Abstract
Vesicular acidification and trafficking are associated with various cellular processes. However, their pathologic relevance to cancer remains elusive. We identified transmembrane protein 9 (TMEM9) as a vesicular acidification regulator. TMEM9 is highly upregulated in colorectal cancer. Proteomic and biochemical analyses show that TMEM9 binds to and facilitates assembly of vacuolar-ATPase (v-ATPase), a vacuolar proton pump, resulting in enhanced vesicular acidification and trafficking. TMEM9-v-ATPase hyperactivates Wnt/β-catenin signalling via lysosomal degradation of adenomatous polyposis coli (APC). Moreover, TMEM9 transactivated by β-catenin functions as a positive feedback regulator of Wnt signalling in colorectal cancer. Genetic ablation of TMEM9 inhibits colorectal cancer cell proliferation in vitro, ex vivo and in vivo mouse models. Moreover, administration of v-ATPase inhibitors suppresses intestinal tumorigenesis of APC mouse models and human patient-derived xenografts. Our results reveal the unexpected roles of TMEM9-controlled vesicular acidification in hyperactivating Wnt/β-catenin signalling through APC degradation, and propose the blockade of TMEM9-v-ATPase as a viable option for colorectal cancer treatment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Microarray data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under accession code GDS2947. The TMEM9 expression data in CRC cells were derived from the cBioportal using the TCGA Research Network (http://cancergenome.nih.gov/) and Genetech data sets. The data set derived from this resource that supports the findings of this study is available in Oncomine (https://www.oncomine.org/resource). TMEM9 expression data were also derived from cBioportal (http://www.cbioportal.org/) and the COSMIC database (Catalogue of Somatic Mutations in Cancer) (https://cancer.sanger.ac.uk/cosmic). Source data for Figs. 1–8 and Supplementary Figs. 1–4 are provided as Supplementary Table 4. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
References
Clevers, H. & Nusse, R. Wnt/β-catenin signaling and disease. Cell 149, 1192–1205 (2012).
Polakis, P. The many ways of Wnt in cancer. Curr. Opin. Genet. Dev. 17, 45–51 (2007).
Bilic, J. et al. Wnt induces LRP6 signalosomes and promotes Dishevelled-dependent LRP6 phosphorylation. Science 316, 1619–1622 (2007).
Zeng, X. et al. Initiation of Wnt signaling: control of Wnt coreceptor Lrp6 phosphorylation/activation via Frizzled, Dishevelled and Axin functions. Development 135, 367–375 (2008).
Cadigan, K. M. & Waterman, M. L. TCF/LEFs and Wnt signaling in the nucleus. Cold Spring Harb. Perspect. Biol. 4, a007906 (2012).
Taelman, V. F. et al. Wnt signaling requires sequestration of glycogen synthase kinase 3 inside multivesicular endosomes. Cell 143, 1136–1148 (2010).
Polakis, P. Wnt signaling in cancer. Cold Spring Harb. Perspect. Biol. 4, a008052 (2012).
Phelps, R. A. et al. A two-step model for colon adenoma initiation and progression caused by APC loss. Cell 137, 623–634 (2009).
Vermeulen, L. et al. Wnt activity defines colon cancer stem cells and is regulated by the microenvironment. Nat. Cell Biol. 12, 468–476 (2010).
Voloshanenko, O. et al. Wnt secretion is required to maintain high levels of Wnt activity in colon cancer cells. Nat. Commun. 4, 2610 (2013).
Jung, Y. S., Jun, S., Lee, S. H., Sharma, A. & Park, J. I. Wnt2 complements Wnt/β-catenin signaling in colorectal cancer. Oncotarget 6, 37257–37268 (2015).
Niehrs, C. & Boutros, M. Trafficking, acidification, and growth factor signaling. Sci. Signal. 3, pe26 (2010).
Marshansky, V. & Futai, M. The V-type H+-ATPase in vesicular trafficking: targeting, regulation and function. Curr. Opin. Cell Biol. 20, 415–426 (2008).
Forgac, M. Vacuolar ATPases: rotary proton pumps in physiology and pathophysiology. Nat. Rev. Mol. Cell Biol. 8, 917–929 (2007).
Mindell, J. A. Lysosomal acidification mechanisms. Annu. Rev. Physiol. 74, 69–86 (2012).
Cruciat, C. M. et al. Requirement of prorenin receptor and vacuolar H+-ATPase-mediated acidification for Wnt signaling. Science 327, 459–463 (2010).
