Abstract
Microtubules are central elements of the eukaryotic cytoskeleton that often function as part of branched networks. Current models for branching include nucleation of new microtubules from severed microtubule seeds or from γ-tubulin recruited to the side of a pre-existing microtubule. Here, we found that microtubules can be directly remodelled into branched structures by the microtubule-remodelling factor SSNA1 (also known as NA14 or DIP13). The branching activity of SSNA1 relies on its ability to self-assemble into fibrils in a head-to-tail fashion. SSNA1 fibrils guide protofilaments of a microtubule to split apart to form daughter microtubules. We further found that SSNA1 localizes at axon branching sites and has a key role in neuronal development. SSNA1 mutants that abolish microtubule branching in vitro also fail to promote axon development and branching when overexpressed in neurons. We have, therefore, discovered a mechanism for microtubule branching and implicated its role in neuronal development.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The cryo-EM structure of the SSNA1–microtubule is available through EMDB with the accession code EMD-4188. The additional tomography images are available in Figshare (https://figshare.com/articles/Microtubule_branch_png/6809795). Source data for Figs. 1, 4, 6 and Supplementary Fig. 6 have been provided as Supplementary Table 3. Other data supporting the findings of this study such as the cryo-tomography data are available from the corresponding author on reasonable request.
References
Ishihara, K., Nguyen, P. A., Groen, A. C., Field, C. M. & Mitchison, T. J. Microtubule nucleation remote from centrosomes may explain how asters span large cells. Proc. Natl Acad. Sci. USA 111, 17715–17722 (2014).
Meunier, S. & Vernos, I. Acentrosomal microtubule assembly in mitosis: the where, when, and how. Trends. Cell Biol. 26, 80–87 (2016).
Petry, S. & Vale, R. D. Microtubule nucleation at the centrosome and beyond. Nat. Cell Biol. 17, 1089–1093 (2015).
Kalil, K. & Dent, E. W. Branch management: mechanisms of axon branching in the developing vertebrate CNS. Nat. Rev. Neurosci. 15, 7–18 (2014).
Lewis, T. L., Courchet, J. & Polleux, F. Cell biology in neuroscience: cellular and molecular mechanisms underlying axon formation, growth, and branching. J. Cell Biol. 202, 837–848 (2013).
Stiess, M. et al. Axon extension occurs independently of centrosomal microtubule nucleation. Science 327, 704–707 (2010).
Yu, W. et al. The microtubule-severing proteins spastin and katanin participate differently in the formation of axonal branches. Mol. Biol. Cell 19, 1485–1498 (2008).
Petry, S., Groen, A. C., Ishihara, K., Mitchison, T. J. & Vale, R. D. Branching microtubule nucleation in Xenopus egg extracts mediated by augmin and TPX2. Cell 152, 768–777 (2013).
Decker, F., Oriola, D., Dalton, B. & Brugués, J. Autocatalytic microtubule nucleation determines the size and mass of Xenopus laevis egg extract spindles. eLife 7, e31149 (2018).
Murata, T. et al. Microtubule-dependent microtubule nucleation based on recruitment of gamma-tubulin in higher plants. Nat. Cell Biol. 7, 961–968 (2005).
Janson, M. E., Setty, T. G., Paoletti, A. & Tran, P. T. Efficient formation of bipolar microtubule bundles requires microtubule-bound gamma-tubulin complexes. J. Cell Biol. 169, 297–308 (2005).
Goshima, G., Mayer, M., Zhang, N., Stuurman, N. & Vale, R. D. Augmin: a protein complex required for centrosome-independent microtubule generation within the spindle. J. Cell Biol. 181, 421–429 (2008).
Kamasaki, T. et al. Augmin-dependent microtubule nucleation at microtubule walls in the spindle. J. Cell Biol. 202, 25–33 (2013).
Sánchez-Huertas, C. et al. Non-centrosomal nucleation mediated by augmin organizes microtubules in post-mitotic neurons and controls axonal microtubule polarity. Nat. Commun. 7, 12187 (2016).
