Abstract
The list of long non-coding RNAs (lncRNAs) involved in the p53 pathway of the DNA damage response is rapidly expanding, but whether lncRNAs have a role in maintaining the de novo structure of DNA is unknown. Here, we demonstrate that the p53-responsive lncRNA GUARDIN is important for maintaining genomic integrity under steady-state conditions and after exposure to exogenous genotoxic stress. GUARDIN is necessary for preventing chromosome end-to-end fusion through maintaining the expression of telomeric repeat-binding factor 2 (TRF2) by sequestering microRNA-23a. Moreover, GUARDIN also sustains breast cancer 1 (BRCA1) stability by acting as an RNA scaffold to facilitate the heterodimerization of BRCA1 and BRCA1-associated RING domain protein 1 (BARD1). As such, GUARDIN silencing triggered apoptosis and senescence, enhanced cytotoxicity of additional genotoxic stress and inhibited cancer xenograft growth. Thus, GUARDIN may constitute a target for cancer treatment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Hoeijmakers, J. H. J. DNA damage, aging, and cancer. N. Engl. J. Med. 361, 1475–1485 (2009).
Hoeijmakers, J. H. J. The key role of DNA damage on cancer, aging and longevity. Environ. Mol. Mutagen. 53, S13 (2012).
Braig, M. & Schmitt, C. A. Oncogene-induced senescence: putting the brakes on tumor development. Cancer Res. 66, 2881–2884 (2006).
Best, B. P. Nuclear DNA damage as a direct cause of aging. Rejuvenation Res. 12, 199–208 (2009).
Lahtz, C. & Pfeifer, G. P. Epigenetic changes of DNA repair genes in cancer. J. Mol. Cell Biol. 3, 51–58 (2011).
Xin, H. W., Liu, D. & Zhou, S. Y. The telosome/shelterin complex and its functions. Genome Biol. 9, 232 (2008).
Palm, W. & de Lange, T. How shelterin protects mammalian telomeres. Annu. Rev. Genet. 42, 301–334 (2008).
Liu, D., O’Connor, M. S., Qin, J. & Songyang, Z. Telosome, a mammalian telomere-associated complex formed by multiple telomeric proteins. J. Biol. Chem. 279, 51338–51342 (2004).
Fumagalli, M. et al. Telomeric DNA damage is irreparable and causes persistent DNA-damage-response activation. Nat. Cell Biol. 14, 355–365 (2012).
van Tuyn, J. & Adams, P. D. Signalling the end of the line. Nat. Cell Biol. 14, 339–341 (2012).
Martinez, P. & Blasco, M. A. Role of shelterin in cancer and aging. Aging Cell 9, 653–666 (2010).
Majidinia, M. & Yousefi, B. DNA damage response regulation by microRNAs as a therapeutic target in cancer. DNA Repair 47, 1–11 (2016).
Idogawa, M. et al. Identification and analysis of large intergenic non-coding RNAs regulated by p53 family members through a genome-wide analysis of p53-binding sites. Hum. Mol. Genet. 23, 2847–2857 (2014).
Huarte, M. et al. A large intergenic noncoding RNA Induced by p53 mediates global gene repression in the p53 response. Cell 142, 409–419 (2010).
Schmitt, A. M. et al. An inducible long noncoding RNA amplifies DNA damage signaling. Nat. Genet. 48, 1370–1376 (2016).
Di Micco, R. et al. Oncogene-induced senescence is a DNA damage response triggered by DNA hyper-replication. Nature 444, 638–642 (2006).
Léveillé, N. et al. Genome-wide profiling of p53-regulated enhancer RNAs uncovers a subset of enhancers controlled by a lncRNA. Nat. Commun. 27, 6520 (2015).
Wang, W., Cheng, B., Miao, L., Mei, Y. & Wu, M. Mutant p53-R273H gains new function in sustained activation of EGFR signaling via suppressing miR-27a expression. Cell Death Dis. 4, e574 (2013).
Tsang, W. P., Ho, F. Y., Fung, K. P., Kong, S. K. & Kwok, T. T. p53-R175H mutant gains new function in regulation of doxorubicin-induced apoptosis. Int. J. Cancer 114, 331–336 (2005).
