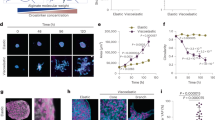
Abstract
It has long been proposed that the cell cycle is regulated by physical forces at the cell–cell and cell–extracellular matrix (ECM) interfaces1,2,3,4,5,6,7,8,9,10,11,12. However, the evolution of these forces during the cycle has never been measured in a tissue, and whether this evolution affects cell cycle progression is unknown. Here, we quantified cell–cell tension and cell–ECM traction throughout the complete cycle of a large cell population in a growing epithelium. These measurements unveil temporal mechanical patterns that span the entire cell cycle and regulate its duration, the G1–S transition and mitotic rounding. Cells subjected to higher intercellular tension exhibit a higher probability to transition from G1 to S, as well as shorter G1 and S–G2–M phases. Moreover, we show that tension and mechanical energy are better predictors of the duration of G1 than measured geometric properties. Tension increases during the cell cycle but decreases 3 hours before mitosis. Using optogenetic control of contractility, we show that this tension drop favours mitotic rounding. Our results establish that cell cycle progression is regulated cooperatively by forces between the dividing cell and its neighbours.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Huang, S., Chen, C. S. & Ingber, D. E. Control of cyclin D1, p27Kip1, and cell cycle progression in human capillary endothelial cells by cell shape and cytoskeletal tension. Mol. Biol. Cell 9, 3179–3193 (1998).
Nelson, C. M. et al. Emergent patterns of growth controlled by multicellular form and mechanics. Proc. Natl Acad. Sci. USA 102, 11594–11599 (2005).
Klein, E. A. et al. Cell-cycle control by physiological matrix elasticity and in vivo tissue stiffening. Curr. Biol. 19, 1511–1518 (2009).
Mih, J. D., Marinkovic, A., Liu, F., Sharif, A. S. & Tschumperlin, D. J. Matrix stiffness reverses the effect of actomyosin tension on cell proliferation. J. Cell Sci. 125, 5974–5983 (2012).
Aragona, M. et al. A mechanical checkpoint controls multicellular growth through YAP/TAZ regulation by actin-processing factors. Cell 154, 1047–1059 (2013).
Streichan, S. J., Hoerner, C. R., Schneidt, T., Holzer, D. & Hufnagel, L. Spatial constraints control cell proliferation in tissues. Proc. Natl Acad. Sci. USA 111, 5586–5591 (2014).
LeGoff, L. & Lecuit, T. Mechanical forces and growth in animal tissues. Cold Spring Harb. Perspect. Biol. 8, a019232 (2015).
Benham-Pyle, B. W., Pruitt, B. L. & Nelson, W. J. Cell adhesion. Mechanical strain induces E-cadherin-dependent Yap1 and β-catenin activation to drive cell cycle entry. Science 348, 1024–1027 (2015).
Pinheiro, D. et al. Transmission of cytokinesis forces via E-cadherin dilution and actomyosin flows. Nature 545, 103–107 (2017).
Gudipaty, S. A. et al. Mechanical stretch triggers rapid epithelial cell division through Piezo1. Nature 543, 118–121 (2017).
Lancaster, O. M. et al. Mitotic rounding alters cell geometry to ensure efficient bipolar spindle formation. Dev. Cell 25, 270–283 (2013).
Vianay, B. et al. Variation in traction forces during cell cycle progression. Biol. Cell 110, 91–96 (2018).
Park, S. et al. Tissue-scale coordination of cellular behaviour promotes epidermal wound repair in live mice. Nat. Cell Biol. 19, 155–163 (2017).
Aigouy, B. et al. Cell flow reorients the axis of planar polarity in the wing epithelium of Drosophila. Cell 142, 773–786 (2010).
Guillot, C. & Lecuit, T. Mechanics of epithelial tissue homeostasis and morphogenesis. Science 340, 1185–1189 (2013).
