Abstract
Recent lineage tracing studies have revealed that mammary gland homeostasis relies on unipotent stem cells. However, whether and when lineage restriction occurs during embryonic mammary development, and which signals orchestrate cell fate specification, remain unknown. Using a combination of in vivo clonal analysis with whole mount immunofluorescence and mathematical modelling of clonal dynamics, we found that embryonic multipotent mammary cells become lineage-restricted surprisingly early in development, with evidence for unipotency as early as E12.5 and no statistically discernable bipotency after E15.5. To gain insights into the mechanisms governing the switch from multipotency to unipotency, we used gain-of-function Notch1 mice and demonstrated that Notch activation cell autonomously dictates luminal cell fate specification to both embryonic and basally committed mammary cells. These functional studies have important implications for understanding the signals underlying cell plasticity and serve to clarify how reactivation of embryonic programs in adult cells can lead to cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Wansbury, O. et al. Transcriptome analysis of embryonic mammary cells reveals insights into mammary lineage establishment. Breast Cancer Res. 13, R79 (2011).
Robinson, G. W. Cooperation of signalling pathways in embryonic mammary gland development. Nat. Rev. Genet. 8, 963–972 (2007).
Veltmaat, J. M., Mailleux, A. A., Thiery, J. P. & Bellusci, S. Mouse embryonic mammogenesis as a model for the molecular regulation of pattern formation. Differentiation 17, 1–17 (2001).
Watson, C. J. & Khaled, W. T. Mammary development in the embryo and adult: a journey of morphogenesis and commitment. Development 135, 995–1003 (2008).
Stingl, J., Raouf, A., Eirew, P. & Eaves, C. J. Deciphering the mammary epithelial cell hierarchy. Cell Cycle 5, 1519–1522 (2006).
Shackleton, M. et al. Generation of a functional mammary gland from a single stem cell. Nature 439, 84–88 (2006).
Rios, A. C., Fu, N. Y., Lindeman, G. J. & Visvader, J. E. In situ identification of bipotent stem cells in the mammary gland. Nature 506, 322–327 (2014).
Van Keymeulen, A. et al. Distinct stem cells contribute to mammary gland development and maintenance. Nature 479, 189–193 (2011).
Wang, D. et al. Identification of multipotent mammary stem cells by protein C receptor expression. Nature 517, 81–84 (2015).
Wuidart, A. et al. Quantitative lineage tracing strategies to resolve multipotency in tissue-specific stem cells. Genes Dev. 30, 1261–1277 (2016).
Rodilla, V. et al. Luminal progenitors restrict their lineage potential during mammary gland development. PLoS Biol. 13, e1002069 (2015).
Blanpain, C. & Simons, B. D. Unravelling stem cell dynamics by lineage tracing. Nat. Rev. Mol. Cell Biol. 14, 489–502 (2013).
Scheele, C. L. et al. Identity and dynamics of mammary stem cells during branching morphogenesis. Nature 542, 313–317 (2017).
Davis, F. M. et al. Single-cell lineage tracing in the mammary gland reveals stochastic clonal dispersion of stem/progenitor cell progeny. Nat. Commun. 7, 13053 (2016).
Koren, S. et al. PIK3CA(H1047R) induces multipotency and multi-lineage mammary tumours. Nature 525, 114–118 (2015).
Prater, M. D. et al. Mammary stem cells have myoepithelial cell properties. Nat. Cell Biol. 16, 942–950 (2014).
Blaas, L. et al. Lgr6 labels a rare population of mammary gland progenitor cells that are able to originate luminal mammary tumours. Nat. Cell Biol. 18, 1346–1356 (2016).
van Amerongen, R., Bowman, A. N. & Nusse, R. Developmental stage and time dictate the fate of Wnt/beta-catenin-responsive stem cells in the mammary gland. Cell Stem Cell 11, 387–400 (2012).
Bouras, T. et al. Notch signaling regulates mammary stem cell function and luminal cell-fate commitment. Cell Stem Cell 3, 429–441 (2008).
Choi, N., Zhang, B., Zhang, L., Ittmann, M. & Xin, L. Adult murine prostate basal and luminal cells are self-sustained lineages that can both serve as targets for prostate cancer initiation. Cancer Cell 21, 253–265 (2011).
Lu, C. P. et al. Identification of stem cell populations in sweat glands and ducts reveals roles in homeostasis and wound repair. Cell 150, 136–150 (2012).