Buechling, T. et al. Wnt/Frizzled signaling requires dPRR, the Drosophila homolog of the prorenin receptor. Curr. Biol. 20, 1263–1268 (2010).
Hermle, T., Saltukoglu, D., Grunewald, J., Walz, G. & Simons, M. Regulation of Frizzled-dependent planar polarity signaling by a V-ATPase subunit. Curr. Biol. 20, 1269–1276 (2010).
Wielenga, V. J. et al. Expression of CD44 in Apc and Tcf mutant mice implies regulation by the WNT pathway. Am. J. Pathol. 154, 515–523 (1999).
Funayama, N., Fagotto, F., McCrea, P. & Gumbiner, B. M. Embryonic axis induction by the armadillo repeat domain of β-catenin: evidence for intracellular signaling. J. Cell Biol. 128, 959–968 (1995).
Nishi, T. & Forgac, M. The vacuolar (H+)-ATPases—nature’s most versatile proton pumps. Nat. Rev. Mol. Cell Biol. 3, 94–103 (2002).
Kinouchi, K. et al. The (pro)renin receptor/ATP6AP2 is essential for vacuolar H+-ATPase assembly in murine cardiomyocytes. Circ. Res. 107, 30–34 (2010).
O’Brien, C. A. et al. ID1 and ID3 regulate the self-renewal capacity of human colon cancer-initiating cells through p21. Cancer Cell 21, 777–792 (2012).
Bowman, E. J., Graham, L. A., Stevens, T. H. & Bowman, B. J. The bafilomycin/concanamycin binding site in subunit c of the V-ATPases from Neurospora crassa and Saccharomyces cerevisiae. J. Biol. Chem. 279, 33131–33138 (2004).
Gross, J. C., Chaudhary, V., Bartscherer, K. & Boutros, M. Active Wnt proteins are secreted on exosomes. Nat. Cell Biol. 14, 1036–1045 (2012).
Urbanelli, L. et al. Signaling pathways in exosomes biogenesis, secretion and fate. Genes 4, 152–170 (2013).
Yang, J. et al. Adenomatous polyposis coli (APC) differentially regulates β-catenin phosphorylation and ubiquitination in colon cancer cells. J. Biol. Chem. 281, 17751–17757 (2006).
Mauvezin, C., Nagy, P., Juhasz, G. & Neufeld, T. P. Autophagosome-lysosome fusion is independent of V-ATPase-mediated acidification. Nat. Commun. 6, 7007 (2015).
Montross, W. T., Ji, H. & McCrea, P. D. A β-catenin/engrailed chimera selectively suppresses Wnt signaling. J. Cell Sci. 113, 1759–1770 (2000).
Su, L. K. et al. Multiple intestinal neoplasia caused by a mutation in the murine homolog of the APC gene. Science 256, 668–670 (1992).
Kane, P. M. Disassembly and reassembly of the yeast vacuolar H+-ATPase in vivo. J. Biol. Chem. 270, 17025–17032 (1995).
Graf, R., Harvey, W. R. & Wieczorek, H. Purification and properties of a cytosolic V1-ATPase. J. Biol. Chem. 271, 20908–20913 (1996).
Smardon, A. M., Tarsio, M. & Kane, P. M. The RAVE complex is essential for stable assembly of the yeast V-ATPase. J. Biol. Chem. 277, 13831–13839 (2002).
Najdi, R., Holcombe, R. F. & Waterman, M. L. Wnt signaling and colon carcinogenesis: beyond APC. J. Carcinog. 10, 5 (2011).
Elias, J. E. & Gygi, S. P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
Gonsalves, F. C. et al. An RNAi-based chemical genetic screen identifies three small-molecule inhibitors of the Wnt/wingless signaling pathway. Proc. Natl Acad. Sci. USA 108, 5954–5963 (2011).
Kuhl, M. & Pandur, P. Dorsal axis duplication as a functional readout for Wnt activity. Methods Mol. Biol. 469, 467–476 (2008).
Park, J. I. et al. Telomerase modulates Wnt signalling by association with target gene chromatin. Nature 460, 66–72 (2009).
Sato, T. et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009).
