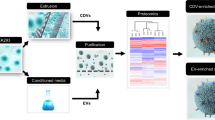
Abstract
The heterogeneity of exosomal populations has hindered our understanding of their biogenesis, molecular composition, biodistribution and functions. By employing asymmetric flow field-flow fractionation (AF4), we identified two exosome subpopulations (large exosome vesicles, Exo-L, 90–120 nm; small exosome vesicles, Exo-S, 60–80 nm) and discovered an abundant population of non-membranous nanoparticles termed ‘exomeres’ (~35 nm). Exomere proteomic profiling revealed an enrichment in metabolic enzymes and hypoxia, microtubule and coagulation proteins as well as specific pathways, such as glycolysis and mTOR signalling. Exo-S and Exo-L contained proteins involved in endosomal function and secretion pathways, and mitotic spindle and IL-2/STAT5 signalling pathways, respectively. Exo-S, Exo-L and exomeres each had unique N-glycosylation, protein, lipid, DNA and RNA profiles and biophysical properties. These three nanoparticle subsets demonstrated diverse organ biodistribution patterns, suggesting distinct biological functions. This study demonstrates that AF4 can serve as an improved analytical tool for isolating extracellular vesicles and addressing the complexities of heterogeneous nanoparticle subpopulations.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Thery, C., Zitvogel, L. & Amigorena, S. Exosomes: composition, biogenesis and function. Nat. Rev. Immunol. 2, 569–579 (2002).
El Andaloussi, S., Mager, I., Breakefield, X. O. & Wood, M. J. Extracellular vesicles: biology and emerging therapeutic opportunities. Nat. Rev. Drug. Discov. 12, 347–357 (2013).
Raposo, G. & Stoorvogel, W. Extracellular vesicles: exosomes, microvesicles, and friends. J. Cell. Biol. 200, 373–383 (2013).
Balaj, L. et al. Tumour microvesicles contain retrotransposon elements and amplified oncogene sequences. Nat. Commun. 2, 180 (2011).
Choi, D. S., Kim, D. K., Kim, Y. K. & Gho, Y. S. Proteomics, transcriptomics and lipidomics of exosomes and ectosomes. Proteomics 13, 1554–1571 (2013).
Thakur, B. K. et al. Double-stranded DNA in exosomes: a novel biomarker in cancer detection. Cell. Res. 24, 766–769 (2014).
Tetta, C., Ghigo, E., Silengo, L., Deregibus, M. C. & Camussi, G. Extracellular vesicles as an emerging mechanism of cell-to-cell communication. Endocrine 44, 11–19 (2013).
Fraunhofer, W. & Winter, G. The use of asymmetrical flow field-flow fractionation in pharmaceutics and biopharmaceutics. Eur. J. Pharm. Biopharm. 58, 369–383 (2004).
Yohannes, G., Jussila, M., Hartonen, K. & Riekkola, M. L. Asymmetrical flow field-flow fractionation technique for separation and characterization of biopolymers and bioparticles. J. Chromatogr. A 1218, 4104–4116 (2011).
Oh, S. et al. Miniaturized asymmetrical flow field-flow fractionation: application to biological vesicles. J. Sep. Sci. 30, 1082–1087 (2007).
Sitar, S. et al. Size characterization and quantification of exosomes by asymmetrical-flow field-flow fractionation. Anal. Chem. 87, 9225–9233 (2015).
Petersen, K. E. et al. A review of exosome separation techniques and characterization of B16-F10 mouse melanoma exosomes with AF4-UV-MALS-DLS-TEM. Anal. Bioanal. Chem. 406, 7855–7866 (2014).
Ashby, J. et al. Distribution profiling of circulating microRNAs in serum. Anal. Chem. 86, 9343–9349 (2014).
Agarwal, K. et al. Analysis of exosome release as a cellular response to MAPK pathway inhibition. Langmuir 31, 5440–5448 (2015).
Beningo, K. A. & Wang, Y. L. Fc-receptor-mediated phagocytosis is regulated by mechanical properties of the target. J. Cell. Sci. 115, 849–856 (2002).
