Abstract
Kelch-like protein 6 (KLHL6) is an uncharacterized gene mutated in diffuse large B-cell lymphoma (DLBCL). Here we report that KLHL6 assembles with cullin3 to form a functional cullin–RING ubiquitin ligase. Mutations in KLHL6 inhibit its ligase activity by disrupting the interaction with cullin3. Loss of KLHL6 favours DLBCL growth and survival both in vitro and in xenograft models. We further established that the mRNA decay factor roquin2 is a substrate of KLHL6. Degradation of roquin2 is dependent on B-cell receptor activation, and requires the integrity of the Tyr691 residue in roquin2 that is essential for its interaction with KLHL6. A non-degradable roquin2(Y691F) mutant requires its RNA-binding ability to phenocopy the effect of KLHL6 loss. Stabilization of roquin2 promotes mRNA decay of the tumour suppressor and NF-κB pathway inhibitor, tumour necrosis factor-α-inducible gene 3. Collectively, our findings uncover the tumour suppressing mechanism of KLHL6.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Yang, Y. & Staudt, L. M. Protein ubiquitination in lymphoid malignancies. Immunol. Rev. 263, 240–256 (2015).
Gupta-Rossi, N. et al. Specific over-expression of deltex and a new kelch-like protein in human germinal center B cells. Mol. Immunol. 39, 791–799 (2003).
Kroll, J. et al. The BTB-kelch protein KLHL6 is involved in B-lymphocyte antigen receptor signaling and germinal center formation. Mol. Cell Biol. 25, 8531–8540 (2005).
Morin, R. D. et al. Frequent mutation of histone-modifying genes in non-Hodgkin lymphoma. Nature 476, 298–303 (2011).
Lohr, J. G. et al. Discovery and prioritization of somatic mutations in diffuse large B-cell lymphoma (DLBCL) by whole-exome sequencing. Proc. Natl Acad. Sci. USA 109, 3879–3884 (2012).
García-Ramírez, I. et al. Crebbp loss cooperates with Bcl2 overexpression to promote lymphoma in mice. Blood 129, 2645–2656 (2017).
Reddy, A. et al. Genetic and functional drivers of diffuse large B cell lymphoma. Cell 171, 481–494 (2017).
Alizadeh, A. A. et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature 403, 503–511 (2000).
Rosenwald, A. et al. Molecular diagnosis of primary mediastinal B cell lymphoma identifies a clinically favorable subgroup of diffuse large B cell lymphoma related to Hodgkin lymphoma. J. Exp. Med. 198, 851–862 (2003).
Staudt, L. M. Oncogenic activation of NF-κB. Cold Spring Harb. Perspect. Biol. 2, a000109 (2010).
Young, R. M., Shaffer, A. L. 3rd, Phelan, J. D. & Staudt, L. M. B-cell receptor signaling in diffuse large B-cell lymphoma. Semin Hematol. 52, 77–85 (2015).
Lenz, G. et al. Oncogenic CARD11 mutations in human diffuse large B cell lymphoma. Science 319, 1676–1679 (2008).
Davis, R. E. et al. Chronic active B-cell-receptor signalling in diffuse large B-cell lymphoma. Nature 463, 88–92 (2010).
Compagno, M. et al. Mutations of multiple genes cause deregulation of NF-κB in diffuse large B-cell lymphoma. Nature 459, 717–721 (2009).
Kato, M. et al. Frequent inactivation of A20 in B-cell lymphomas. Nature 459, 712–716 (2009).
Fu, M. & Blackshear, P. J. RNA-binding proteins in immune regulation: a focus on CCCH zinc finger proteins. Nat. Rev. Immunol. 17, 130–143 (2017).
Kataoka, K. et al. Aberrant PD-L1 expression through 3′-UTR disruption in multiple cancers. Nature 534, 402–406 (2016).
Leppek, K. et al. Roquin promotes constitutive mRNA decay via a conserved class of stem-loop recognition motifs. Cell 153, 869–881 (2013).