Campinho, P. et al. Tension-oriented cell divisions limit anisotropic tissue tension in epithelial spreading during zebrafish epiboly. Nat. Cell Biol. 15, 1405–1414 (2013).
Saxton, R. A. & Sabatini, D. M. mTOR signaling in growth, metabolism, and disease. Cell 169, 361–371 (2017).
Yu, F. X., Zhao, B. & Guan, K. L. Hippo pathway in organ size control, tissue homeostasis, and cancer. Cell 163, 811–828 (2015).
Lloyd, A. C. The regulation of cell size. Cell 154, 1194–1205 (2013).
Folkman, J. & Moscona, A. Role of cell shape in growth control. Nature 273, 345–349 (1978).
Watt, F. M., Jordan, P. W. & O’Neill, C. H. Cell shape controls terminal differentiation of human epidermal keratinocytes. Proc. Natl Acad. Sci. USA 85, 5576–5580 (1988).
Chen, C. S., Mrksich, M., Huang, S., Whitesides, G. M. & Ingber, D. E. Geometric control of cell life and death. Science 276, 1425–1428 (1997).
Roca-Cusachs, P. et al. Micropatterning of single endothelial cell shape reveals a tight coupling between nuclear volume in G1 and proliferation. Biophys. J. 94, 4984–4995 (2008).
Son, S. et al. Direct observation of mammalian cell growth and size regulation. Nat. Methods 9, 910–912 (2012).
Nelson, C. M., Vanduijn, M. M., Inman, J. L., Fletcher, D. A. & Bissell, M. J. Tissue geometry determines sites of mammary branching morphogenesis in organotypic cultures. Science 314, 298–300 (2006).
Poujade, M. et al. Collective migration of an epithelial monolayer in response to a model wound. Proc. Natl Acad. Sci. USA 104, 15988–15993 (2007).
Serra-Picamal, X. et al. Mechanical waves during tissue expansion. Nat. Phys. 8, 628–634 (2012).
Trepat, X. et al. Physical forces during collective cell migration. Nat. Phys. 5, 426–430 (2009).
Tambe, D. T. et al. Collective cell guidance by cooperative intercellular forces. Nat. Mater. 10, 469–475 (2011).
Ginzberg, M. B., Chang, N., Kafri, R. & Kirschner, M. W. Cell size sensing in animal cells coordinates growth rates and cell cycle progression to maintain cell size uniformity. bioRxiv https://dx.doi.org/10.1101/123851 (2017).
Cadart, C. et al. An adder behavior in mammalian cells achieves size control by modulation of growth rate and cell cycle duration. bioRxiv https://dx.doi.org/10.1101/152728 (2017).
Guo, M. et al. Cell volume change through water efflux impacts cell stiffness and stem cell fate. Proc. Natl Acad. Sci. USA 114, E8618–E8627 (2017).
Wagenmakers, E. J. A practical solution to the pervasive problems of P values. Psychon. Bull. Rev. 14, 779–804 (2007).
Schwarz, G. Estimating the dimension of a model. Ann. Stat. 6, 461–464 (1978).
Valon, L., Marin-Llaurado, A., Wyatt, T., Charras, G. & Trepat, X. Optogenetic control of cellular forces and mechanotransduction. Nat. Commun. 8, 14396 (2017).
Reinsch, S. & Karsenti, E. Orientation of spindle axis and distribution of plasma membrane proteins during cell division in polarized MDCKII cells. J. Cell Biol. 126, 1509–1526 (1994).
Higashi, T., Arnold, T. R., Stephenson, R. E., Dinshaw, K. M. & Miller, A. L. Maintenance of the epithelial barrier and remodeling of cell–cell junctions during cytokinesis. Curr. Biol. 26, 1829–1842 (2016).
Burton, K. & Taylor, D. L. Traction forces of cytokinesis measured with optically modified elastic substrata. Nature 385, 450–454 (1997).