Key, J. et al. Soft discoidal polymeric nanoconstructs resist macrophage uptake and enhance vascular targeting in tumors. ACS Nano 9, 11628–11641 (2015).
Colombo, M., Raposo, G. & Thery, C. Biogenesis, secretion, and intercellular interactions of exosomes and other extracellular vesicles. Annu. Rev. Cell. Dev. Biol. 30, 255–289 (2014).
Hessvik, N. P. & Llorente, A. Current knowledge on exosome biogenesis and release. Cell. Mol. Life Sci. 75, 193–208 (2017).
Molinari, M. & Helenius, A. Chaperone selection during glycoprotein translocation into the endoplasmic reticulum. Science 288, 331–333 (2000).
Fukuda, M. N., Masri, K. A., Dell, A., Luzzatto, L. & Moremen, K. W. Incomplete synthesis of N-glycans in congenital dyserythropoietic anemia type II caused by a defect in the gene encoding α-mannosidase II. Proc. Natl Acad. Sci. USA 87, 7443–7447 (1990).
Yang, W. H. et al. An intrinsic mechanism of secreted protein aging and turnover. Proc. Natl Acad. Sci. USA 112, 13657–13662 (2015).
Martiniuk, F., Ellenbogen, A. & Hirschhorn, R. Identity of neutral alpha-glucosidase AB and the glycoprotein processing enzyme glucosidase II. Biochemical and genetic studies. J. Biol. Chem. 260, 1238–1242 (1985).
Kowal, J. et al. Proteomic comparison defines novel markers to characterize heterogeneous populations of extracellular vesicle subtypes. Proc. Natl Acad. Sci. USA 113, E968–977 (2016).
Pinho, S. S. & Reis, C. A. Glycosylation in cancer: mechanisms and clinical implications. Nat. Rev. Cancer 15, 540–555 (2015).
Escrevente, C. et al. Sialoglycoproteins and N-glycans from secreted exosomes of ovarian carcinoma cells. PLoS ONE 8, e78631 (2013).
Batista, B. S., Eng, W. S., Pilobello, K. T., Hendricks-Munoz, K. D. & Mahal, L. K. Identification of a conserved glycan signature for microvesicles. J. Proteom. Res. 10, 4624–4633 (2011).
Saraswat, M. et al. N-linked (N-) glycoproteomics of urinary exosomes. Mol. Cell. Proteom. 14, 263–276 (2015).
Thery, C., Amigorena, S., Raposo, G. & Clayton, A. Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Curr. Protoc. Cell Biol. 3, 22.1–22.29 (2006).
Merchant, M. L. et al. Microfiltration isolation of human urinary exosomes for characterization by MS. Proteom. Clin. Appl. 4, 84–96 (2010).
Lasser, C., Eldh, M. & Lotvall, J. Isolation and characterization of RNA-containing exosomes. J. Vis. Exp. 59, e3037 (2012).
Chen, C. et al. Microfluidic isolation and transcriptome analysis of serum microvesicles. Lab. Chip 10, 505–511 (2010).
Jorgensen, M. et al. Extracellular vesicle (EV) array: microarray capturing of exosomes and other extracellular vesicles for multiplexed phenotyping. J. Extracell. Vesicles 2, 20920 (2013).
Tauro, B. J. et al. Comparison of ultracentrifugation, density gradient separation, and immunoaffinity capture methods for isolating human colon cancer cell line LIM1863-derived exosomes. Methods 56, 293–304 (2012).
Gardiner, C. et al. Extracellular vesicles, tissue factor, cancer and thrombosis—discussion themes of the ISEV 2014 Educational Day. J. Extracell. Vesicles 4, 26901 (2015).
Liang, Y. et al. Complex N-linked glycans serve as a determinant for exosome/microvesicle cargo recruitment. J. Biol. Chem. 289, 32526–32537 (2014).
White, M. J., Roife, D. & Gomer, R. H. Galectin-3 binding protein secreted by breast cancer cells inhibits monocyte-derived fibrocyte differentiation. J. Immunol. 195, 1858–1867 (2015).