Murakawa, Y. et al. RC3H1 post-transcriptionally regulates A20 mRNA and modulates the activity of the IKK/NF-κB pathway. Nat. Commun. 6, 7367 (2015).
Glasmacher, E. et al. Roquin binds inducible costimulator mRNA and effectors of mRNA decay to induce microRNA-independent post-transcriptional repression. Nat. Immunol. 11, 725–733 (2010).
Vogel, K. U. et al. Roquin paralogs 1 and 2 redundantly repress the Icos and Ox40 costimulator mRNAs and control follicular helper T cell differentiation. Immunity 38, 655–668 (2013).
Schlundt, A. et al. Structural basis for RNA recognition in roquin-mediated post-transcriptional gene regulation. Nat. Struct. Mol. Biol. 21, 671–678 (2014).
Yu, D. et al. Roquin represses autoimmunity by limiting inducible T-cell co-stimulator messenger RNA. Nature 450, 299–303 (2007).
Puente, X. S. et al. Non-coding recurrent mutations in chronic lymphocytic leukaemia. Nature 526, 519–524 (2015).
Lohr, J. G. et al. Widespread genetic heterogeneity in multiple myeloma: implications for targeted therapy. Cancer Cell 25, 91–101 (2014).
Green, M. R. et al. Transient expression of Bcl6 is sufficient for oncogenic function and induction of mature B-cell lymphoma. Nat. Commun. 5, 3904 (2014).
Lydeard, J. R., Schulman, B. A. & Harper, J. W. Building and remodelling Cullin–RING E3 ubiquitin ligases. EMBO Rep. 14, 1050–1061 (2013).
Lo, S. C., Li, X. C., Henzl, M. T., Beamer, L. J. & Hannink, M. Structure of the Keap1:Nrf2 interface provides mechanistic insight into Nrf2 signaling. Embo J. 25, 3605–3617 (2006).
Meriranta, L. et al. Low expression and somatic mutations of the KLHL6 gene predict poor survival in patients with activated B-cell like diffuse large B-cell lymphoma. Blood 128, 2926 (2016).
Kunder, C. A. et al. KLHL6 is preferentially expressed in germinal center-derived B-cell lymphomas. Am. J. Clin. Pathol. 148, 465–476 (2017).
Satpathy, S. et al. Systems-wide analysis of BCR signalosomes and downstream phosphorylation and ubiquitylation. Mol. Syst. Biol. 11, 810 (2015).
Lenz, G. et al. Aberrant immunoglobulin class switch recombination and switch translocations in activated B cell-like diffuse large B cell lymphoma. J. Exp. Med. 204, 633–643 (2007).
Milhollen, M. A. et al. MLN4924, a NEDD8-activating enzyme inhibitor, is active in diffuse large B-cell lymphoma models: rationale for treatment of NF-κB-dependent lymphoma. Blood 116, 1515–1523 (2010).
Zhao, B. et al. The NF-κB genomic landscape in lymphoblastoid B cells. Cell Rep. 8, 1595–1606 (2014).
Zhou, A., Scoggin, S., Gaynor, R. B. & Williams, N. S. Identification of NF-B-regulated genes induced by TNFα utilizing expression profiling and RNA interference. Oncogene 22, 2054–2064 (2003).
Tian, B., Nowak, D. E., Jamaluddin, M., Wang, S. & Brasier, A. R. Identification of direct genomic targets downstream of the nuclear factor-κB transcription factor mediating tumor necrosis factor signaling. J. Biol. Chem. 280, 17435–17448 (2005).
Mansouri, L. et al. Frequent NFKBIE deletions are associated with poor outcome in primary mediastinal B-cell lymphoma. Blood 128, 2666–2670 (2016).
Boice, M. et al. Loss of the HVEM tumor suppressor in lymphoma and restoration by modified CAR-T cells. Cell 167, 405–418 (2016).
Chu, Y. Y. et al. B cells lacking the tumor suppressor TNFAIP3/A20 display impaired differentiation and hyperactivation and cause inflammation and autoimmunity in aged mice. Blood 117, 2227–2236 (2011).
