Abstract
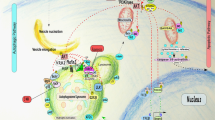
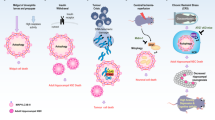
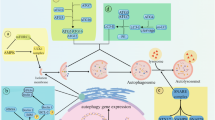
Autophagy influences cell survival through maintenance of cell bioenergetics and clearance of protein aggregates and damaged organelles. Several lines of evidence indicate that autophagy is a multifaceted regulator of cell death, but controversy exists over whether autophagy alone can drive cell death under physiologically relevant circumstances. Here, we review the role of autophagy in cell death and examine how autophagy interfaces with other forms of cell death including apoptosis and necrosis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Green, D. R. & Llambi, F. Cell death signalling. Cold Spring Harb. Perspect. Biol. 7, a006080 (2015).
Kerr, J. F., Wyllie, A. H. & Currie, A. R. Apoptosis: a basic biological phenomenon with wide-ranging implications in tissue kinetics. Br. J. Cancer 26, 239–257 (1972).
Galluzzi, L. et al. Molecular mechanisms of cell death: recommendations of the nomenclature committee on cell death 2018. Cell Death Differ. 25, 486–541 (2018).
Shi, Y. Mechanisms of caspase activation and inhibition during apoptosis. Mol. Cell 9, 459–470 (2002).
De Duve, C. & Wattiaux, R. Functions of lysosomes. Annu. Rev. Physiol. 28, 435–492 (1966).
Proskuryakov, S. Y., Konoplyannikov, A. G. & Gabai, V. L. Necrosis: a specific form of programmed cell death? Exp. Cell Res. 283, 1–16 (2003).
Denton, D., Nicolson, S. & Kumar, S. Cell death by autophagy: facts and apparent artefacts. Cell Death Differ. 19, 87–95 (2012).
Mizushima, N. & Komatsu, M. Autophagy: renovation of cells and tissues. Cell 147, 728–741 (2011).
Thumm, M. et al. Isolation of autophagocytosis mutants of Saccharomyces cerevisiae. FEBS Lett. 349, 275–280 (1994).
Tsukada, M. & Ohsumi, Y. Isolation and characterization of autophagy-defective mutants of Saccharomyces cerevisiae. FEBS Lett. 333, 169–174 (1993).
Russell, R. C. et al. ULK1 induces autophagy by phosphorylating Beclin-1 and activating VPS34 lipid kinase. Nat. Cell Biol. 15, 741–750 (2013).
Mizushima, N. et al. A protein conjugation system essential for autophagy. Nature 395, 395–398 (1998).
Mizushima, N., Noda, T. & Ohsumi, Y. Apg16p is required for the function of the Apg12p-Apg5p conjugate in the yeast autophagy pathway. EMBO J. 18, 3888–3896 (1999).
Kirisako, T. et al. Formation process of autophagosome is traced with Apg8/Aut7p in yeast. J. Cell Biol. 147, 435–446 (1999).
Huang, W. P., Scott, S. V., Kim, J. & Klionsky, D. J. The itinerary of a vesicle component, Aut7p/Cvt5p, terminates in the yeast vacuole via the autophagy/Cvt pathways. J. Biol. Chem. 275, 5845–5851 (2000).
Lang, T. et al. Aut2p and Aut7p, two novel microtubule-associated proteins are essential for delivery of autophagic vesicles to the vacuole. EMBO J. 17, 3597–3607 (1998).
Ichimura, Y. et al. A ubiquitin-like system mediates protein lipidation. Nature 408, 488–492 (2000).
Ohsumi, Y. Molecular dissection of autophagy: two ubiquitin-like systems. Nat. Rev. Mol. Cell Biol. 2, 211–216 (2001).
Glick, D., Barth, S. & Macleod, K. F. Autophagy: cellular and molecular mechanisms. J. Pathol. 221, 3–12 (2010).
Yu, L., Chen, Y. & Tooze, S. A. Autophagy pathway: Cellular and molecular mechanisms. Autophagy 14, 207–215 (2018).
Inoki, K., Zhu, T. & Guan, K. L. TSC2 mediates cellular energy response to control cell growth and survival. Cell 115, 577–590 (2003).