Liu, D. P., Song, H. & Xu, Y. A common gain of function of p53 cancer mutants in inducing genetic instability. Oncogene 29, 949–956 (2010).
Jiang, P. et al. p53 regulates biosynthesis through direct inactivation of glucose-6-phosphate dehydrogenase. Nat. Cell Biol. 13, 310–316 (2011).
Georgakilas, A. G. et al. Are common fragile sites merely structural domains or highly organized “functional” units susceptible to oncogenic stress? Cell. Mol. Life Sci. 71, 4519–4544 (2014).
Henrich, K. O., Schwab, M. & Westermann, F. 1p36 tumor suppression—a matter of dosage? Cancer Res. 72, 6079–6088 (2012).
Hunten, S. et al. p53-regulated networks of protein, mRNA, miRNA, and lncRNA expression revealed by integrated pulsed stable isotope labeling with amino acids in cell culture (pSILAC) and next generation sequencing (NGS) analyses. Mol. Cell. Proteom. 14, 2609–2629 (2015).
Ashouri, A. et al. Pan-cancer transcriptomic analysis associates long non-coding RNAs with key mutational driver events. Nat. Commun. 7, 13197 (2016).
Sarkar, S. et al. Different combinations of genetic/epigenetic alterations inactivate the p53 and pRb pathways in invasive human bladder cancers. Cancer Res. 60, 3862–3871 (2000).
Vikhanskaya, F., Lee, M. K., Mazzoletti, M., Broggini, M. & Sabapathy, K. Cancer-derived p53 mutants suppress p53-target gene expression—potential mechanism for gain of function of mutant p53. Nucleic Acids Res. 35, 2093–2104 (2007).
Li, Q. et al. C23 promotes tumorigenesis via suppressing p53 activity. Oncotarget. 7, 58274–58285 (2016).
Debacq-Chainiaux, F., Erusalimsky, J. D., Campisi, J. & Toussaint, O. Protocols to detect senescence-associated β-galactosidase (SA-β-gal) activity, a biomarker of senescent cells in culture and in vivo. Nat. Protoc. 4, 1798–1806 (2009).
Dimri, G. P. et al. A biomarker that identifies senescent human-cells in culture and in aging skin in vivo. Proc. Natl Acad. Sci. USA 92, 9363–9367 (1995).
Sharma, V. et al. A BRCA1-interacting lncRNA regulates homologous recombination. EMBO Rep. 16, 1520–1534 (2015).
Kartha, R. V & Subramanian, S. Competing endogenous RNAs (ceRNAs): new entrants to the intricacies of gene regulation. Front. Genet. 5, 8 (2014).
Tay, Y., Rinn, J. & Pandolfi, P. P. The multilayered complexity of ceRNA crosstalk and competition. Nature 505, 344–352 (2014).
Dodd, D. W., Gagnon, K. T. & Corey, D. R. Digital quantitation of potential therapeutic target RNAs. Nucleic Acid. Ther. 23, 188–194 (2013).
Luo, Z. H. et al. mir-23a induces telomere dysfunction and cellular senescence by inhibiting TRF2 expression. Aging Cell 14, 391–399 (2015).
Choi, K. H., Farrell, A. S., Lakamp, A. S. & Ouellette, M. M. Characterization of the DNA binding specificity of shelterin complexes. Nucleic Acids Res. 39, 9206–9223 (2011).
Yan, Y., Liang, Z., Du, Q., Yang, M. & Geller, D. A. MicroRNA-23a downregulates the expression of interferon regulatory factor-1 in hepatocellular carcinoma cells. Oncol. Rep. 36, 633–640 (2016).
Baer, R. & Ludwig, T. The BRCA1/BARD1 heterodimer, a tumor suppressor complex with ubiquitin E3 ligase activity. Curr. Opin. Genet. Dev. 12, 86–91 (2002).
Hashizume, R. et al. The RING heterodimer BRCA1–BARD1 is a ubiquitin ligase inactivated by a breast cancer-derived mutation. J. Biol. Chem. 276, 14537–14540 (2001).
Brzovic, P. S., Rajagopal, P., Hoyt, D. W., King, M. C. & Klevit, R. E. Structure of a BRCA1–BARD1 heterodimeric RING–RING complex. Nat. Struct. Biol. 8, 833–837 (2001).