Leiserson, M. D. M., Reyna, M. A. & Raphael, B. J. A weighted exact test for mutually exclusive mutations in cancer. Bioinformatics 32, i736–i745 (2016).
Fontan, L. et al. MALT1 small molecule inhibitors specifically suppress ABC-DLBCL in vitro and in vivo. Cancer Cell 22, 812–824 (2012).
Puente, X. S. et al. Whole-genome sequencing identifies recurrent mutations in chronic lymphocytic leukaemia. Nature 475, 101–105 (2011).
Pasqualucci, L. et al. Hypermutation of multiple proto-oncogenes in B-cell diffuse large-cell lymphomas. Nature 412, 341–346 (2001).
Honma, K. et al. TNFAIP3/A20 functions as a novel tumor suppressor gene in several subtypes of non-Hodgkin lymphomas. Blood 114, 2467–2475 (2009).
Schmitz, R. et al. TNFAIP3 (A20) is a tumor suppressor gene in Hodgkin lymphoma and primary mediastinal B cell lymphoma. J. Exp. Med. 206, 981–989 (2009).
Busino, L. et al. Fbxw7α- and GSK3-mediated degradation of p100 is a pro-survival mechanism in multiple myeloma. Nat. Cell Biol. 14, 375–385 (2012).
Florens, L. & Washburn, M. P. Proteomic analysis by multidimensional protein identification technology. Methods Mol. Biol. 328, 159–175 (2006).
Washburn, M. P., Wolters, D. & Yates, J. R. III. Large-scale analysis of the yeast proteome by multidimensional protein identification technology. Nat. Biotechnol. 19, 242–247 (2001).
McDonald, W. H., Ohi, R., Miyamoto, D. T., Mitchison, T. J. & Yates, J. R. III. Comparison of three directly coupled HPLC MS/MS strategies for identification of proteins from complex mixtures: single-dimension LC-MS/MS, 2-phase MudPIT, and 3-phase MudPIT. Int. J. Mass Spectrom. 219, 245–251 (2002).
Eng, J. K., McCormack, A. L. & Yates, J. R. III. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, 976–989 (1994).
Tabb, D. L., McDonald, W. H. & Yates, J. R.III. DTASelect and Contrast: tools for assembling and comparing protein identifications from shotgun proteomics. J. Proteome Res. 1, 21–26 (2002).
Florens, L. et al. Analyzing chromatin remodeling complexes using shotgun proteomics and normalized spectral abundance factors. Methods 40, 303–311 (2006).
Paoletti, A. C. et al. Quantitative proteomic analysis of distinct mammalian Mediator complexes using normalized spectral abundance factors. Proc. Natl Acad. Sci. USA 103, 18928–18933 (2006).
Zybailov, B. et al. Statistical analysis of membrane proteome expression changes in Saccharomyces cerevisiae. J. Proteome Res. 5, 2339–2347 (2006).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Monti, S. et al. Integrative analysis reveals an outcome-associated and targetable pattern of p53 and cell cycle deregulation in diffuse large B cell lymphoma. Cancer Cell 22, 359–372 (2012).
Wright, G. et al. A gene expression-based method to diagnose clinically distinct subgroups of diffuse large B cell lymphoma. Proc. Natl Acad. Sci. USA 100, 9991–9996 (2003).
Reich, M. et al. GenePattern 2.0. Nat. Genet. 38, 500–501 (2006).
Lenz, G. et al. Stromal gene signatures in large-B-cell lymphomas. N. Engl. J. Med. 359, 2313–2323 (2008).
Barbie, D. A. et al. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature 462, 108–122 (2009).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Parkhomchuk, D. et al. Transcriptome analysis by strand-specific sequencing of complementary DNA. Nucleic Acids Res. 37, e123 (2009).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Wang, L., Feng, Z., Wang, X., Wang, X. & Zhang, X. DEGseq: an R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics 26, 136–138 (2010).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37, 1–13 (2009).