Kroemer, G. & Levine, B. Autophagic cell death: the story of a misnomer. Nat. Rev. Mol. Cell Biol. 9, 1004–1010 (2008).
Boya, P. et al. Inhibition of macroautophagy triggers apoptosis. Mol. Cell Biol. 25, 1025–1040 (2005).
Tanaka, Y. et al. Accumulation of autophagic vacuoles and cardiomyopathy in LAMP-2-deficient mice. Nature 406, 902–906 (2000).
Zhu, H. et al. The fusion of autophagosome with lysosome is impaired in L-arginine-induced acute pancreatitis. Int. J. Clin. Exp Pathol. 8, 11164–11170 (2015).
Gonzalez-Polo, R. A. et al. The apoptosis/autophagy paradox: autophagic vacuolization before apoptotic death. J. Cell Sci. 118, 3091–3102 (2005).
Schweichel, J. U. & Merker, H. J. The morphology of various types of cell death in prenatal tissues. Teratology 7, 253–266 (1973).
Fulda, S. & Kogel, D. Cell death by autophagy: emerging molecular mechanisms and implications for cancer therapy. Oncogene 34, 5105–5113 (2015).
Xu, T., Nicolson, S., Denton, D. & Kumar, S. Distinct requirements of autophagy-related genes in programmed cell death. Cell Death Differ. 22, 1792–1802 (2015).
Das, G., Shravage, B. V. & Baehrecke, E. H. Regulation and function of autophagy during cell survival and cell death. Cold Spring Harb Perspect. Biol. 4, a008813 (2012).
Neufeld, T. P. & Baehrecke, E. H. Eating on the fly: function and regulation of autophagy during cell growth, survival and death in Drosophila. Autophagy 4, 557–562 (2008).
Zhang, H. & Baehrecke, E. H. Eaten alive: novel insights into autophagy from multicellular model systems. Trends Cell Biol. 25, 376–387 (2015).
Anding, A. L. & Baehrecke, E. H. Autophagy in cell life and cell death. Curr. Top. Dev. Biol. 114, 67–91 (2015).
Zhao, Z. et al. Autophagosome-independent essential function for the autophagy protein Atg5 in cellular immunity to intracellular pathogens. Cell Host Microbe 4, 458–469 (2008).
Miller, B. C. et al. The autophagy gene ATG5 plays an essential role in B lymphocyte development. Autophagy 4, 309–314 (2008).
Subramani, S. & Malhotra, V. Non-autophagic roles of autophagy-related proteins. EMBO Rep. 14, 143–151 (2013).
Cadwell, K. & Debnath, J. Beyond self-eating: The control of nonautophagic functions and signaling pathways by autophagy-related proteins. J. Cell Biol. 217, 813–822 (2018).
Codogno, P., Mehrpour, M. & Proikas-Cezanne, T. Canonical and non-canonical autophagy: variations on a common theme of self-eating? Nat. Rev. Mol. Cell Biol. 13, 7–12 (2011).
Scarlatti, F., Maffei, R., Beau, I., Ghidoni, R. & Codogno, P. Non-canonical autophagy: an exception or an underestimated form of autophagy? Autophagy 4, 1083–1085 (2008).
Fuchs, Y. & Steller, H. Programmed cell death in animal development and disease. Cell 147, 742–758 (2011).
Nezis, I. P., Vaccaro, M. I., Devenish, R. J. & Juhasz, G. Autophagy in development, cell differentiation, and homeodynamics: from molecular mechanisms to diseases and pathophysiology. Biomed. Res. Int. 2014, 349623 (2014).
Jiang, C., Baehrecke, E. H. & Thummel, C. S. Steroid regulated programmed cell death during Drosophila metamorphosis. Development 124, 4673–4683 (1997).
Martin, D. N. & Baehrecke, E. H. Caspases function in autophagic programmed cell death in Drosophila. Development 131, 275–284 (2004).
Lee, C. Y., Simon, C. R., Woodard, C. T. & Baehrecke, E. H. Genetic mechanism for the stage- and tissue-specific regulation of steroid triggered programmed cell death in. Drosophila. Dev. Biol. 252, 138–148 (2002).
Lee, C. Y., Cooksey, B. A. & Baehrecke, E. H. Steroid regulation of midgut cell death during Drosophila development. Dev. Biol. 250, 101–111 (2002).