Shrivastav, M., De Haro, L. P. & Nickoloff, J. A. Regulation of DNA double-strand break repair pathway choice. Cell Res. 18, 134–147 (2008).
Vogelstein, B., Lane, D. & Levine, A. J. Surfing the p53 network. Nature 408, 307–310 (2000).
Galanos, P. et al. Chronic p53-independent p21 expression causes genomic instability by deregulating replication licensing. Nat. Cell Biol. 18, 777–789 (2016).
Yin, D. et al. miR-34a functions as a tumor suppressor modulating EGFR in glioblastoma multiforme. Oncogene 32, 1155–1163 (2013).
Okada, N. et al. A positive feedback between p53 and miR-34 miRNAs mediates tumor suppression. Genes Dev. 28, 438–450 (2014).
Wynford-Thomas, D. p53: guardian of cellular senescence. J. Pathol. 180, 118–121 (1996).
Stiewe, T. The p53 family in differentiation and tumorigenesis. Nat. Rev. Cancer 7, 165–168 (2007).
Karlseder, J., Broccoli, D., Dai, Y. M., Hardy, S. & de Lange, T. p53- and ATM-dependent apoptosis induced by telomeres lacking TRF2. Science 283, 1321–1325 (1999).
Takai, H., Smogorzewska, A. & de Lange, T. DNA damage foci at dysfunctional telomeres. Curr. Biol. 13, 1549–1556 (2003).
Wang, Y. et al. BASC, a super complex of BRCA1-associated proteins involved in the recognition and repair of aberrant DNA structures. Genes Dev. 14, 927–939 (2000).
Friedenson, B. The BRCA1/2 pathway prevents hematologic cancers in addition to breast and ovarian cancers. BMC Cancer 7, 152 (2007).
Tapia, T. et al. Promoter hypermethylation of BRCA1 correlates with absence of expression in hereditary breast cancer tumors. Epigenetics 3, 157–163 (2008).
Shen, J., Ambrosone, C. B. & Zhao, H. Novel genetic variants in microRNA genes and familial breast cancer. Int. J. Cancer 124, 1178–1182 (2009).
Beck, M. et al. The quantitative proteome of a human cell line. Mol. Syst. Biol. 7, 549 (2011).
Badie, S. et al. BRCA1 and CtIP promote alternative non-homologous end-joining at uncapped telomeres. EMBO J. 34, 410–424 (2015).
Isono, M. et al. BRCA1 directs the repair pathway to homologous recombination by promoting 53BP1 dephosphorylation. Cell Rep. 18, 520–532 (2017).
Powell, S. N. & Kachnic, L. A. Roles of BRCA1 and BRCA2 in homologous recombination, DNA replication fidelity and the cellular response to ionizing radiation. Oncogene 22, 5784–5791 (2003).
Feng, Z. & Zhang, J. A dual role of BRCA1 in two distinct homologous recombination mediated repair in response to replication arrest. Nucleic Acids Res. 40, 726–738 (2012).
Bau, D. T., Mau, Y. C. & Shen, C. Y. The role of BRCA1 in non-homologous end-joining. Cancer Lett. 240, 1–8 (2006).
Jiang, G. et al. BRCA1–Ku80 protein interaction enhances end-joining fidelity of chromosomal double-strand breaks in the G1 phase of the cell cycle. J. Biol. Chem. 288, 8966–8976 (2013).
Brown, J. S., O’Carrigan, B., Jackson, S. P. & Yap, T. A. Targeting DNA repair in cancer: beyond PARP inhibitors. Cancer Discov. 7, 20–37 (2017).
Audeh, M. W. et al. Oral poly(ADP-ribose) polymerase inhibitor olaparib in patients with BRCA1 or BRCA2 mutations and recurrent ovarian cancer: a proof-of-concept trial. Lancet 376, 245–251 (2010).
Tutt, A. et al. Oral poly(ADP-ribose) polymerase inhibitor olaparib in patients with BRCA1 or BRCA2 mutations and advanced breast cancer: a proof-of-concept trial. Lancet 376, 235–244 (2010).