Pfannenschmid, F. et al. Chlamydomonas DIP13 and human NA14: a new class of proteins associated with microtubule structures is involved in cell division. J. Cell Sci. 116, 1449–1462 (2003).
Lai, C. K. et al. Functional characterization of putative cilia genes by high-content analysis. Mol. Biol. Cell 22, 1104–1119 (2011).
Goyal, U., Renvoisé, B., Chang, J. & Blackstone, C. Spastin-interacting protein NA14/SSNA1 functions in cytokinesis and axon development. PLoS ONE 9, e112428 (2014).
Blanchoin, L. et al. Direct observation of dendritic actin filament networks nucleated by Arp2/3 complex and WASP/Scar proteins. Nature 404, 1007–1011 (2000).
Volkmann, N. et al. Structure of Arp2/3 complex in its activated state and in actin filament branch junctions. Science 293, 2456–2459 (2001).
Itzhak, D. N., Tyanova, S., Cox, J. & Borner, G. H. Global, quantitative and dynamic mapping of protein subcellular localization. eLife 5, e16950 (2016).
Woodruff, J. B. et al. The centrosome is a selective condensate that nucleates microtubules by concentrating tubulin. Cell 169, 1066–1077.e10 (2017).
Wilde, A. & Zheng, Y. Stimulation of microtubule aster formation and spindle assembly by the small GTPase Ran. Science 284, 1359–1362 (1999).
Hyman, A. A., Salser, S., Drechsel, D. N., Unwin, N. & Mitchison, T. J. Role of GTP hydrolysis in microtubule dynamics: information from a slowly hydrolyzable analogue, GMPCPP. Mol. Biol. Cell 3, 1155–1167 (1992).
Bechstedt, S. & Brouhard, G. J. Doublecortin recognizes the 13-protofilament microtubule cooperatively and tracks microtubule ends. Dev. Cell 23, 181–192 (2012).
Des Georges, A. et al. Mal3, the Schizosaccharomyces pombe homolog of EB1, changes the microtubule lattice. Nat. Struct. Mol. Biol. 15, 1102–1108 (2008).
Ramos-Morales, F., Infante, C., Fedriani, C., Bornens, M. & Rios, R. M. NA14 is a novel nuclear autoantigen with a coiled-coil domain. J. Biol. Chem. 273, 1634–1639 (1998).
Price, H. P. et al. The orthologue of Sjögren’s syndrome nuclear autoantigen 1 (SSNA1) in Trypanosoma brucei is an immunogenic self-assembling molecule. PLoS ONE 7, e31842 (2012).
Rodríguez-Rodríguez, M. et al. Characterization of the structure and self-recognition of the human centrosomal protein NA14: implications for stability and function. Protein Eng. Des. Sel. 24, 883–892 (2011).
Janke, C. The tubulin code: molecular components, readout mechanisms, and functions. J. Cell Biol. 206, 461–472 (2014).
Wang, Q., Crevenna, A. H., Kunze, I. & Mizuno, N. Structural basis for the extended CAP-Gly domains of p150(glued) binding to microtubules and the implication for tubulin dynamics. Proc. Natl Acad. Sci. USA 111, 11347–11352 (2014).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. E. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Witte, H. & Bradke, F. The role of the cytoskeleton during neuronal polarization. Curr. Opin. Neurobiol. 18, 479–487 (2008).
Conde, C. & Cáceres, A. Microtubule assembly, organization and dynamics in axons and dendrites. Nat. Rev. Neurosci. 10, 319–332 (2009).
Kapitein, L. C. & Hoogenraad, C. C. Building the neuronal microtubule cytoskeleton. Neuron 87, 492–506 (2015).
Mizuno, N., Narita, A., Kon, T., Sutoh, K. & Kikkawa, M. Three-dimensional structure of cytoplasmic dynein bound to microtubules. Proc. Natl Acad. Sci. USA 104, 20832–20837 (2007).