Acknowledgements
We thank C. Vinuesa for providing roquin cDNAs, A. Thomas-Tikhonenko, L. Pasqualucci and Y. Yang for providing DLBCL cell lines, M. Pagano for providing FBP cDNAs, B. Kim for helping with the 3D Matrigel colony formation assay; R. Saffie, D. Brady, E. Witze and R. Greenberg for critically reading the manuscript. This work was supported in part by grant R00-CA166181-04, R01-CA207513-01 from the National Cancer Institute and Gilead Sciences Research Scholars Program in Hematology/Oncology to L.B. R.B. is supported by an NIH Innovator Award (DP2MH107055), the Searle Scholars Program (15-SSP-102), the March of Dimes Foundation (1-FY-15-344) and the W.W. Smith Charitable Trust (C1404).
Author information
Authors and Affiliations
Contributions
L.B. conceived, directed the project and oversaw the results. J.C. designed and performed most experiments. K.L. helped J.C. with experiments in Figs. 1i,3b,d,4h,6a,7h,j and Supplementary Figs. 3b,4h. K.I. helped J.C. with experiments in Fig. 6b. R.B. helped with the bioinformatics analysis of RNA-seq data. A.S., L.F. and M.P.W. performed the mass spectrometry analysis of the KLHL6 complex purified by L.B. M.R.G. and S.T. helped with Figs. 1a,b and 3a,7g and Supplementary Figs. 1a,b,7e,f. L.B. and J.C. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Copy number and transcriptional analysis of KLHL6 in primary DLBCLs.
a, The DNA copy number of chromosome 3 is shown. DNA copy number data from high-resolution SNP microarray analysis of 609 primary DLBCL tumours were utilized form a previously published study26. The position of KLHL6 is annotated for 21 DLBCL tumours with copy number <1.8. KLHL6 loss is detectable in 3.4% of patients. b, A row-normalized heat map is shown for probe sets corresponding to KLHL6. Gene expression microarray from 249 tumours with matched DNA copy number data were obtained from a previously published study26. The data are annotated for the cell of origin subtype and DNA copy loss of KLHL6 as shown in a. 6% of DLBCL cases with expression ≤1 s.d. below the mean are indicated.
Supplementary Figure 2 KLHL6 WT, but not KLHL6 BTB-domain mutant, promotes roquin2 ubiquitylation and degradation.
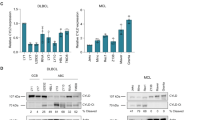
a, Immunoblot analysis from immunoprecipitated endogenous KLHL6 in U2932 cells. IgG antibody immunoprecipitates = negative control. A representative blot from two independent experiments is shown. *Non-specific band. b, Immunoblot analysis from immunoprecipitated Flag–KLHL6 wild-type (WT) and mutants in HEK293T cells. EV, empty vector. c, Quantification of (x axis) and KLHL6 (y axis) protein levels in each DLBCL cell line. n = 11 DLBCL cell lines. r, Pearson correlation coefficient (95% confidence interval). d, Quantification of roquin1 and roquin2 immunoblots. Relative intensities were plotted over time for OCI-LY10 (left panel) and U2932 (right panel) (mean ± s.d., n = 3 independent experiments, two-way ANOVA, *P ≤ 0.05; **P ≤ 0.01; ***P ≤ 0.001; n.s., not significant). e, Immunoblot analysis of whole-cell lysates from HEK293T (TET)-OFF cells transduced with retroviruses encoding a doxycycline (DOX) inducible expression of Flag-tagged KLHL6 carrying a hygromycin cassette (top panel). DOX was added and/or washed at the indicated times. Bottom panel shows the immunoblot analysis of the indicated proteins in HBL1 cells transduced with lentiviruses encoding a doxycycline (DOX) inducible expression of KLHL6 wild-type (WT) carrying a puromycin cassette. The cells were treated with DOX for 12 h. f, Immunoblot analysis of whole-cell lysates from OCI-LY8 cells transduced with retroviruses encoding empty vector (EV), KLHL6 wild-type (WT) or BTB-domain mutants (L65P, S94I and F97L) carrying a puromycin cassette. g, Immunoblot analysis of whole-cell lysates from HEK293T cells transduced with lentiviruses encoding a doxycline (DOX) inducible expression of KLHL6 carrying a puromycin cassette and infected with lentiviruses encoding empty vector(EV) or KLHL6 BTB-domain mutants (L65P and S94I) carrying a GFP marker. The cells were treated with DOX for 12 h. h, Immunoblot analysis of immunoprecipitated endogenous roquin2 in U2932 KLHL6+/+ and KLHL6−/− (clone-derived) cells treated with or without MG132 for 6 h. A representative blot from two independent experiments is shown. Unprocessed original scans of immunoblots for a,b,e–h are shown in Supplementary Fig. 8, and statistical source data for c,d and exact P values for d can be found in Supplementary Table 6. Unless otherwise noted, immunoblots are representative of three independent experiments.