Ousset, M. et al. Multipotent and unipotent progenitors contribute to prostate postnatal development. Nat. Cell Biol. 14, 1131–1138 (2012).
Fre, S. et al. Notch lineages and activity in intestinal stem cells determined by a new set of knock-in mice. PLoS One 6, e25785 (2011).
Howard, B. A. & Veltmaat, J. M. Embryonic mammary gland development; a domain of fundamental research with high relevance for breast cancer research. J. Mammary Gland Biol. 18, 89–91 (2013).
Zvelebil, M. et al. Embryonic mammary signature subsets are activated in Brca1-/- and basal-like breast cancers. Breast Cancer Res. 15, R25(2013).
Spike, B. T. et al. A mammary stem cell population identified and characterized in late embryogenesis reveals similarities to human breast cancer. Cell Stem Cell 10, 183–197 (2012).
Muzumdar, M. D., Tasic, B., Miyamichi, K., Li, L. & Luo, L. A global double-fluorescent Cre reporter mouse. Genesis 45, 593–605 (2007).
Asselin-Labat, M. L. et al. Gata-3 is an essential regulator of mammary-gland morphogenesis and luminal-cell differentiation. Nat. Cell Biol. 9, 201–209 (2007).
Boras-Granic, K., Dann, P. & Wysolmerski, J. J. Embryonic cells contribute directly to the quiescent stem cell population in the adult mouse mammary gland. Breast Cancer Res. 16, 487 (2014).
Mills, A. A. et al. p63 is a p53 homologue required for limb and epidermal morphogenesis. Nature 398, 708–713 (1999).
Moumen, M. et al. The mammary myoepithelial cell. Int J. Dev. Biol. 55, 763–771 (2011).
Sun, P., Yuan, Y., Li, A., Li, B. & Dai, X. Cytokeratin expression during mouse embryonic and early postnatal mammary gland development. Histochem. Cell Biol. 133, 213–221 (2010).
Wendling, O., Bornert, J. M., Chambon, P. & Metzger, D. Efficient temporally-controlled targeted mutagenesis in smooth muscle cells of the adult mouse. Genesis. 47, 14–18 (2009).
Snippert, H. J. et al. Intestinal crypt homeostasis results from neutral competition between symmetrically dividing Lgr5 stem cells. Cell 143, 134–144 (2010).
Wang, C., Christin, J. R., Oktay, M. H. & Guo, W. Lineage-biased stem cells maintain estrogen-receptor-positive and -negative mouse mammary luminal lineages. Cell Rep. 18, 2825–2835 (2017).
Van Keymeulen, A. et al. Lineage-restricted mammary stem cells sustain the development, homeostasis, and regeneration of the estrogen receptor positive lineage. Cell Rep. 20, 1525–1532 (2017).
Murtaugh, L. C., Stanger, B. Z., Kwan, K. M. & Melton, D. A. Notch signaling controls multiple steps of pancreatic differentiation. Proc. Natl Acad. Sci. USA 100, 14920–14925 (2003).
Van Keymeulen, A. et al. Reactivation of multipotency by oncogenic PIK3CA induces breast tumour heterogeneity. Nature 525, 119–123 (2015).
Indra, A. K. et al. Temporally-controlled site-specific mutagenesis in the basal layer of the epidermis: comparison of the recombinase activity of the tamoxifen-inducible Cre-ER(T) and Cre-ER(T2) recombinases. Nucleic Acids Res. 27, 4324–4327 (1999).
Lim, E. et al. Transcriptome analyses of mouse and human mammary cell subpopulations reveal multiple conserved genes and pathways. Breast Cancer Res. 12, R21 (2010).
Ye, X. et al. Distinct EMT programs control normal mammary stem cells and tumour-initiating cells. Nature 525, 256–260 (2015).
Wuidart, A. et al. Early lineage segregation of multipotent embryonic mammary gland progenitors https://doi.org/10.1038/s41556-018-0095-2 (2018).
Lee, M. Y. et al. Ectodermal influx and cell hypertrophy provide early growth for all murine mammary rudiments, and are differentially regulated among them by Gli3. PLoS One 6, e26242 (2011).
Tadeu, A. M. & Horsley, V. Notch signaling represses p63 expression in the developing surface ectoderm. Development 140, 3777–3786 (2013).