Errico, A., Claudiani, P., D’Addio, M. & Rugarli, E. I. Spastin interacts with the centrosomal protein NA14, and is enriched in the spindle pole, the midbody and the distal axon. Hum. Mol. Genet. 13, 2121–2132 (2004).
Shelanski, M. L., Gaskin, F. & Cantor, C. R. Microtubule assembly in the absence of added nucleotides. Proc. Natl Acad. Sci. USA 70, 765–768 (1973).
Chrétien, D., Kenney, J. M., Fuller, S. D. & Wade, R. H. Determination of microtubule polarity by cryo-electron microscopy. Structure 4, 1031–1040 (1996).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat. Methods 10, 584–590 (2013).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Shang, Z. et al. High-resolution structures of kinesin on microtubules provide a basis for nucleotide-gated force-generation. eLife 3, e04686 (2014).
Zhang, R. & Nogales, E. A new protocol to accurately determine microtubule lattice seam location. J. Struct. Biol. 192, 245–254 (2015).
Zhang, R., Alushin, G. M., Brown, A. & Nogales, E. Mechanistic origin of microtubule dynamic instability and its modulation by EB proteins. Cell 162, 849–859 (2015).
Alushin, G. M. et al. High-resolution microtubule structures reveal the structural transitions in αβ-tubulin upon GTP hydrolysis. Cell 157, 1117–1129 (2014).
Grigorieff, N. Frealign: an exploratory tool for single-particle cryo-EM. Methods Enzymol. 579, 191–226 (2016).
Cardone, G., Heymann, J. B. & Steven, A. C. One number does not fit all: mapping local variations in resolution in cryo-EM reconstructions. J. Struct. Biol. 184, 226–236 (2013).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Crevenna, A. H. et al. Side-binding proteins modulate actin filament dynamics. eLife 4, e04599 (2015).
Wieczorek, M., Bechstedt, S., Chaaban, S. & Brouhard, G. J. Microtubule-associated proteins control the kinetics of microtubule nucleation. Nat. Cell Biol. 17, 907–916 (2015).
Kaech, S. & Banker, G. Culturing hippocampal neurons. Nat. Protoc. 1, 2406–2415 (2006).
Lahaye, X. et al. The capsids of HIV-1 and HIV-2 determine immune detection of the viral cDNA by the innate sensor cGAS in dendritic cells. Immunity 39, 1132–1142 (2013).
Magiera, M. M. & Janke, C. Investigating tubulin posttranslational modifications with specific antibodies. Methods Cell Biol. 115, 247–267 (2013).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Longair, M. H., Baker, D. A. & Armstrong, J. D. Simple Neurite Tracer: open source software for reconstruction, visualization and analysis of neuronal processes. Bioinformatics 27, 2453–2454 (2011).
Hollingworth, T. & Berry, M. Network analysis of dendritic fields of pyramidal cells in neocortex and Purkinje cells in the cerebellum of the rat. Phil. Trans. R. Soc. Lond. B 270, 227–264 (1975).
Vormberg, A., Effenberger, F., Muellerleile, J. & Cuntz, H. Universal features of dendrites through centripetal branch ordering. PLoS Comput. Biol. 13, e1005615 (2017).
Schnitzbauer, J., Strauss, M. T., Schlichthaerle, T., Schueder, F. & Jungmann, R. Super-resolution microscopy with DNA-PAINT. Nat. Protoc. 12, 1198–1228 (2017).
Huang, B., Wang, W., Bates, M. & Zhuang, X. Three-dimensional super-resolution imaging by stochastic optical reconstruction microscopy. Science 319, 810–813 (2008).
Xu, T. et al. SOAX: a software for quantification of 3D biopolymer networks. Sci. Rep. 5, 9081 (2015).