Supplementary Figure 3 Loss of KLHL6 promotes proliferation of ABC-DLBCL cells.
a, Immunoblot analysis of whole-cell lysates from U2932-Cas9 cells infected with lentiviruses encoding scrambled gRNA or gRNAs against KLHL6 exon 1 carrying a puromycin cassette. b, Left panel shows a representative image of U2932 cell colonies expressing indicated gRNAs and plated into a matrigel. After 14 days, the matrigel was dissolved and recovered. Cells were counted and plotted as shown on the right panel (mean ± s.d., n = 4 independent experiments, one-way ANOVA, ****P ≤ 0.0001). Scale bar, 150 μm. c, Immunoblot analysis of whole-cell lysates from U2932 cells infected with lentiviruses encoding the indicated shRNAs carrying a puromycin cassette. d, MTS assay for U2932 cells infected with lentiviruses encoding the indicated shRNAs and grown in media containing 1 μg ml−1 of F(ab′)2–IgM. Values were normalized to the shCTRL cells at time 0 h and set to 100% (mean ± s.d., n = 3 independent experiments, two-way ANOVA; ***P ≤ 0.001, ****P ≤ 0.0001). e, Immunoblot analysis of whole-cell lysates from OCI-LY10 cells infected with lentiviruses encoding indicated shRNAs carrying a puromycin cassette. f, Left panel shows a representative image of OCI-LY10 cell colonies infected with indicated shRNAs and plated in Matrigel. After 14 days, the Matrigel was dissolved and recovered. Cells were counted and plotted as shown on the right panel (mean ± s.d., n = 4 independent experiments, one-way ANOVA, ****P ≤ 0.0001). Scale bar, 150 μm. g, Gating strategy for Fig. 4c. The box indicates the stained cells. Unprocessed original scans of immunoblots for a,c,e are shown in Supplementary Fig. 8, and statistical source data and exact P values for b,d,f can be found in Supplementary Table 6. Unless otherwise noted, immunoblots are representative of three independent experiments.
Supplementary Figure 4 The roquin2(Y691F) mutant is resistant to KLHL6-mediated degradation.