Blanpain, C. & Fuchs, E. Stem cell plasticity. Plasticity of epithelial stem cells in tissue regeneration. Science 344, 1242281 (2014).
Yalcin-Ozuysal, O. et al. Antagonistic roles of Notch and p63 in controlling mammary epithelial cell fates. Cell Death Differ. 17, 1600–1612 (2010).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10 (2011).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq--a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Ritchie, M. E. et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43, 47–47 (2015).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Acknowledgements
The authors thank S. Artavanis-Tsakonas for sharing the N1CreERT2 mice and P. Chambon and D. Metzger for providing the SMACreERT2 (Acta2-CreERT2) and K5-CreERT2 mice, and S. Tajbakhsh for the mTmG reporter line and H. Clevers for sharing the Confetti reporter mouse. The authors thank members of the Glukhova laboratory, especially M. Faraldo and M.-A. Deugnier, for technical advice and constructive discussions. The authors acknowledge the Flow Cytometry and Cell Sorting Platform at Institute Curie for their expertise, in particular Z. Maciorowski; the In Vivo Experimental Facility, mainly S. Jannet, for help in the maintenance and care of our mouse colony; and the Experimental Pathology facility at Curie Hospital for paraffin sample preparation. This work was supported by Paris Sciences et Lettres (PSL* Research University), the French National Research Agency (ANR) grant no.s ANR-15-CE13-0013-01, the Canceropole Ile-de-France (grant no. 2015-2-APD-01-ICR-1) and by Labex DEEP ANR-Number 11-LBX-0044 (to S.F.), the Wellcome Trust (grant no. 098357/Z/12/Z to B.D.S. and 110326/Z/15/Z to E.H.). A.M.L. was funded by a post-doctoral fellowship from the Fondation de France. E.H. was funded by a Junior Research Fellowship from Trinity College (Cambridge University), a Sir Henry Wellcome Fellowship from the Wellcome Trust, and acknowledges the Bettencourt-Schueller Young Researcher Prize for support. The PICT-IBiSA imaging platform was funded by ANR-10-INBS-04 (France-BioImaging), ANR-11 BSV2 012 01, ERC ZEBRATECTUM no. 311159, ARC SFI20121205686 and the Schlumberger Foundation. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
A.M.L, V.R., M.H., E.H. and S.F. conceived and designed the experiments and analysed the data. A.M.L, V.R., C.L. and M.H. performed all experiments. O.R. and O.L. performed the 3D image analysis. S.R. analysed the RNA sequencing data. A.M.L, V.R., E.H., B.D.S and S.F. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
All authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated Supplementary Information
Supplementary Figure 1 All embryonic mammary cells express Notch1.
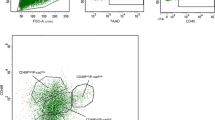
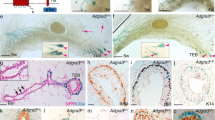
a. Representative sections of embryonic buds of N1Cre/mTmG embryos induced with tamoxifen at E15.5 and analysed 24h later by RNAscope in situ hybridisation for Notch1 (red punctate dots, left panel in a) or POLR2A (red punctate dots, right panel in a) followed by immunofluorescence anti-GFP (to detect N1Cre-labelled cells in green) and DAPI (to stain nuclei in blue). n = 3 biologically independent animals. Scale bars correspond to 20 μm. b. Representative dot plots of the used FACS gating strategy. First, FCS/SSC gating allows discarding cell debris (1), then SSC-A/SSC-W selects single cells (2), DAPI exclusion selects live cells (3), Lin exclusion (CD45/CD31/Ter118) eliminates hematopoietic cells (4), CD29/CD24 are used to visualise Mammary Epithelial Cells (MEC) (5) and GFP/Tomato selects fluorescent cells (6).
Supplementary Figure 2 ERαpos and ERαneg LC cells are maintained by unipotent progenitors.