Acknowledgements
The authors thank X. Wang and A. Sharma for technical assistance. This work was supported by the University of Texas McGovern Medical School (startup funding to R.K.M.), the Cancer Prevention Research Institute of Texas (RP140563 to J-.I.P.), the National Institutes of Health (R01 CA193297-01 to J-.I.P.; 5R01 GM107079 to P.D.M.; K01DK092320 to R.K.M.; R01 GM126048 to W.W.), the Department of Defense Peer Reviewed Cancer Research Program (CA140572 to J-.I.P.), a Duncan Family Institute for Cancer Prevention and Risk Assessment Grant (IRG-08-061-01 to J-.I.P.), a Center for Stem Cell and Developmental Biology Transformative Grant (MD Anderson Cancer Center to J-.I.P.), an Institutional Research Grant (MD Anderson Cancer Center to J-.I.P.), a New Faculty Award (MD Anderson Cancer Center Support Grant to J-.I.P.), a Metastasis Research Center Grant (MD Anderson to J-.I.P.) and a Uterine SPORE Career Enhancement Program (MD Anderson to J-.I.P.). The core facility (DNA sequencing and Genetically Engineered Moue Facility) was supported by an MD Anderson Cancer Center Support Grant (CA016672).
Author information
Authors and Affiliations
Contributions
Y-.S.J., S.J., H.Y.J., and J-.I.P. conceived and designed experiments. Y-.S.J., S.J., M.J.K., S.H.L., H.N.S., E.M.L., H-.Y.J., S.L., J.Z., J-.I.Y., H.J., W.W., R.K.M., and J-.I.P. performed experiments. J.Y.W. and S.K. provided samples. Y-.S.J., W.W., S.L., H-.Y.J., R.K.M., J.C., P.D.M. and J-.I.P. analysed data. Y-.S.J. and J-.I.P. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interest
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Activation of Wnt/β-catenin signalling by TMEM9.
(a-c), Screening of cell signallings affected by TMEM9. qRT-PCR of CRC cells (MSI [a] versus MSS [b]) and IECs (c). d, Activation of β-catenin by TMEM9 ectopic expression. HeLa cells (Ctrl versus TMEM9-FLAG) were analyzed for β-catenin protein half-life using cycloheximide (CHX; 100 μg/ml), and quantified by ImageJ. (e and f), Upregulation of β-catenin transcriptional activity by TMEM9 in IECs. 48hr after overexpression of TMEM9, cells were analyzed by luciferase activity (e) and AXIN2 qRT-PCR (f). (g–i), Decreased β-catenin transcription activity by shTMEM9. Depletion of endogenous TMEM9 using multiple shRNAs (#1–6) in HCT116. HCT116 cells were stably transduced with lentiviruses encoding six different shRNAs and analyzed by IB (g). Eleven CRC cells were analyzed for determination of the effect of TMEM9 on Wnt/β-catenin signalling hyperactivation. (h). TOP/FOP-FLASH luciferase activity (i). (j and k), Establishment of TMEM9 KO CRC cells. Exon2 of TMEM9 was deleted using CRISPR/Cas9 gene targeting. PCR genotyping of TMEM9 displayed deletion of TMEM9 (WT: 281bp; KO; 233bp; j). TMEM9 protein was not expressed in TMEM9 KO CRC cells (k). NS: Not significant; Experiments were performed three times with similar results; Error bars: mean ± S.D.; Two-sided unpaired t-test.
Supplementary Figure 2 TMEM9 facilitates assembly of v-ATPase.
(a), The endogenous interaction of TMEM9 with ATP6AP2 and APT6V0D1. Co-IP of HCT116 cells (TMEM9 WT versus KO). IgG H.C.: immunoglobulin heavy chain. Experiments were performed three times with similar results. (b), Oligomerization of TMEM9. 293T cells were transfected with each plasmid (TMEM9-FLAG or TMEM9-HA) and were analyzed by co-IP assays. Experiments were performed three times with similar results. (c), Subcellular localization of ectopically expressed TMEM9. HeLa cells were transfected with TMEM9-FLAG plasmid. After fixation cells were stained with FLAG antibody. (d and e), Decreased MVB acidification by TMEM9 depletion. CRC (d) and 293T (e) cells were transfected with shTMEM9-GFP or TMEM9-FLAG plasmid for 24hr, respectively. After transfection cells were stained with Lysotracker for monitoring of MVB acidification. (f), Expression of TMEM9B, ATP6AP2, and ATP6V0D1. GEO datasets (GDS2947) from NCBI were analyzed for each gene expression in normal intestine and the matched CRC samples (32 patient samples). Of note, TMEM9B and other v-ATPase subunits are not upregulated in CRC. Representative images of three independent experiments with similar results;; Scale bars=20 μm.
Supplementary Figure 3 TMEM9 activates Wnt/β-catenin signalling via v-ATPase-mediated lysosomal degradation of APC.