Ginzberg, M. B., Kafri, R. & Kirschner, M. Cell biology. On being the right (cell) size. Science 348, 1245075 (2015).
Acknowledgements
We thank N. Castro for technical assistance, A. Elosegui-Artola for assistance with the data analysis and L. Hufnagel for generously providing MDCK-Fucci cells. We are grateful to B. Baum and members of our groups for insightful comments and discussion. This work was supported by the Spanish Ministry of Economy and Competitiveness/FEDER (BES-2013-062633 grant to M.U., BFU2016-79916-P and BFU2014-52586-REDT to P.R.-C., BFU2015-65074-P to X.T., and BFU2016-75101-P and RYC-2014-15559 to V.C.), the Generalitat de Catalunya (2014-SGR-927 to X.T. and CERCA programme), the European Research Council (CoG-616480 to X.T.), European Commission (Grant Agreement SEP-210342844 to P.R.-C. and X.T.), Obra Social “La Caixa”, a Career Integration Grant within the seventh European Community Framework Programme (PCIG10-GA-2011-303848 to P.R.-C.), Fundació la Marató de TV3 (project 20133330 to P.R.-C.). IBEC is recipient of a Severo Ochoa Award of Excellence from the MINECO.
Author information
Authors and Affiliations
Contributions
M.U., X.S.-P. and X.T. conceived the study and designed the experiments. M.U. and S.W. performed the experiments. M.U. and X.S.-P. implemented the data analysis software. M.U. analysed the data. M.S.-P. and R.G. carried out the predictive model analysis. V.C. and P.R.-C. contributed data interpretation. All authors discussed the results. M.U. and X.T. wrote the manuscript with feedback from all authors. X.T. oversaw the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated Supplementary Information
Supplementary Figure 1 Monolayer height evolution during expansion.
(a) Maximum intensity projection image of Fucci cells expressing CIBN-CAAX-GFP as a membrane marker 12 h after removing the PDMS membrane. (b) Map of monolayer height corresponding to (a). Images are representative of n = 4 independent experiments. c-e, Monolayer height (c), monolayer area (d), and monolayer volume (e), at different times after removing the PDMS membrane (n = 4 experiments). (f) Monolayer height profile at different times after removing the PDMS membrane. Data is representative of n = 4 independent experiments. (g) Cell height as a function of cell area for cells at G1 during an arbitrary time point of monolayer expansion (19 h, n = 185 cells from 3 independent experiments). (h) Area of cells in G1 binned in area quintiles (5 groups of 37 cells each). (i) Cell height as a function of tension for each of the n = 185 cells. Each of the groups shown in (h) is labelled using a different color/symbol. Height and tension are weakly correlated for the whole cell population (black dashed line) but correlation is lost within each group (r1 = −0.04, r2 = −0.11, r3 = −0.21, r4 = 0.19, r5 = −0.13, rall = −0.17, Pearson Correlation). Thus, for cells with the same area, tension and height are not correlated. Note, however, that a precise reconstruction of tension using Monolayer Stress Microscopy would require knowledge of the geometry of the stress bearing elements of the cell in 3D. All error bars represent SD.
Supplementary Figure 2 Monolayer mechanics and density in confinement.
(a-c) Traction and tension followed a very similar temporal pattern in expansion (Fig. 1) and confinement (no expansion), characterized by a transient increase followed by a slow decrease and a plateau. As expected, however, cell density followed opposite trends in the two conditions. (a) Time evolution of mean traction in monolayers confined to rectangular patterns. (b) Time evolution of mean tension in monolayers confined to rectangular patterns. (c) Time evolution of cell area in monolayers confined to rectangular patterns. Cells were seeded in rectangular collagen patterns of the same size than those used for expansion studies (Fig. 1). Data recording started 4 h after seeding (as in the expansion case). Data are averages from n = 9 patterns from 2 independent experiments. Shaded areas are SD.