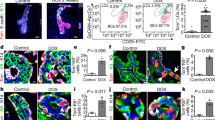
a. FACS dot plots showing CD133 and GFP (left panel) or Sca1 and GFP (right panel) expression in N1Cre/mTmG mice 6 weeks after tamoxifen induction at E13.5. CD133pos and Sca1pos cells (corresponding to ERαpos cells) are gated within green boxes, while ERαneg cells are gated within black boxes. n=3 biologically independent animals. b. FACS quantification of the percentage of CD133pos (left panel) and Sca1pos (right panel) total luminal (LC in red) or Notch1-derived lineages (GFP in green) 6 weeks after tamoxifen induction of N1Cre/mTmG mice at the indicated developmental times. n=5, 3, 6, 4, 7, and 3 biologically independent animals for N1Cre/mTmG induced at E13.5, P0.5, P3, P10, P15 and P21, respectively. Left panel from left to right: p=0.23, p= 0.06, p=0.0003, p=0.003; p=6.93e-08, p=0.001); right panel from left to right: p=0.055, p=0.064, p=7.8e-05, p=0.018, p=3.25e-07, p=0.021, using two-tailed unpaired multiple t-test. Graphs indicate average values ± SEM. c. Representative immunofluorescent staining for ERα (in green) in 6-week-old N1Cre/Confetti mice after tamoxifen induction at P0. Notch1-derived lineages are labelled in red by RFP. DAPI stains nuclei in blue. n=2 biologically independent animals. Scale bars correspond to 20 μm. Source data are available in Supplementary Table 1.
Supplementary Figure 3 Recombination efficiency of N1Cre/Confetti mice at different embryonic developmental times and tamoxifen doses.
Representative single Z-stacks of wholemount immunostaining of embryonic mammary buds from N1Cre/Confetti mice induced with tamoxifen at E13.5 (a), and E17.5 (b) at doses 0.1mg (upper images), or 2 µg (lower image), and analysed 48h later. Immunostaining for K5 is shown in white. Scale bars correspond to 100 μm, and to 50 μm in the magnifications; n=20 glands in 11 embryos (a) and 31 glands in 13 embryos (b). c-e. Quantification of the number of cells per coloured clone (c), total number of cells per gland (d) or number of colours per gland (e) at 48h-chase from females induced at the indicated time points during embryogenesis with full dose (0.1 mg) or diluted doses (0.05 mg or 2 µg) of tamoxifen; n=98 glands in 30 embryos. p=0.0004 and p=0.036 (c); p<0.0001 and p=0.004 (d); p=0.0001 and p=0.0118 (e). Graphs indicate average values ± SEM. *p<0.05, **p<0.01, ***p< 0.001, ****p< 0.0001 using two-tailed non-parametric Mann Whitney t-test. Source data are available in Supplementary Table 1.
Supplementary Figure 4 Wholemount immunostaining of N1Cre/Confetti mammary glands.
a-d. Representative single Z-stacks of wholemount immunostaining of mammary glands from N1Cre/Confetti mice induced with 0.1mg of tamoxifen at E13.5 and analysed at P7 (a; n=5 biologically independent animals); with 0.1 mg of tamoxifen at E15.5 and analysed at P7 (b; n=2 biologically independent animals) and with 0.1mg (c; n=3 biologically independent animals) or 2μg (d; n=3 biologically independent animals) of tamoxifen at E17.5 and analysed at P7. Immunostaining for K5 in white marks basal cells. Immunofluorescence for GFP (green), Cyan (blue), YFP (yellow) and RFP (red) represents Notch1 lineages considered as unique clones derived from a single embryonic stem cell. The number of floxed cells is directly proportional to the dose of tamoxifen used to induce Cre recombination (compare c and d). Images were acquired with 3x3 tiles (a, d), 5x4 tiles (b) and 2x3 tiles (c) and stitched without overlap by juxtaposition using the Zeiss software Zen Black 14.0.8.201. The dotted square in b demarcates 2 tiles that were not imaged as there was no epithelium in that area. Scale bars correspond to 100 μm.
Supplementary Figure 5 Statistical analysis of bipotency and distribution of coloured clones at different embryonic time points.