Berry, D. L. & Baehrecke, E. H. Growth arrest and autophagy are required for salivary gland cell degradation in Drosophila. Cell 131, 1137–1148 (2007).
Gorski, S. M. et al. A SAGE approach to discovery of genes involved in autophagic cell death. Curr. Biol. 13, 358–363 (2003).
Lee, C. Y. et al. Genome-wide analyses of steroid- and radiation-triggered programmed cell death in. Drosophila. Curr. Biol. 13, 350–357 (2003).
Lee, C. Y. & Baehrecke, E. H. Steroid regulation of autophagic programmed cell death during development. Development 128, 1443–1455 (2001).
Denton, D. et al. Autophagy, not apoptosis, is essential for midgut cell death in. Drosophila. Curr. Biol. 19, 1741–1746 (2009).
Chang, T. K. et al. Uba1 functions in Atg7- and Atg3-independent autophagy. Nat. Cell Biol. 15, 1067–1078 (2013).
Nezis, I. P. et al. Autophagic degradation of dBruce controls DNA fragmentation in nurse cells during late Drosophila melanogaster oogenesis. J. Cell Biol. 190, 523–531 (2010).
Gump, J. M. et al. Autophagy variation within a cell population determines cell fate through selective degradation of Fap-1. Nat. Cell Biol. 16, 47–54 (2014).
Hou, Y. C., Chittaranjan, S., Barbosa, S. G., McCall, K. & Gorski, S. M. Effector caspase Dcp-1 and IAP protein Bruce regulate starvation-induced autophagy during Drosophila melanogaster oogenesis. J. Cell Biol. 182, 1127–1139 (2008).
Scherfer, C., Han, V. C., Wang, Y., Anderson, A. E. & Galko, M. J. Autophagy drives epidermal deterioration in a Drosophila model of tissue aging. Aging 5, 276–287 (2013).
McPhee, C. K. & Baehrecke, E. H. Autophagy in Drosophila melanogaster. Biochim. Biophys. Acta 1793, 1452–1460 (2009).
Martin, D. N. et al. Proteomic analysis of steroid-triggered autophagic programmed cell death during Drosophila development. Cell Death Differ. 14, 916–923 (2007).
McPhee, C. K. et al. Identification of factors that function in Drosophila salivary gland cell death during development using proteomics. Cell Death Differ. 20, 218–225 (2013).
Batlevi, Y. et al. Dynein light chain 1 is required for autophagy, protein clearance, and cell death in Drosophila. Proc. Natl Acad. Sci. USA 107, 742–747 (2010).
Nelson, C., Ambros, V. & Baehrecke, E. H. miR-14 regulates autophagy during developmental cell death by targeting ip3-kinase 2. Mol. Cell 56, 376–388 (2014).
Dutta, S. & Baehrecke, E. H. Warts is required for PI3K-regulated growth arrest, autophagy, and autophagic cell death in Drosophila. Curr. Biol. 18, 1466–1475 (2008).
Denton, D. et al. UTX coordinates steroid hormone-mediated autophagy and cell death. Nat. Commun. 4, 2916 (2013).
Ress, C., Holtmann, M., Maas, U., Sofsky, J. & Dorn, A. 20-Hydroxyecdysone-induced differentiation and apoptosis in the Drosophila cell line, l(2)mbn. Tissue Cell 32, 464–477 (2000).
Tracy, K., Velentzas, P. D. & Baehrecke, E. H. Ral GTPase and the exocyst regulate autophagy in a tissue-specific manner. EMBO Rep. 17, 110–121 (2016).
McPhee, C. K., Logan, M. A., Freeman, M. R. & Baehrecke, E. H. Activation of autophagy during cell death requires the engulfment receptor Draper. Nature 465, 1093–1096 (2010).
Lin, L. et al. Complement-related regulates autophagy in neighbouring cells. Cell 170, 158–171 (2017).
McPhee, C. K. & Baehrecke, E. H. The engulfment receptor Draper is required for autophagy during cell death. Autophagy 6, 1192–1193 (2010).
Anding, A. L. et al. Vps13D encodes a ubiquitin-binding protein that is required for the regulation of mitochondrial size and clearance. Curr. Biol. 28, 287–295 (2018).