Acknowledgements
We thank C. Grashoff, E. Lorentzen and J. von Blume for assistance at various stages of the project, E. Conti, W. Baumeister and the imaging, biochemistry and cryo-EM core facilities for resources and infrastructure (Max Planck Institute of Biochemistry). We thank A. Carter (University of Cambridge) and C. Kelley (Max Planck Institute of Biochemistry) for careful proofreading of the manuscript and helpful discussions. We thank PICT-IBiSA@Orsay and S. Leboucher (Institut Curie, Orsay) for technical assistance. We are grateful to G. Woehlke (Technical University Munich, Germany) for combined efforts of purification of tubulin, and to C. Gonzalez-Billault and C. Villablanca (University of Chile, Santiago) for help with the analysis of neurons. We also acknowledge C. Sindelar and R. Zhang for providing the software package for microtubule analysis and giving guidance on its use. The EM map of the SSNA1–microtubule complex is deposited with EMDB ID: EMD-4188. N.M., R.J. and C.B. acknowledge the Max Planck Society for the financial support. This study was supported by the DFG through grants within GRK1721, MI 1745/1, and the European Research Council (ERC-CoG-724209 to N.M., ERC-StG-680241 to R.J.). N.M. is a recipient of an EMBO Young Investigator award and a Boehringer Ingelheim Foundation Plus 3 Program grant. R.J. is supported by the DFG Emmy Noether Program (DFG JU 2957/1-1). C.J. is supported by the Institut Curie, the CNRS, the ANR-10-LBX-0038, ANR-10-IDEX-0001-02 PSL, ANR-12-BSV2-0007, INCA 2013-1-PLBIO-02-ICR-1 and 2014-PLBIO-11-ICR-1. A.H.C. acknowledges LISBOA-01-0145-FEDER-007660 and FCT.
Author information
Authors and Affiliations
Contributions
N.B. and N.M. performed electron and light microscopy, designed mutant constructs and performed biochemistry experiments, analysed the data and prepared the figures. S.B. and M.M.M. performed experiments with neuron primary culture. N.B. and A.H.C. performed light microscopy experiments. N.B., C.B. and M.T. cloned and purified proteins. N.M., N.B., H.N., S.B., M.M.M. and C.J. analysed neuron data and G.C. facilitated the automation of neuron analysis. N.M., N.B. and H.N. performed electron microscopic data collection and analysed the data. H.N. and T.S. performed super-resolution light microscopy experiments and N.M., H.N., G.C., C.B and R.J. analysed the data. Experiments were designed by N.M. and the manuscript was written by N.M. with contributions from the other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Gallery of primary neurons (DIV3) showing the localization of SSNA1 (red) at the axon branching sites.
β3-tubulin is shown in green. These images are representing three independent experiments.
Supplementary Figure 2
(A) Purified CrSSNA1 FL observed under negative stain EM, showing no aggregation. Fibril formation can also occur concomitantly, which is shown in Fig. S5A. (B) SDS-PAGE of purified protein fragments reported in the study. (C) Circular dichroism spectra (Mean residue ellipticity (MRE: □) of the purified CrSSNA1 FL at different temperatures, showing well folded alpha-helical configurations. (D) Snapshots of microtubule branching during nucleation observed by cryo-EM in addition to the images shown in Fig. 2b. In addition to typically observed split of microtubules (“split”, see Fig. 2b), widely or narrowly opened split (“split-wide” and “split-narrow”), microtubules branched into 3 splits (“fork-like”) or a microtubule splitting both ways was observed. Red arrowheads indicate the split protofilaments or the cloud of molecules that are joining to the polymerizing microtubules. (E) Representative 2D class averages of microtubule branches (n = 226 branches) showing wide-range of branching formation. Due to the flexible junction points, 2D averages do not resolve details. (F) Gallery of cryo-electron tomographic slices of microtubule branches. At the branching points, the breakage of the protofilament lattice can be observed. These images represent 15 cryo-tomograms.