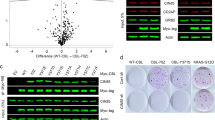
a, Schematic representation of the sequence of the biotinylated roquin2 peptides. b, A pull-down assay using the indicated amount of biotinylated roquin2 peptides incubated with cell extracts from HEK293T cells stably expressing KLHL6. AP, Affinity purification. c,d, Same as in b except that Flag-tagged in vitro translated proteins as indicated were used instead of cell extracts. Immunoblot analysis for the indicated proteins was performed using Flag antibody. e, Quantification of immunoblots from BJAB cells stably expressing roquin2(WT) or roquin2(Y691F). Relative intensities were plotted over time under cycloheximide (CHX) treatment (mean ± s.d., n = 3 independent experiments, two-way ANOVA, *P ≤ 0.05; **P ≤ 0.01). f, Immunoblot analysis of whole-cell lysates from HEK2932T cells stably expressing KLHL6(WT) or KLHL6(L65P) carrying hygromycin cassette and further infected with retroviruses encoding an empty vector (EV), roquin2(WT) or roquin2(Y691F) carrying a puromycin cassette. A low exposure (l.e.) and high exposure (h.e.) are shown. g, Immunoblot analysis for indicated amounts of recombinant roquin2 (set as the standard) along with whole-cell lysates from U2932 stably expressing HA–roquin2(WT) and HA–roquin2(Y691F) (left panel) or U2932 KLHL6+/+ and KLHL6−/− cells (clone-derived) (middle panel). Right panel shows intensity of quantified roquin2 bands compared to the standard. A representative blot from one experiment is shown. h, Flow cytometry analysis of GFP+-live OCI-LY10 KLHL6+/+ and KLHL6−/− (clone-derived) cells infected with lentiviruses encoding the indicated shRNAs carrying a GFP marker. Cells were grown in media containing 2 μg ml−1 of F(ab′)2–IgM and normalized to the shCTRL cells set as 100% (mean ± s.d., n = 3 independent experiments, two-way ANOVA, **P ≤ 0.01; ****P ≤ 0.0001). i, Immunoblot analysis of whole-cell lysates from GFP-sorted U2932 (left panel) and OCI-LY10 (right panel) KLHL6+/+ and KLHL6−/− (clone-derived) infected with lentiviruses encoding the indicated shRNAs carrying a GFP marker. A representative blot from two independent experiments is shown. Unprocessed original scans of immunoblots for b–d,f,g,i are shown in Supplementary Fig. 8, and source data for g and statistical source data and exact P values for e,h can be found in Supplementary Table 6. Unless otherwise noted, immunoblots are representative of three independent experiments.
Supplementary Figure 5 roquin 2 is degraded upon BCR stimulation in a KLHL6-dependent manner.
a, Flow cytometry analysis of a panel of human DLBCLs, as indicated, stained with anti-IgM or anti-IgG antibody to detect surface expression. A darker curve indicates a positive signal. A representative image from two independent experiments is shown. b, MTS assay for U2932 cells KLHL6+/+ and KLHL6−/− treated with increasing amounts of ibrutinib for 48 h. Values were normalized to the non-treated cells and set as 100% (mean ± s.d., n = 3 independent experiments, two-way ANOVA; *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001). c, Immunoblot analysis of whole-cell lysates from U2932 cells electroporated with a siRNA scramble (siCTRL) or siRNA targeting KLHL6 (siKLHL6) and treated with increasing concentrations of F(ab′)2–IgM for 6 h. d, Immunoblot analysis of whole-cell lysates from OCI-LY10 cells electroporated with siRNA scramble (siCTRL) or siRNA targeting KLHL6 (siKLHL6) and treated with 10 μg ml−1 F(ab′)2–IgM for the indicated times. e, Gating strategy for a. The box indicates the stained cells. Unprocessed original scans of immunoblots for c,d are shown in Supplementary Fig. 8, and statistical source data and exact P values for b can be found in Supplementary Table 6. Unless otherwise noted, immunoblots are representative of three independent experiments.
Supplementary Figure 6 Analysis of transcripts deregulated by non-degradable roquin2(Y691F) mutant.
a, Gene ontology (GO) analysis of genes regulated by the non-degradable roquin2(Y691F) mutant. Bar plot for the −log10 of the P value of the top 10 enriched GO terms of genes regulated by roquin2(Y691F) as determined by the hypergeometric distribution. −reg., negative regulation; +reg., positive regulation. Statistical analysis and genes list are provided in Supplementary Table 4. b, Ranking of roquin2-regulated genes by the percentage of genetic alteration in human DLBCLs from the TCGA database along with the base mean expression from the RNA-seq analysis. The cut-off was set at 6%. 'Yes' indicates transcripts that are dependent on the ROQ domain of roquin2; 'No' indicates transcripts that are not dependent. (c) Level of indicated mRNAs analysed by quantitative PCR from U2932 cells stably expressing HA–roquin2(WT), HA–roquin2(Y691F) or HA–roquin2(Y691FΔROQ) and treated with 10 μg ml−1 of F(ab′)2–IgM for 12 h. The value for each PCR product present in HA–roquin2(WT) cells was set as 100%. Rescued transcripts are defined as ones whose levels reach at least 70% of the roquin2(WT) control (indicated by the dashed line) (mean ± s.d., n = 3 independent experiments). Source data for c can be found in Supplementary Table 6.
