginal (green) cells were isolated and cultured as described in the Methods section. The white dashed rectangles show close-ups of the angles measured (yellow marks) at the triplet edges using CellFIT-3D software. h, Plot of the relative TST calculated by the CellFIT-3D software based on the measured angles (see Methods) with marginal cells displaying increased TST compared to central cells ex vivo, but not in vivo (centre in vivo, n = 41 interfaces; margin in vivo, n = 120 interfaces; centre ex vivo, n = 50 interfaces; margin ex vivo, n = 48 interfaces). i, Representative micropipette aspiration (MPA) analysis of TST from aspirated central and marginal cells. Plot on the right shows the calculated TST from the aspiration experiments with marginal cells displaying higher TST than central cells at their interface to the buffer in the pipette (centre, n = 30 embryos; margin, n = 22 embryos). The observation that 3D-VFM and MPA reveal similar tension distributions at comparable interfaces (cell-to-buffer for MPA and cell-to-culture medium for 3D-VFM) suggests that 3D-VFM is a valid method for analysing interfacial tensions both in vivo and ex vivo. Images are representative of 3 embryos each (c) or at least 5 embryos/cultures (g) and 2 (a), 4 (e, f), 5 (h) and 9 (i) biological replicates (different embryos in a, c; independent embryo batches in e, f, h, i). Data are mean ± s.e.m; Kruskal-Wallis test (a, d-g); two-tailed Mann-Whitney test (h, i). Scale bars: (a, c) 20 µm; (e, f) 100 µm; (g, close up) 10 μm.
Supplementary information
Supplementary Information
Supplementary Figures 1–7, Supplementary Table and Supplementary Video legends.
Supplementary Table 1
Doming simulation parameters.
Supplementary Table 2
Relative TST measurements in transplants using CellFIT-3D.
Supplementary Table 3
Statistics Source Data.
Supplementary Video 1
Zebrafish doming.
Supplementary Video 2
Deep-cell tissue micropipette aspiration.
Supplementary Video 3
Simulated deformation of deep cell tissue with varying viscosity.
Supplementary Video 4
Simulated doming movements in embryos with varying viscosity.
Supplementary Video 5
Doming of transplanted embryos containing central, marginal or earlier-staged central deep cells in the central blastoderm.
Supplementary Video 6
Deep cell cohesion during doming.
Supplementary Video 7
Mitotic rounding-induced deep cell–cell contact disassembly.
Supplementary Video 8
Deep cell–cell contact dynamics in HUA-treated embryos.
Supplementary Video 9
Doming of transplanted embryos containing HUA pre-treated central cells.
Supplementary Video 10
Deep cell tissue micropipette aspiration in slb mutants.
Supplementary Video 11
Doming of transplanted embryos containing WT, slb or slb+ CARhoA marginal deep cells in the central blastoderm.
Supplementary Note
Supplementary Theory.
Rights and permissions
About this article
Cite this article
Petridou, N.I., Grigolon, S., Salbreux, G. et al. Fluidization-mediated tissue spreading by mitotic cell rounding and non-canonical Wnt signalling. Nat Cell Biol 21, 169–178 (2019). https://doi.org/10.1038/s41556-018-0247-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0247-4
This article is cited by
-
Mechanical state transitions in the regulation of tissue form and function
Nature Reviews Molecular Cell Biology (2024)
-
Adherens junctions as molecular regulators of emergent tissue mechanics
Nature Reviews Molecular Cell Biology (2024)
-
Cell cycle dynamics control fluidity of the developing mouse neuroepithelium
Nature Physics (2023)
-
Live imaging and conditional disruption of native PCP activity using endogenously tagged zebrafish sfGFP-Vangl2
Nature Communications (2022)
-
Single-molecule tracking of Nodal and Lefty in live zebrafish embryos supports hindered diffusion model
Nature Communications (2022)