Dong, L. et al. Ets-1 mediates upregulation of Mcl-1 downstream of XBP-1 in human melanoma cells upon ER stress. Oncogene 30, 3716–3726 (2011).
Dodd, D. W., Gagnon, K. T. & Corey, D. R. Digital quantitation of potential therapeutic target RNAs. Nucleic Acid. Ther. 23, 188–194 (2013).
Karbaschi, M. & Cooke, M. S. Novel method for the high-throughput processing of slides for the comet assay. Sci. Rep. 4, 7200 (2014).
Liu, X. Y. et al. RIP1 kinase is an oncogenic driver in melanoma. Cancer Res. 75, 1736–1748 (2015).
Jin, L. et al. MicroRNA-149*, a p53-responsive microRNA, functions as an oncogenic regulator in human melanoma. Proc. Natl Acad. Sci. USA 108, 15840–15845 (2011).
Guzman, C., Bagga, M., Kaur, A., Westermarck, J. & Abankwa, D. ColonyArea: an ImageJ plugin to automatically quantify colony formation in clonogenic assays. PLoS ONE 9, e92444 (2014).
Acknowledgements
We thank X. Xu of Shenzhen University for providing U2OS-HR and U2OS-NHEJ cells and T. Ohta of St. Marianna University for providing the HA-BARD1 plasmid. We thank L. Kong and Q. Cheng of Henan Provincial People’s Hospital for their assistance with preparation of tissue sections and immunohistochemistry experiments. This work was supported by grants from the National Key R&D Program of China (2016YFC1302302) and the National Natural Science Foundation of China (81430065, 31371388, 31601117 and 81471551).
Author information
Authors and Affiliations
Contributions
W.L.H., L.J., A.X., X.D.Z. and M.W. designed the research. W.L.H., L.J. and A.X. performed most of the experiments and data analysis. Y.F.W. participated in the experiments and data analysis. R.F.T. participated in the data analysis and manuscript preparation. M.W., R.F.T. and X.D.Z. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 lncRNA#6, GUARDIN, is regulated by wildtype p53 and its depletion induces apoptosis and reduced cell viability.
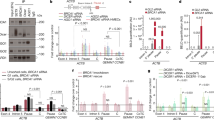
(a) p53-upregulated lncRNAs, 1, 2, 6, 7 and 8 identified by lncRNA profiling along with lincRNA-p21 and DINO were validated after induction of p53 in H1299 cells carrying an inducible wild-type p53 expression system using qPCR. Induced expression of p53 is shown as inset. Mean ± s.e.m.; n=3 independent experiments, two-tailed Student’s t-test. (b) Silencing of lncRNA#6 but not 1, 2, 7 or 8 triggered activation of caspase-3 and cleavage of PARP in HCT116 cells. Data shown represent three independent experiments. (c) Silencing of lncRNA#6 but not 1, 2, 7 or 8 caused reduction in cell viability in HCT116 cells. Mean ± s.e.m.; n=3 independent experiments, two-tailed Student’s t-test. (d) shRNA knockdown efficacy of lncRNA#1, 2, 6, 7 and 8 in HCT116 cells. Mean ± s.e.m.; n=3 independent experiments, two-tailed Student’s t-test. (e) The longest isoform of GUARDIN, RP3-510D11.2-1, was markedly more abundant than the others (RP3-510D11.2-2 and RP3-510D11.2-3) in H1299 cells with or without induction of p53. Mean ± s.e.m.; n=3 independent experiments, two-tailed Student’s t-test. (f) Ectopic expression wild-type p53 but not p53 mutants, p53R175H and p53R273H caused upregulation of GUARDIN in H1299 cells. Mean ± s.e.m.; n=3 independent experiments, two-tailed Student’s t-test. Statistics source data for a and c-f are provided in Supplementary Table 7. Uncropped images of blots for a, b and f in Supplementary Fig. 7.
Supplementary Figure 2 GUARDIN expression in colon cancer cells and tissues and the relation between miR-34a and MIR34AHG expression, copy number loss and p53 status.