Supplementary Figure 7 KLHL6 regulates TNFAIP3 levels in a roquin2-dependent manner.
a, Immunoblot analysis of whole-cell lysates from GFP-sorted U2932 KLHL6+/+, KLHL6−/− or KLHL6−/− cells infected with lentiviruses encoding the indicated shRNAs carrying a GFP marker and treated with 10 μg ml−1 of F(ab′)2–IgM for 6 h. b, Immunoblot analysis of whole-cell lysates from GFP-sorted U2932 KLHL6−/− (clone-derived) cells infected with retroviruses encoding empty vector (EV) or KLHL6(WT) carrying a GFP marker and treated with 10 μg ml−1 of F(ab′)2–IgM for 6 h. c, Same as in b except that cells were treated with 10 μg ml−1 F(ab′)2–IgM for the indicated times (min). d, GFP-sorted U2932 KLHL6+/+, KLHL6−/− or KLHL6−/− cells infected with lentiviruses encoding the indicated shRNAs carrying a GFP marker were fractionated into cytoplasmic and nuclear extracts and analysed by immunoblotting for the indicated proteins. e, A heat map showing the presence of biallelic deletion (dark blue), monoallelic deletion (light blue) and monoallelic mutation (green) of TNFAIP3 in DLBCL tumours sequenced at UNMC6 and DCI7 (n = 1,175). Deleterious mutations of KLHL6 BTB-domain are shown in these same cases. Based on DCI database, exclusivity analysis of KLHL6 and TNFAIP3 mutations is provided in Supplementary Table 5 (n = 1,001). f, A heat map showing tumour gene alterations matched with gene expression profiling data available at UNMC. Single sample gene set enrichment analysis (GSEA) was utilized to infer NF-κB activity via expression of target gene sets from the molecular signatures database (NFkB_Q and NFkB_C)26,59. The enrichment score is displayed as a row-normalized heat map. Unprocessed original scans of immunoblots for a–d are shown in Supplementary Fig. 8. Unless otherwise noted, immunoblots are representative of three independent experiments.
Supplementary Figure 8
Unprocessed blots.
Supplementary information
Supplementary Information
Supplementary Figures 1–8 and Supplementary Table legends
Supplementary Table 2
Proteomic analysis of KLHL6 complex
Supplementary Table 3
Transcriptomic analysis of roquin2-regulated genes
Supplementary Table 4
GO enrichments of roquin2-regulated genes
Supplementary Table 5
Mutually exclusive analysis of KLHL6 BTB-mutations in DLBCL
Supplementary Table 6
Statistical Source Data
Rights and permissions
About this article
Cite this article
Choi, J., Lee, K., Ingvarsdottir, K. et al. Loss of KLHL6 promotes diffuse large B-cell lymphoma growth and survival by stabilizing the mRNA decay factor roquin2. Nat Cell Biol 20, 586–596 (2018). https://doi.org/10.1038/s41556-018-0084-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0084-5
This article is cited by
-
Regulation of inflammatory diseases via the control of mRNA decay
Inflammation and Regeneration (2024)
-
Lymphoid clonal hematopoiesis: implications for malignancy, immunity, and treatment
Blood Cancer Journal (2023)
-
Clinical significance of FBXW7 loss of function in human cancers
Molecular Cancer (2022)
-
Disease-associated KBTBD4 mutations in medulloblastoma elicit neomorphic ubiquitylation activity to promote CoREST degradation
Cell Death & Differentiation (2022)
-
Disorders of ubiquitylation: unchained inflammation
Nature Reviews Rheumatology (2022)