(a–d), Activation of Wnt/β-catenin signalling by TMEM9-activated v-ATPase. TMEM9-induced β-catenin stabilization via v-ATPase (a and b). HeLa cells stably expressing control Vec or TMEM9-FLAG were treated with BAF (10nM, 24hr) and analyzed by IB (a) and IF staining (b). The requirement of TMEM9-TMD for TMEM9-induced β-catenin reporter activation (c and d). 293T (c) and CRC (HT29 and SW620; d) cells were transfected with WT or TMD deleted MT (ΔTMD) TMEM9 plasmids and analyzed by luciferase assays. The firefly luciferase plasmids and SV40-renilla luciferase expression plasmids (internal control for the transfection efficiency) were transfected into CRC cells for measurement of luciferase activity. 24hr after transfection, cell lysates were assessed by using Dual luciferase assay kit (Promega). Then, the firefly luciferase activity was normalized by the renilla luciferase activity for quantification. e and f, ATP6AP2 depletion inhibits TMEM9-activated β-catenin reporter. 293T cells stably expressing shRNAs (shCtrl or shATP6AP2 [#1 and #2; two different shRNAs]) were confirmed by IB (e), and transfected with the β-catenin reporter and TMEM9 expression plasmids for luciferase assays (f). (g and h), No effect of ATP6AP2 and TMEM9 on β-catenin reporter activity in TMEM9 or ATP6AP2 depleted CRC cells. shCtrl, shTMEM9 (g), or shATP6AP2 (h) plasmids were co-transfected with ATP6AP2 (g) or TMEM9 (h) plasmids, respectively. i and j, Rescue of β-catenin transcription activity by β-catenin overexpression in ATP6AP2 depleted CRC cells. 24hr after transfection, cells were collected for assessment of luciferase activity (i) and AXIN2 expression (j). (k and l), Activation of β-catenin transcription activity by TMEM9 independently of Wnt agonist or antagonist. IECs (k) and CRC cells (l) were treated with Wnt3a (50ng/ml) or Dkk-1 (100 ng/ml) for 12 hr and analyzed for AXIN2 qRT-PCR. (m), Downregulation of Wnt/β-catenin signaling by shTMEM9 independently of Wnt ligand secretion. After transfection, cells were incubated with IWP-2 (2 μm) for 12 hr, and AXIN2 expression was analyzed by qRT-PCR. (n), Upregulated APC protein by TMEM9 depletion. TMEM9 WT and KO cells analyzed by IF staining. GFP-expression marks shTMEM9-transduced cells (green dotted line). Experiments were performed three times with similar results. Representative images of three independent experiments with similar results; Scale bars=20 μm; NS: Not significant; Error bars: mean ± S.D.; Two-sided unpaired t-test.
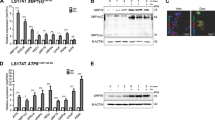
Supplementary Figure 4 Suppression of intestinal tumorigenesis by blockade of TMEM9.
(a), Correlation of TMEM9 expression with low survival in human CRC. The Kaplan–Meier plot of CRC specimens demonstrates significant (log-rank test) lower survival with TMEM9-high expression. Plots were analyzed from PrognoScan, a publicly available database (www.prognoscan.org). pmin: parallel minima, pcor: partial correlation. b and c, Transactivation of TMEM9 by Wnt/β-Catenin signaling. TMEM9 is required for CRC cell proliferation (b). Each CRC cell line (shCtrl versus shTMEM9) was analyzed for cell proliferation by cell counting. IF staining of HCT116 (shCtrl and shTMEM9; c) for Ki67. Experiments were performed three times with similar results. (d–f), Reduced CRC cell proliferation by TMEM9 depletion ex vivo. Each mouse (n = 4 biologically independent samples) was subcutaneously injected with 1×107 cells into the left flank (HT29 [control]) and the right flanks (TMEM9 KO-HT29). 15 days after transplantation, tumors were harvested for tumor assessment (d; n = 4 mice). HCT116 cells were subcutaneously injected into immunocompromised mice. 28 days later, tumors were collected for imaging and weight analyses (e; n = 8 mice). IF analyses of TMEM9-depleted tumors from xenograft (Ki67, CD44; f). (g), The targeting strategy of TMEM9 KO mouse model. (h), No defects in the Paneth cell differentiation by TMEM9 KO. IHC analysis of Lysozyme, a marker of Paneth cells, was performed in TMEM9 KO mouse. (i), Median survival of APCMIN, APCMIN:TMEM9±, and APCMIN:TMEM9-/- mice. (j–m), Suppression of intestinal tumorigenesis by TMEM9 KO. Cyclin D1 IHC of small intestine samples from APCMIN, APCMIN:TMEM9±, and APCMIN:TMEM9-/- mice (j). No alteration of cell death in APCMIN and APCMIN:TMEM9-/- tumors. IHC of cleaved caspase-3 (c-Cas3; k). EMT marker analysis of APCMIN and APCMIN:TMEM9-/- small intestine tumors. Markers of mesenchymal cell (N-cadherin and Vimentin) were not detected in tumors of both strains. The staining results of the mesenchymal cell in the APCMIN normal intestine served as a positive control. DAPI was stained for the nuclei. No increase in cell death by TMEM9 KO in vivo (l). No change in E-cadherin expression in APCMIN and APCMIN:TMEM9 KO adenomas (m). E-cadherin expression was monitored by Super Resolution Level-Confocal Microscope (LSM880-Airyscan). Representative images of three independent experiments with similar results; Scale bars=20 μm; NS: Not significant; Error bars: ± S.D. except for s5e ( ± S.E.M); Two-sided unpaired t-test. Centre: Average.