a. Quantification of the percentage of exclusively unipotent clones in any of the four confetti colours scored post-natally (between P7 and P15) in N1Cre/Confetti mice induced with tamoxifen at the indicated embryonic stages. 46 analysed clones from n=23 distinct glands from 8 embryos of 5 litters for E12; 33 analysed clones from n=14 distinct glands from 5 embryos of 3 litters for E13; 25 analysed clones from n=11 distinct glands from 5 embryos of 3 litters for E15 diluted 1:2; 44 analysed clones from n=17 distinct glands from 3 embryos for E17 diluted 1:50. p=0.0132 with unpaired t test with Welch’s correction. b. Comparison between the theoretical probability of “chance bipotency” arising from merging of individual labelling events of unipotent progenitors (in red) and the experimentally observed bipotency (in grey), at different embryonic induction times and tamoxifen doses. Females were injected with tamoxifen 0.1mg/g of mouse body weight at E12 (n=8 mice, 66.42±6.19, p<0.0001), E13 (n=5 mice, 51.29±18.55, p<0.0001), and E15 0.1mg (n=5 mice, 73.61±6.94, p=0.5), with 0.05 mg/g of mouse body weight at E15 0.05 mg (n=3 mice, 39.16±16.54, p=0.12) and with 2µg/g of mouse body weight at E17 (n=3 mice, 45.37±10.67, p=0.43). All colours were grouped together in this analysis. c-f. Comparisons between the theoretical probability of “chance” bipotency arising from merging of individual labelling events of unipotent progenitors (in red) and the experimentally observed bipotency (in grey) in the same animals analysed in b, for each individual confetti colour, and at the indicated induction times: E12 (c; n=6, 5 and 7 animals presenting GFP, YFP and RFP clones respectively out of 8 mice, p=0.008, p=0.013 and p=0.0027), E13 (d; n=5, 4 and 5 animals presenting GFP, YFP and RFP clones respectively out of 5 mice, p=0.003, p=0.005 and p=0.12), E15 (e; n=5, 3 and 5 animals presenting GFP, YFP and RFP clones respectively out of 5 mice, p=0.99, p=0.58 and p=0.52) and E17 (f; n=3, 3 and 3 animals presenting GFP, YFP and RFP clones respectively out of 3 mice, p=0.31, p=0.99 and p=0.99). g-h. Theoretical relationship between the average number of independent recombination events per mammary gland and the average number of colours per gland (g) and the average number of colours per gland and the fraction of “chance” bipotency (h). These curves are generated from stochastic simulations of random induction, using the relative chimerism extracted either from the E12 (purple line, GFP: 41.4%, YFP: 21.4%, RFP: 34.3%, CFP: 2.9%) or E17 dataset (green line, GFP: 45.2%, YFP: 24.2%, RFP: 29.0%, CFP: 1.6%). Error bars represent mean and SD. P-values were calculated from two-tailed binomial tests. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001. i-k. Comparisons between the theoretical probabilities (in grey) and the observed colour recombination (in red) to get 0, 1, 2, 3 or 4 different colours in the same gland in N1Cre/Confetti females induced at E13.5 (i; n=5 mice), E15.5 (j; n= 5 mice) and E17.5 (k; n=3 mice) and analysed at P7. Non-statistical significance has been observed between the predicted model and the observed quantifications. l-m. Comparisons between the theoretical (in red) and the observed potency (in grey) for E15.5: 0.22±0.07, 0.72±0.11, 0.04±0.06 (l) and E17.5: 0.46±0.13, 0.35±0.19, 0.18±0.06 (m) inductions, segregating unipotent basal and unipotent luminal in separate categories. No statistical differences between experimental and predicted distributions; n=5 biologically independent animals, p=0.25 at E15 in l, n=3 biologically independent animals, p=0.79 at E17 in m, using Chi-squared tests, demonstrating robustness of our results. Error bars represent mean±SD. p-values, unless explicitly specified otherwise were calculated from two-tailed binomial tests. Source data are available in Supplementary Table 1.
Supplementary Figure 6 Distribution of epithelial and stromal populations after embryonic Notch1 activation.