Laubli, H. et al. Lectin galactoside-binding soluble 3 binding protein (LGALS3BP) is a tumor-associated immunomodulatory ligand for CD33-related Siglecs. J. Biol. Chem. 289, 33481–33491 (2014).
Hellstern, S. et al. Functional studies on recombinant domains of Mac-2-binding protein. J. Biol. Chem. 277, 15690–15696 (2002).
Sasaki, T., Brakebusch, C., Engel, J. & Timpl, R. Mac-2 binding protein is a cell-adhesive protein of the extracellular matrix which self-assembles into ring-like structures and binds beta1 integrins, collagens and fibronectin. EMBO J. 17, 1606–1613 (1998).
van Meer, G., Voelker, D. R. & Feigenson, G. W. Membrane lipids: where they are and how they behave. Nat. Rev. Mol. Cell Biol. 9, 112–124 (2008).
Peinado, H. et al. Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nat. Med. 18, 883–891 (2012).
Zhang, H. & Lyden, D. A protocol for asymmetric-flow field-flow fractionation (AF4) of small extracellular vesicles. Protocol Exchange https://doi.org/10.1038/protex.2018.002 (2018).
Langlois, E. D., Shaw, G. A., Kramar, J. A., Pratt, J. R. & Hurley, D. C. Spring constant calibration of atomic force microscopy cantilevers with a piezosensor transfer standard. Rev. Sci. Instrum. 78, 093705 (2007).
Costa-Silva, B. et al. Pancreatic cancer exosomes initiate pre-metastatic niche formation in the liver. Nat. Cell Biol. 17, 816–826 (2015).
Hoshino, A. et al. Tumour exosome integrins determine organotropic metastasis. Nature 527, 329–335 (2015).
Bolstad, B. M., Irizarry, R. A., Astrand, M. & Speed, T. P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193 (2003).
Monti, S., Tamayo, P., Mesirov, J. & Golub, T. Consensus clustering: a resampling-based method for class discovery and visualization of gene expression microarray data. Mach. Learn. 52, 91–118 (2003).
Golub, T. R. et al. Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science 286, 531–537 (1999).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Ferreira, J. A. et al. Synthesis and optimization of lectin functionalized nanoprobes for the selective recovery of glycoproteins from human body fluids. Anal. Chem. 83, 7035–7043 (2011).
Kolarich, D., Windwarder, M., Alagesan, K. & Altmann, F. Isomer-specific analysis of released N-glycans by LC-ESI MS/MS with porous graphitized carbon. Methods Mol. Biol. 1321, 427–435 (2015).
Jensen, P. H., Karlsson, N. G., Kolarich, D. & Packer, N. H. Structural analysis of N- and O-glycans released from glycoproteins. Nat. Protoc. 7, 1299–1310 (2012).
Ceroni, A. et al. GlycoWorkbench: a tool for the computer-assisted annotation of mass spectra of glycans. J. Prote. Res. 7, 1650–1659 (2008).
Acknowledgements
The authors acknowledge technical support from Wyatt Technology and especially J. Champagne. The authors also acknowledge the Genomics Resource Core facility (WCM) for their high-quality service. The authors thank C. Ghajar and J. Weiss for feedback on the manuscript and members of the Lyden laboratory for discussions. Our study was supported by the National Cancer Institute (U01-CA169538 to D.L.), the National Institutes of Health (NIH; R01-CA169416 to D.L. and H.P.; R01-CA218513 to D.L. and H.Z.), the US Department of Defense (W81XWH-13-10249 to D.L.), W81XWH-13-1-0425 (to D.L., J.Br.), the Sohn Conference Foundation (D.L., I.M., H.P. and H.Z.), the Children’s Cancer and Blood Foundation (D.L.), The Manning Foundation (A.H. and D.L.), The Hartwell Foundation (D.L.), The Nancy C. and Daniel P. Paduano Foundation (D.L.), The Starr Cancer Consortium (H.P. and D.L.; D.L. and H.Z.), the Pediatric Oncology Experimental Therapeutic Investigator Consortium (POETIC; D.L.), the James Paduano Foundation (D.L. and H.P.), the NIH/WCM CTSC (NIH/NCATS: UL1TR00457 to H.M. and H.Z.; UL1TR002384 to D.L., H.M. and