Seong, E. et al. Mutations in VPS13D lead to a new recessive ataxia with spasticity and mitochondrial defects. Ann. Neurol. 83, 1075–1088 (2018).
Gauthier, J. et al. Recessive mutations in >VPS13D cause childhood onset movement disorders. Ann. Neurol. 83, 1089–1095 (2018).
Santhanam, A. et al. Ecdysone-induced receptor tyrosine phosphatase PTP52F regulates Drosophila midgut histolysis by enhancement of autophagy and apoptosis. Mol. Cell Biol. 34, 1594–1606 (2014).
Dasari, S. K. et al. Death by over-eating: the Gaucher disease associated gene GBA1, identified in a screen for mediators of autophagic cell death, is necessary for developmental cell death in Drosophila midgut. Cell Cycle 16, 2003–2010 (2017).
Arakawa, S. et al. Role of Atg5-dependent cell death in the embryonic development of Bax/Bak double-knockout mice. Cell Death Differ. 24, 1598–1608 (2017).
Imagawa, Y., Saitoh, T. & Tsujimoto, Y. Vital staining for cell death identifies Atg9a-dependent necrosis in developmental bone formation in mouse. Nat. Commun. 7, 13391 (2016).
Shimizu, S. et al. Role of Bcl-2 family proteins in a non-apoptotic programmed cell death dependent on autophagy genes. Nat. Cell Biol. 6, 1221–1228 (2004).
Reef, S. et al. A short mitochondrial form of p19ARF induces autophagy and caspase-independent cell death. Mol. Cell 22, 463–475 (2006).
Yu, S. W. et al. Autophagic death of adult hippocampal neural stem cells following insulin withdrawal. Stem Cells 26, 2602–2610 (2008).
Xie, C. et al. Neuroprotection by selective neuronal deletion of Atg7 in neonatal brain injury. Autophagy 12, 410–423 (2016).
Yamaguchi, T. et al. The CCR4-NOT deadenylase complex controls Atg7-dependent cell death and heart function. Sci Signal 11, eaan3638 (2018).
Rojas-Rios, P. et al. Translational control of autophagy by orb in the Drosophila germline. Dev. Cell 35, 622–631 (2015).
Liu, Y. et al. Autosis is a Na+, K+-ATPase-regulated form of cell death triggered by autophagy-inducing peptides, starvation, and hypoxia-ischemia. Proc. Natl Acad. Sci. USA 110, 20364–20371 (2013).
Zhang, G., Luk, B. T., Hamidy, M., Zhang, L. & Spector, S. A. Induction of a Na+/K+-ATPase-dependent form of autophagy triggers preferential cell death of human immunodeficiency virus type-1-infected macrophages. Autophagy 2018, 1–17 (2018).
Elgendy, M., Sheridan, C., Brumatti, G. & Martin, S. J. Oncogenic Ras-induced expression of Noxa and Beclin-1 promotes autophagic cell death and limits clonogenic survival. Mol. Cell 42, 23–35 (2011).
Byun, J. Y. et al. The Rac1/MKK7/JNK pathway signals upregulation of Atg5 and subsequent autophagic cell death in response to oncogenic Ras. Carcinogenesis 30, 1880–1888 (2009).
Byun, J. Y. et al. Oncogenic Ras signals through activation of both phosphoinositide 3-kinase and Rac1 to induce c-Jun NH2-terminal kinase-mediated, caspase-independent cell death. Mol. Cancer Res. 7, 1534–1542 (2009).
Chen, Y., McMillan-Ward, E., Kong, J., Israels, S. J. & Gibson, S. B. Oxidative stress induces autophagic cell death independent of apoptosis in transformed and cancer cells. Cell Death Differ. 15, 171–182 (2008).
Liao, G. et al. Phycocyanin inhibits tumorigenic potential of pancreatic cancer cells: role of apoptosis and autophagy. Sci. Rep. 6, 34564 (2016).
Azad, M. B. et al. Hypoxia induces autophagic cell death in apoptosis-competent cells through a mechanism involving BNIP3. Autophagy 4, 195–204 (2008).
Yu, L. et al. Regulation of an ATG7-beclin 1 program of autophagic cell death by caspase-8. Science 304, 1500–1502 (2004).