Supplementary Figure 5 Suppression of intestinal tumorigenesis by v-ATPase inhibitors.
(a), No defects in IEC differentiation by v-ATPase inhibitors. IHC for Chromogranin A (ChgA), a marker for the enteroendocrine cells, and Lysozyme, a marker for the Paneth cells in the non-tumor region of APCMIN mice treated with v-ATPase inhibitors. (b), Reduced CD44 expression and tumor cell growth by v-ATPase inhibitors. IHC for CD44 and Ki67. Representative images of three independent experiments with similar results; Scale bars=20 μm.
Supplementary Figure 6 Decreased PDX growth by v-ATPase inhibitor.
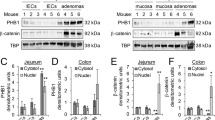
(a), Mutation status of PDXs. (b), Suppression of PDX growth by BAF. Immunocompromised mice (BALB/c nude) were subcutaneously transplanted with three different CRC tissues from the patients into both right and left flanks. 7 days after transplantation, mice were injected with vehicle (corn oil) or BAF (1 mg/kg and 3 mg/kg) every 3 days for 15 days. At 18 days post injection, CRC samples were collected for quantification. Experiment was performed once. (c), Decreased cell proliferation by BAF in PDXs. IHC for phosphorylated-Histone H3 (pHH3; a marker of mitosis) and Ki67 (a marker of proliferative cells), and H&E. (d), Increased APC expression by BAF. HT29-parental and HT29-APC KO cells served as a positive and negative control for IF staining of APC, respectively. BAF-treated PDXs displays the increased expression of APC protein. (e), Redistribution of β-catenin by BAF. BAF-treated PDXs exhibited the redistribution of β-catenin protein mainly in the cytosol and cell-cell adhesion, whereas control PDXs showed the nuclear localization of β-catenin. Representative images of three independent experiments with similar results; White scale bars=20 μm; Blue scale bars=200 μm; Red scale bars=1 cm.
Supplementary information
Supplementary information
Supplementary Figures 1–7 and Supplementary Table legends.
Supplementary Table 1
List of genes highly expressed in CRC.
Supplementary Table 2
Primer information.
Supplementary Table 3
Antibody information.
Supplementary Table 4
Statistics Source Data.
Rights and permissions
About this article
Cite this article
Jung, YS., Jun, S., Kim, M.J. et al. TMEM9 promotes intestinal tumorigenesis through vacuolar-ATPase-activated Wnt/β-catenin signalling. Nat Cell Biol 20, 1421–1433 (2018). https://doi.org/10.1038/s41556-018-0219-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0219-8
This article is cited by
-
Ubiquitin ligase subunit FBXO9 inhibits V-ATPase assembly and impedes lung cancer metastasis
Experimental Hematology & Oncology (2024)
-
Cellular senescence triggers intracellular acidification and lysosomal pH alkalinized via ATP6AP2 attenuation in breast cancer cells
Communications Biology (2023)
-
Activating Wnt/β-catenin signaling by autophagic degradation of APC contributes to the osteoblast differentiation effect of soy isoflavone on osteoporotic mesenchymal stem cells
Acta Pharmacologica Sinica (2023)
-
Reprogramming of palmitic acid induced by dephosphorylation of ACOX1 promotes β-catenin palmitoylation to drive colorectal cancer progression
Cell Discovery (2023)
-
Genome-wide analysis of genetic predisposition to common polygenic cancers
Journal of Applied Genetics (2022)