a. FACS dot plots showing LC and BC distribution gated in Lin (CD31/Ter119/CD45)neg from N1Cre/mTmG (left panel) and N1Cre/N1ICD (right panel) mice 6 weeks after tamoxifen induction at E13.5. Right panel in a: quantification of the proportion of LC (in red, CD24posCD29low) and BC (in blue, CD24posCD29high) analysed in MEC (Mammary Epithelial Cells, in orange gate in FACS plots) comparing N1Cre/mTmG and N1Cre/N1ICD. Graphs indicate average values ± SEM; non-significant differences were observed in the LC/BC distribution in MEC with unpaired Welch's t test (n=5 and 6 biologically independent animals, respectively). N1Cre/mTmG: 24.04±4.34 (BC) and 75.96±4.34 (LC); N1Cre/N1ICD: 23.77±1.90 (BC) and 77.43±1.90 (LC). b. Representative immunofluorescent staining for K8 in red) in mammary buds of N1Cre/mTmG females induced at E11.5 and analysed 24h later. Green indicates the Notch1-expressing cells and DAPI stains the nuclei in blue; n=2 biologically independent animals. The insets show magnifications of two GFP-labelled cells (’ and ’’) within the mammary epithelium. c. Representative immunofluorescent staining for K8 in red) and K14 (in white) in mammary glands from N1Cre/N1ICD females induced at E13.5 and analysed 6 weeks later. Nuclear GFP labels the progeny derived from Notch1pos cells targeted during embryogenesis. White arrows indicate stromal GFPpos cells; n=3 biologically independent animals. Scale bars correspond to 20 μm. d. FACS dot plots showing the distribution of GFPpos cells within the stromal gate (circled in black, CD24neg/CD29pos) from N1Cre/mTmG (left panel) and N1Cre/N1ICD (right panel) mice induced at E13.5 and analysed after 6 weeks. e. Percentage of GFPpos cells within the stromal compartment comparing mammary glands from N1Cre/mTmG and N1Cre/N1ICD. Graphs indicate average values ± SEM; non-significant differences were observed (n=5 and 6 biologically independent animals, respectively). *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001, using multiple unpaired t-test. Source data are available in Supplementary Table 1.
Supplementary Figure 7 Notch1 ectopic activation in basal cells induces a progressive reprogramming towards a luminal phenotype.
a. Representative FACS dot plots and quantification of the percentage of BC (blue gate) within the GFPpos population of SMACre/N1ICD mice induced with tamoxifen at P21 and analysed after 72h, 1 week, 3 weeks, 4 weeks, 5 weeks and 6 weeks, as indicated; n=22 biologically independent animals. b. Representative cryosections of mammary ducts from SMACre/N1ICD females induced with tamoxifen at P21 and analysed at the same chase time points as in (a). Upper panels: K8 expression is shown in red and K14 in white; middle panels: K8 expression is shown in red and EdU incorporation in white; bottom panels: K8 expression is shown in red and Cleaved caspase-3 in white. SMA-derived cells expressing N1ICD are labelled in green by nuclear GFP and DAPI stains nuclei in all panels. The arrow in the P21+3w upper panel indicates a GFPpos cell co-expressing luminal K8 and basal K14. Insets on the right of each panel represent magnifications of the boxed areas. Scale bars correspond to 20 μm and to 10 μm in the magnifications. Graphs represent the quantification of GFPpos and GFPneg cells expressing cleaved caspase-3 (n=5 for P21+72h and 6 for P21+1w independent immunofluorescence sections), or the proportion of GFPpos cells expressing K14 only (grey), K8 only (in red) or co-expressing K14 and K8 (in yellow) at each chase time point (n=3, 4, 2, 2, 2 and 3 independent immunofluorescence sections). Upper graph indicates average values ± SEM: 11.78±6.51 and 11.11±8.24 Caspase-3pos/GFPpos cells after 72h and 1-week chase, respectively, and 88.22±6.51 and 88.88±8.24 Caspase-3pos/GFPneg cells after 72h and 1-week chase, respectively. Source data are available in Supplementary Table 1.
Supplementary information
Supplementary Information
Supplementary Figures 1–7, Supplementary Table legends
Supplementary Table 1
Statistics source data
Rights and permissions
About this article
Cite this article
Lilja, A.M., Rodilla, V., Huyghe, M. et al. Clonal analysis of Notch1-expressing cells reveals the existence of unipotent stem cells that retain long-term plasticity in the embryonic mammary gland. Nat Cell Biol 20, 677–687 (2018). https://doi.org/10.1038/s41556-018-0108-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0108-1
This article is cited by
-
Extracting, filtering and simulating cellular barcodes using CellBarcode tools
Nature Computational Science (2024)
-
Lineage plasticity enables low-ER luminal tumors to evolve and gain basal-like traits
Breast Cancer Research (2023)
-
Kindlin-2 in myoepithelium controls luminal progenitor commitment to alveoli in mouse mammary gland
Cell Death & Disease (2023)
-
Reconstruction of dynamic mammary mini gland in vitro for normal physiology and oncogenesis
Nature Methods (2023)
-
Modeling the complexity of mammary gland in vitro
Nature Methods (2023)