Supplementary Figure 3 Extended correlation matrix between measured properties and cell cycle duration.
a-b, Pearson's linear correlation coefficients between all physical properties and the duration of G1 (a), and S-G2-M (b). (c) Duration of G1, duration of the interval between the beginning of the cycle and the Cdt1 peak, and duration of the interval between the Cdt1 peak and the G1-S transition. Mean±SD are represented. (d) Pearson's linear correlation coefficients between the duration of G1 and the population averaged cell properties over the 0h-Cdt1peak interval. (e) The duration of S-G2-M correlates with that of G1, indicating that S-G2-M has memory of G1. Dashed lines represent linear fits. All graphs include n = 120 cells from 3 independent experiments. Error bars represent SD.
Supplementary Figure 4 Correlations between geometric and mechanical properties.
(a) Cell area at the G1-S transition as a function of cell area at birth. (b) Cell area increment during G1 as a function of cell area at birth. The observed positive correlation between the area added during G1 and the initial cell area rules out an adder behavior in area. (c) Mean cell area growth rate during G1 as a function of mean cell tension during G1. (d) Pearson's linear correlation coefficients between the duration of G1, tG1, and cellular tension 6 h after the beginning of the cycle, σc6h. Correlations are computed for each area growth rate sextile 6 h after the beginning of the cycle. For each sextile correlation is negative, supporting that tension is predictive of G1 duration independently of area growth rate. (e) Area at the G1-S transition as a function of the mean cell area growth rate during G1. (f) Area at the G1-S transition as a function of the mean nuclear area growth rate during G1. (g) Mean cell area during G1 as a function of the mean nuclear area during G1. Dashed lines represent linear fits. All graphs include n = 120 cells from 3 independent experiments.
Supplementary Figure 5 Bayes factors of product models.
(a) Bayes factors of the product models that predict G1 duration. (b) Bayes factors of the product models used to predict S-G2-M duration. All Bayes factors were computed using n = 120 cells from 3 independent experiments.
Supplementary information
Supplementary Information
Supplementary Figures 1–5, Supplementary Table and Supplementary Video legends
Supplementary Table 1
Ranked list of one-property linear models to predict the duration of G1, tG1 (top), or the duration of S–G2–M, tSG2M (bottom).
Supplementary Table 2
Ranked list of two-property product models to predict the duration of G1, tG1.
Supplementary Table 3
Ranked list of two-property product models to predict the duration of S-G2-M, tSG2M.
Supplementary Table 4
Ranked list of two-term product models to predict the duration of G1, tG1.
Supplementary Table 5
Ranked list of two-term product models to predict the duration of S-G2-M, tSG2M.
Supplementary Video 1
Force maps during MDCK-Fucci growth.
Supplementary Video 2
MDCK monolayers expand at constant density.
Supplementary Video 3
Illustration of force average calculation.
Supplementary Video 4
Cytoskeletal dynamics in the dividing cell and its neighbours.
Supplementary Video 5
E-cadherin dynamics in the dividing cell and its neighbours.
Rights and permissions
About this article
Cite this article
Uroz, M., Wistorf, S., Serra-Picamal, X. et al. Regulation of cell cycle progression by cell–cell and cell–matrix forces. Nat Cell Biol 20, 646–654 (2018). https://doi.org/10.1038/s41556-018-0107-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0107-2
This article is cited by
-
“Patchiness” in mechanical stiffness across a tumor as an early-stage marker for malignancy
BMC Ecology and Evolution (2024)
-
Monitoring the mass, eigenfrequency, and quality factor of mammalian cells
Nature Communications (2024)
-
LATS1 controls CTCF chromatin occupancy and hormonal response of 3D-grown breast cancer cells
The EMBO Journal (2024)
-
DNA mechanocapsules for programmable piconewton responsive drug delivery
Nature Communications (2024)
-
Centripetal nuclear shape fluctuations associate with chromatin condensation in early prophase
Communications Biology (2023)