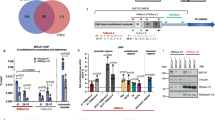
(a) Schematic illustration of the genomic location of GUARDIN at chromosome 1p36.22 and its relation to the genes encoding miR-34a, MIR34AHG, and H6PD. (b) GUARDIN expression was positively correlated with the expression of miR-34a and MIR34AHG in laser capture micro-dissected (LCM) cancer cells from fresh surgical samples of 40 colon cancers. Linear regression analysis. (c) GUARDIN expression along with the expression of miR-34a and MIR34AHG were reduced in LCM-dissected colon cancer cells with copy number loss of their genes. n=40 biologically independent samples, two-tailed Student’s t-test. (d) Silencing or overexpression of GUARDIN did not alter the expression of miR-34a and its target Snail as well as MIR34AHG. n=3 independent experiments, two-tailed Student’s t-test. (e) Introduction of anti-miR-23a or knockdown of MIR34AHG did not affect expression of GUARDIN. n=3 independent experiments, two-tailed Student’s t-test. (f) Schematic illustrations of the putative p53-binding region (p53-BR) located 305-329 bp upstream of the GUARDIN transcriptional start site and the pGL3-basic based GUARDIN promoter reporter constructs. (g) In situ hybridization analysis of GUARDIN expression in HCT116 cells and HCT116 cells with GUARDIN knocked down by shRNA that were used as positive and negative biological controls, respectively. Data shown represent three independent experiments. (h) GUARDIN expression in LCM-dissected colon cancer cells from tumours without copy number loss of its gene, similar to the expression of p21 mRNA, was lower in mutant TP53 cases compared with paired LCM pre-neoplastic epithelial cells (hyperplastic polyps (n=5 biologically independent samples) and adenomas (n=7 biologically independent samples)), whereas there was no significant difference in GUARDIN and p21 mRNA expression between LCM-dissected pre-neoplastic and paired normal colon epithelial cells. n=12 biologically independent samples. two-tailed Student’s t-test. Statistics source data for b-e and h are provided in Supplementary Table 7.
Supplementary Figure 3 Rescue of the inhibitory effects of GUARDIN depletion by pan-caspase inhibitor z-VAD-fmk and the identification of complementarity between GUARDIN and miR-23a.
(a) Knockdown of GUARDIN in HCT116 cells using independent shRNAs. Mean ± s.e.m.; n=3 independent experiments, two-tailed Student’s t-test. (b) Treatment with z-VAD-fmk abolished activation of caspase-3 and cleavage of PARP caused by silencing of GUARDIN in HCT116 cells. Data shown represent three independent experiments. (c) Treatment with z-VAD-fmk partially reversed inhibition of cell number expansion caused by silencing of GUARDIN in HCT116, U2OS, and A549 cells. Mean ± s.e.m.; n=3 independent experiments, two-tailed Student’s t-test. (d) Schematic Illustration of base pairing between miR-23a and GUARDIN. (e) Schematic illustration of psiCHECK2-based luciferase reporter constructs containing wild-type GUARDIN (psiCHECK2-GUARDIN-WT) and a mutant reporter construct in which three putative miR-23a binding sites were mutated (psiCHECK2-GUARDIN-mt). Mutated bases are depicted in red. Statistics source data for a and c are provided in Supplementary Table 7. Uncropped images of blots for b are shown in Supplementary Fig. 7.
Supplementary Figure 4 EMSA assays confirm the GUARDIN-BRCA1-BARD1 interaction with domain mapping experiments identifying distinct regions of GUARDIN which bind to BRCA1 and BARD1.
(a) GUARDIN physically associates with BRCA1 and BARD1 in electrophoretic mobility-shift assays. Data shown represent three independent experiments. (b) Schematic illustration of division of GUARDIN into three fragments corresponding to individual exons of the GUARDIN gene (E1, E2 and E3) along with corresponding deletion mutants used. (c) The fragment of GUARDIN corresponding to exon 3 of its gene was required for its association with BARD1. Data shown represent three independent experiments. (d) The fragments of GUARDIN corresponding to exon 1 and 2 of its gene were necessary for its association with BRCA1. Data shown represent three independent experiments. Uncropped images of blots for a, c and d are shown in Supplementary Fig. 7.
Supplementary Figure 5 GUARDIN binding to BRCA1 and BARD1 occurs through their respective amino-terminal domains but independently of embodied RING domains.