Yu, L. et al. Autophagic programmed cell death by selective catalase degradation. Proc. Natl Acad. Sci. USA 103, 4952–4957 (2006).
Hou, W. et al. Autophagy promotes ferroptosis by degradation of ferritin. Autophagy 12, 1425–1428 (2016).
Chen, Z. et al. The autophagic degradation of Cav-1 contributes to PA-induced apoptosis and inflammation of astrocytes. Cell Death Dis 9, 771 (2018).
Thorburn, J. et al. Autophagy controls the kinetics and extent of mitochondrial apoptosis by regulating PUMA levels. Cell Rep. 7, 45–52 (2014).
Jiang, K. et al. Autophagic degradation of FOXO3a represses the expression of PUMA to block cell apoptosis in cisplatin-resistant osteosarcoma cells. Am. J. Cancer Res. 7, 1407–1422 (2017).
Young, M. M. et al. Autophagosomal membrane serves as platform for intracellular death-inducing signaling complex (iDISC)-mediated caspase-8 activation and apoptosis. J. Biol. Chem. 287, 12455–12468 (2012).
Basit, F., Cristofanon, S. & Fulda, S. Obatoclax (GX15–070) triggers necroptosis by promoting the assembly of the necrosome on autophagosomal membranes. Cell Death Differ. 20, 1161–1173 (2013).
Sakamaki, J. I. & Ryan, K. M. Autophagy Determines the Path on the TRAIL to Death. Dev. Cell 37, 291–293 (2016).
Goodall, M. L. et al. The autophagy machinery controls cell death switching between apoptosis and necroptosis. Dev. Cell 37, 337–349 (2016).
Yousefi, S. et al. Calpain-mediated cleavage of Atg5 switches autophagy to apoptosis. Nat. Cell Biol. 8, 1124–1132 (2006).
Zhu, Y. et al. Beclin 1 cleavage by caspase-3 inactivates autophagy and promotes apoptosis. Protein Cell 1, 468–477 (2010).
Wirawan, E. et al. Caspase-mediated cleavage of Beclin-1 inactivates Beclin-1-induced autophagy and enhances apoptosis by promoting the release of proapoptotic factors from mitochondria. Cell Death Dis. 1, e18 (2010).
Betin, V. M. & Lane, J. D. Caspase cleavage of Atg4D stimulates GABARAP-L1 processing and triggers mitochondrial targeting and apoptosis. J. Cell Sci. 122, 2554–2566 (2009).
Radoshevich, L. et al. ATG12 conjugation to ATG3 regulates mitochondrial homeostasis and cell death. Cell 142, 590–600 (2010).
Szondy, Z., Sarang, Z., Kiss, B., Garabuczi, E. & Koroskenyi, K. Anti-inflammatory mechanisms triggered by apoptotic cells during their clearance. Front. Immunol. 8, 909 (2017).
Green, D. R., Oguin, T. H. & Martinez, J. The clearance of dying cells: table for two. Cell Death Differ. 23, 915–926 (2016).
Qu, X. et al. Autophagy gene-dependent clearance of apoptotic cells during embryonic development. Cell 128, 931–946 (2007).
Mellen, M. A., de la Rosa, E. J. & Boya, P. The autophagic machinery is necessary for removal of cell corpses from the developing retinal neuroepithelium. Cell Death Differ. 15, 1279–1290 (2008).
Michaud, M. et al. Autophagy-dependent anticancer immune responses induced by chemotherapeutic agents in mice. Science 334, 1573–1577 (2011).
Ko, A. et al. Autophagy inhibition radiosensitizes in vitro, yet reduces radioresponses in vivo due to deficient immunogenic signalling. Cell Death Differ. 21, 92–99 (2014).
Konishi, A., Arakawa, S., Yue, Z. & Shimizu, S. Involvement of Beclin 1 in engulfment of apoptotic cells. J. Biol. Chem. 287, 13919–13929 (2012).
Huang, S., Jia, K., Wang, Y., Zhou, Z. & Levine, B. Autophagy genes function in apoptotic cell corpse clearance during C. elegans embryonic development. Autophagy 9, 138–149 (2013).
Ruck, A. et al. The Atg6/Vps30/Beclin 1 ortholog BEC-1 mediates endocytic retrograde transport in addition to autophagy in C. elegans. Autophagy 7, 386–400 (2011).