Supplementary Figure 3
(A) Dynamic microtubules (green) on microtubule seeds (red) in the presence of high concentration of SSNA1 (3 or 30 µM) without molecular crowding agent, to achieve globally concentrated conditions. At 3 µM, branch-like microtubules started appearing (20%, 115 out of 559 microtubules observed) and at 30 µM, 50% (448 out of 895 microtubules) had branch-like protrusion of microtubules. (B) Snapshots of microtubules showing ‘branch-like’ formations. Branches were categorized as ‘splitting’, ‘end-joining’, ‘side-branching’ or ‘dynamic-branching’. (C) Negative-stain EM snapshots of branched microtubules with conditions tested in a fluorescence microscopy-based dynamic assay, showing branching happens under various conditions. (D) Snapshot of branched microtubules protruding out of a nucleation center in the presence of GMPCPP, used for cryo-EM ultrastructure observation. (E) The sequence based alignment of the SSNA1 proteins. Secondary structure elements, based on the prediction from PHYRE2 are depicted below the sequences with red bars for α-helices. Coiled-coil prediction from the Marcoil server is shown above the sequences. The charged amino acids are colored in blue for the positive, and red for the negative charge. The colors or grey-scale (for uncharged amino acids) are intensified based on the degree of conservation of the amino acids. The green box highlights residues E20/E(D)21, and the red box shows the unstructured tail region, which are both essential for microtubule branching. (F) A snapshot of SSNA1-FL fibrils forming sheet-like structure after 8 hours and 24 hours of incubation. The red arrows show the direction of a fibril. Fibrils laterally assemble together making an ordered sheet. This allows us to assess the basic arrangement of individual fibrils, showing 11-nm of repeat within a fibril as indicated in power spectrum of an average of fibril sheets. These images are representing three independent experiments.
Supplementary Figure 4
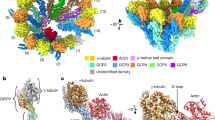
(A) Snapshots showing ‘cluster’ formation embedded in vitreous ice for cryo-EM observation. The centers of microtubule nucleation clusters have high densities of microtubules. (B) Fourier Shell Correlation (FSC) of the 3D reconstruction of the SSNA1-microtubule complex. The global resolution of the 3D reconstruction is 6.1 Å according to the FSC = 0.143 criteria, though it is only effective for the core of tubulin according to the local resolution mapping shown in D. (C and D) Local resolution representation of the 3D reconstruction according to the resolution-color code on the right color bar. (C) with a threshold that includes the SSNA1 decoration and (D) with the threshold that allows the visualization of the secondary structure elements in the tubulin core. While the tubulin core part shows a resolution ~ 6 Å, the decorated SSNA1 fibril is not resolved due to the symmetry mismatch between microtubules and SSNA1. The surface of the microtubules is not resolved either, presumably due to the coverage of SSNA1 knob-like pattern with 11 nm periodicity, blurring surrounding densities. Note the 11-nm knob-like pattern is averaged out due to the symmetry mismatch to the microtubule symmetry. (E) Chemical crosslinking of microtubules in the absence (top) and presence (bottom) of SSNA1. Subtilisin proteolyzes tubulin E-hooks, and subsequent crosslinking shows the loss of SSNA1-tubulin binding.
Supplementary Figure 5
(A) Gallery of electron micrographs of various CrSSNA1 truncation series tested for microtubule nucleation and branching. From left, observation of the purified protein at 0 h incubation (i.e. immediately after purification), 24 h incubation at RT, an overview after the addition of tubulin resulting in co-polymerized microtubules, and a magnified view of the copolymerized microtubules. Microtubule branching is shown with FL, 20-C and 1–105, while other protein fragments do not facilitate branching. For the proteins that do not cause the branching, examples of typical crossing of microtubules (red and blue bars at the scheme at the right column), instead of branching are shown. (B) Gallery of electron micrographs of various CrSSNA1 swap mutants. Left – all the mutants form cable-like fibrils. Right - branch formation is not induced with the E20K/D21K/K105E/K106E/K107E mutant. These images are representing three independent experiments.