(a) Schematic illustration of BRCA1 and the corresponding mutants with deletion of individual fragments. FL: Full length; NT: N Terminus; SCD: serine cluster domain; CT: C Terminus; ΔRING: RING domain deletion. (b) Schematic illustration of BARD1 and the corresponding mutants with deletion of individual fragments. FL: Full length; NT: N Terminus; FAR: Four ankyrin repeats; CT: C Terminus; ΔRING: RING domain deletion. (c) RNA immunoprecipitation assays were performed in HCT116 cells transfected with indicated BRCA1 segment cDNA. The NT of BRCA1 was required for its association with endogenous GUARDIN. Data shown represent three independent experiments. (d) RNA immunoprecipitation assays were performed in HCT116 cells transfected with the indicated BARD1 construct. The NT of BARD1 was required for its association with endogenous GUARDIN. Data shown represent three independent experiments. (e) RNA immunoprecipitation assays were performed in HCT116 cells transfected with indicated BRCA1 segment cDNA. The RING domain encompassed within the NT of BRCA1 was not required for its association with endogenous GUARDIN. Data shown represent three independent experiments. (f) RNA immunoprecipitation assays were performed in HCT116 cells transfected with indicated BARD1 segment cDNA. The RING domain encompassed within the NT of BARD1 was not required for its association with endogenous GUARDIN. Data shown represent three independent experiments. Uncropped images of blots for c-f are shown in Supplementary Fig. 7.
Supplementary Figure 6 GUARDIN depletion induces cell cycle arrest and defective DNA repair through effects on BRCA1, TRF2 and/or miR-23a.
(a) Silencing of GUARDIN caused cell cycle arrest in the G0/G1 phase in HCT116 cells. Mean ± s.e.m.; n=3 independent experiments, two-tailed Student’s t-test. (b) Treatment with z-VAD-fmk did not affect reduction in activation of the HR and NHEJ repair pathways caused by silencing of GUARDIN. Mean ± s.e.m.; n=3 independent experiments, two-tailed Student’s t-test. (c) Neither overexpression of TRF2, introduction of anti-miR-23a, nor overexpression of BRCA1 alone significantly affected inhibition of DDR activation caused by GUARDIN knockdown, whereas co-overexpression of TRF2 and BRCA1 or co-introduction of anti-miR-23a and BRCA1-expressing constructs abolished reduction in DDR activation in cells with GUARDIN knocked down. Mean ± s.e.m.; n=3 independent experiments, two-tailed Student’s t-test. (d) Overexpression of TRF2, introduction of anti-miR-23a, or overexpression of BRCA1 alone did not rescue HCT116 cells from GUARDIN knockdown, whereas the combination of TRF2 overexpression or anti-miR-23a and BRCA1 overexpression abolished inhibition of cell viability caused by GUARDIN knockdown. Mean ± s.e.m.; n=3 independent experiments, two-tailed Student’s t-test. (e) Model depicting the GUARDIN-mediated pathway required to maintain genomic stability. Statistics source data for a-d are provided in Supplementary Table 7.
Supplementary information
Supplementary Information
Supplementary Figures 1–7 and legends for Supplementary Tables 1–7
Rights and permissions
About this article
Cite this article
Hu, W.L., Jin, L., Xu, A. et al. GUARDIN is a p53-responsive long non-coding RNA that is essential for genomic stability. Nat Cell Biol 20, 492–502 (2018). https://doi.org/10.1038/s41556-018-0066-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0066-7
This article is cited by
-
KLF15 transcriptionally activates LINC00689 to inhibit colorectal cancer development
Communications Biology (2024)
-
Genome instability-related LINC02577, LINC01133 and AC107464.2 are lncRNA prognostic markers correlated with immune microenvironment in pancreatic adenocarcinoma
BMC Cancer (2023)
-
p53-regulated lncRNAs in cancers: from proliferation and metastasis to therapy
Cancer Gene Therapy (2023)
-
Tumor suppressor p53 mediates interleukin-6 expression to enable cancer cell evasion of genotoxic stress
Cell Death Discovery (2023)
-
Long noncoding RNAs as versatile molecular regulators of cellular stress response and homeostasis
Human Genetics (2023)