Li, W. et al. Autophagy genes function sequentially to promote apoptotic cell corpse degradation in the engulfing cell. J. Cell Biol. 197, 27–35 (2012).
Singh, S. R. et al. The lipolysis pathway sustains normal and transformed stem cells in adult Drosophila. Nature 538, 109–113 (2016).
Sanjuan, M. A., Milasta, S. & Green, D. R. Toll-like receptor signaling in the lysosomal pathways. Immunol. Rev. 227, 203–220 (2009).
Sanjuan, M. A. et al. Toll-like receptor signalling in macrophages links the autophagy pathway to phagocytosis. Nature 450, 1253–1257 (2007).
Martinez, J. et al. Microtubule-associated protein 1 light chain 3 alpha (LC3)-associated phagocytosis is required for the efficient clearance of dead cells. Proc. Natl Acad. Sci. USA 108, 17396–17401 (2011).
Kim, S. E. & Overholtzer, M. Autophagy proteins regulate cell engulfment mechanisms that participate in cancer. Semin. Cancer Biol. 23, 329–336 (2013).
Martinez, J. et al. Molecular characterization of LC3-associated phagocytosis reveals distinct roles for Rubicon, NOX2 and autophagy proteins. Nat. Cell Biol. 17, 893–906 (2015).
Huang, J. et al. Activation of antibacterial autophagy by NADPH oxidases. Proc. Natl Acad. Sci. USA 106, 6226–6231 (2009).
Ma, J., Becker, C., Lowell, C. A. & Underhill, D. M. Dectin-1-triggered recruitment of light chain 3 protein to phagosomes facilitates major histocompatibility complex class II presentation of fungal-derived antigens. J. Biol. Chem. 287, 34149–34156 (2012).
Romao, S. et al. Autophagy proteins stabilize pathogen-containing phagosomes for prolonged MHC II antigen processing. J. Cell Biol. 203, 757–766 (2013).
Florey, O., Kim, S. E., Sandoval, C. P., Haynes, C. M. & Overholtzer, M. Autophagy machinery mediates macroendocytic processing and entotic cell death by targeting single membranes. Nat. Cell Biol. 13, 1335–1343 (2011).
Gutierrez, M. G. et al. Autophagy is a defense mechanism inhibiting BCG and Mycobacterium tuberculosis survival in infected macrophages. Cell 119, 753–766 (2004).
Nakagawa, I. et al. Autophagy defends cells against invading group A Streptococcus. Science 306, 1037–1040 (2004).
Overholtzer, M. et al. A nonapoptotic cell death process, entosis, that occurs by cell-in-cell invasion. Cell 131, 966–979 (2007).
Sun, Q., Cibas, E. S., Huang, H., Hodgson, L. & Overholtzer, M. Induction of entosis by epithelial cadherin expression. Cell Res. 24, 1288–1298 (2014).
Wang, M. et al. Impaired formation of homotypic cell-in-cell structures in human tumor cells lacking alpha-catenin expression. Sci. Rep. 5, 12223 (2015).
Hamann, J. C. et al. Entosis is induced by glucose starvation. Cell Rep. 20, 201–210 (2017).
Sun, L. et al. TM9SF4 is a novel factor promoting autophagic flux under amino acid starvation. Cell Death Differ. 25, 368–379 (2018).
Bergeret, E. et al. TM9SF4 is required for Drosophila cellular immunity via cell adhesion and phagocytosis. J. Cell Sci. 121, 3325–3334 (2008).
Cornillon, S. et al. Phg1p is a nine-transmembrane protein superfamily member involved in dictyostelium adhesion and phagocytosis. J. Biol. Chem. 275, 34287–34292 (2000).
Lozupone, F. et al. TM9SF4 is a novel V-ATPase-interacting protein that modulates tumor pH alterations associated with drug resistance and invasiveness of colon cancer cells. Oncogene 34, 5163–5174 (2015).
Lozupone, F. et al. The human homologue of Dictyostelium discoideum phg1A is expressed by human metastatic melanoma cells. EMBO Rep. 10, 1348–1354 (2009).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Doherty, J., Baehrecke, E.H. Life, death and autophagy. Nat Cell Biol 20, 1110–1117 (2018). https://doi.org/10.1038/s41556-018-0201-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0201-5