Supplementary Figure 6
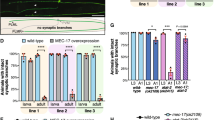
(A) Unmerged images of primary neurons (DIV3) overexpressing various SSNA1 variants shown in Fig. 6. The axons are labeled in red by Tau1 antibody (first row, red), while the dendrites are marked with the MAP2 antibody (second row, green). The expression of the SSNA1 proteins was confirmed by concomitant GFP expression (third row, cyan). In the merged view of the SSNA1 WT, axon is guided with a dotted line. (B-E) Neuron morphology analysis of various overexpression conditions B) Distribution of neurons based on total branch length/axon length and pie graphs showing the distribution of the number of processes (major branches plus minor protrusions along axon). Sample size: Control (n = 266 cells), wild type (n = 289 cells), 1–104 (n = 537 cells), 21-C (n = 274 cells), 5 A (n = 358 cells) pooled from 3 independent experiments and (C) Pie graphs showing the distribution of the total number of processes. Sample size: Control (n = 266 cells), wild type (n = 289 cells), 1–104 (n = 537 cells), 21-C (n = 274 cells), 5 A (n = 358 cells) pooled from 3 independent experiments. In (B) and (C), statistics of 5 A show significant difference (χ2 = 23.0, p < 0.001 and 36.3, p < 0.001 respectively) compared to control, indicating a negative effect of the 5 A mutant overexpression to neurons. In (C), wild type overexpressed neurons show the significant difference (χ2 = 12.83, p < 0.01). (D) Scatter dot plots of axon length under over-expression of various SSNA1 swap mutants. The promotion of axon development occurs in over-expression of swap-KK/EE, while slight dominant negative effect (shortening of axon) was observed in over-expression of swap-KK/EEE. Every cell is represented by a single point: Control (n = 1348 cells), swap-KK/EE (n = 789 cells), swap-KK/EEE (n = 1129 cells) and the overlaid box-and-whisker plots cover 50% (boxes) and 90% (whiskers) of the entire population, with median values indicated as lines within the boxes. (E) Pie graphs showing the distribution of the number of branches. Distributions of the branches in swap-mutants expressed neurons differ significantly from control (GFP over-expression) according to χ2 two-sample test (χ2 = 14.4, p < 0.005 and 29.1, p < 0.000005, respectively). Sample size: Control (n = 1348 cells), swap-KK/EE (n = 789 cells), swap-KK/EEE (n = 1127 cells). (F) Purification of chTOG and EB3. (G) Mixtures of tubulin with chTOG (upper) and EB3 (lower) were treated in the same way as SSNA1-tubulin mixture to test the induction of microtubule branches. No microtubule branching was observed in the tested conditions.
Supplementary information
Supplementary Information
Supplementary Figures 1–6, and legends for Supplementary Tables 1–3 and Supplementary Videos 1–4.
Supplementary Table 1
Series of truncations and mutations created and their microtubule branching activities.
Supplementary Table 2
Information of antibodies used in immunofluorescence and DNA-PAINT experiments.
Supplementary Table 3
Statistics source data.
Supplementary Video 1
Aster-like microtubule formation in the presence of SSNA1.
Supplementary Video 2
Nucleation of microtubules from the side of a microtubule with 100 nM SSNA1.
Supplementary Video 3
Nucleation of microtubules from the side of a microtubule with 30 µM SSNA1.
Supplementary Video 4
Nucleation of microtubules from the end of the microtubule seed.
Cryo-EM Table
Cryo-EM data collection, refinement and validation statistics.
Rights and permissions
About this article
Cite this article
Basnet, N., Nedozralova, H., Crevenna, A.H. et al. Direct induction of microtubule branching by microtubule nucleation factor SSNA1. Nat Cell Biol 20, 1172–1180 (2018). https://doi.org/10.1038/s41556-018-0199-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0199-8
This article is cited by
-
Feedback regulation of crystal growth by buffering monomer concentration
Nature Communications (2020)
-
From mechanical resilience to active material properties in biopolymer networks
Nature Reviews Physics (2019)