H.Z.), the Malcolm Hewitt Wiener Foundation (D.L.), the Champalimaud Foundation (D.L.), the Thompson Family Foundation (D.L., R.S.), U01-CA210240 (D.L.), the Beth Tortolani Foundation (J.Br.), the Charles and Marjorie Holloway Foundation (J.Br.), the Sussman Family Fund (J.Br.), the Lerner Foundation (J.Br.), the Breast Cancer Alliance (J.Br.), the Manhasset Women’s Coalition Against Breast Cancer (J.Br.), the National Institute on Minority Health and Health Disparities (NIMHD) of the NIH (MD007599 to H.M.), NIH/NCATS (UL1TR00457 to H.M.). C.R., A.M., D.F., A.F., A.S. and H.O. acknowledge FEDER (Fundo Europeu de Desenvolvimento Regional funds through COMPETE 2020) POCI, Portugal 2020 (NORTE-01-0145-FEDER-000029) and FCT – Fundação para a Ciência e a Tecnologia in the framework of the project ‘Institute for Research and Innovation in Health Sciences’ (POCI-01-0145-FEDER-007274) and the FCT project POCI-01-0145-FEDER-016585 (PTDC/BBB-EBI/0567/2014). The authors acknowledge FCT for grants to A.M. (SFRH/BPD/75871/2011) and A.F. (SFRH/BPD/111048/2015). D.F. acknowledges FCT (SFRH/BD/110636/2015), the BiotechHealth PhD Programme (PD/0016/2012) and the American Portuguese Biomedical Research Fund.
Author information
Authors and Affiliations
Contributions
H.Z. designed the experimental approach, performed the experimental work, analysed the data, coordinated the project and wrote the manuscript. K.F., S.R., J.F. and H.M. performed zeta potential and stiffness measurements. C.A.R., D.F. A.Mag., J.A.F., A.M.S. and H.O. conducted the glycomics analysis. M.T.M. and H.M. performed and analysed exosome MS. Z.L. conducted and analysed lipidomic MS. H.S.K. conducted proteomic data analysis. H.C., L.B., A.B.M., M.N., A.P.M., P.S., W.B., H.W., A.Mas., G.G. and J.R.C.-R. facilitated exosome isolation/fractionation and other experimental work. J.P.J. and L.C.-G. performed electron microscopy. N.P., M.B. and K.M.-T. performed atomic force microscopy. A.H., I.M. and C.K. contributed to manuscript reading, editing and providing feedback. P.G., A.M.C., J.Bl., R.S., H.P. and J.Br. discussed the hypothesis and contributed to data interpretation and manuscript writing. M.S.B. contributed to studies involving human specimens. D.L. conceived the hypothesis, led the project, interpreted the data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Characterization of AF4 fractions using TEM imaging and NTA analyses and examination of AF4 profiles of nanoparticles derived from cells under different culture and storage conditions.
(a) TEM analysis of particles in AF4 peaks P1 and P5 of B16-F10. The experiment was repeated independently 3 times with similar result. Scale bar, 500 nm. (b) Comparison of the hydrodynamic diameter of each fraction determined by AF4-QELS versus NTA. Individual fractions (time slice, 0.5 min/fraction) were taken every 2 min from 20 to 44 min during the AF4 time course, and subjected to NTA. Results shown are mean ± SEM (n=3 independent samples). Mode size from NTA was utilized. X-axis, time course of AF4 (min); Y-axis, hydrodynamic diameter (nm). (c) The size distribution profiles of representative fractions by NTA (input, unfractionated samples; fractions at 20, 32, and 44 min). Multiple peaks were detected for fractions at 20 and 44 min by NTA. a mode size of 126 nm of Input indicates that NTA cannot efficiently resolve polydisperse samples and is biased towards large particles. This experiment was repeated 3 times independently with similar results. (d) Particle concentration of each fraction measured by NTA. The hydrodynamic diameter of the peak fraction (28 min) was 77nm. Results shown are mean ± SEM (n=3 independent samples). (e-i) AF4 profiles of B16-F10 sEVs collected from technical (blue lines, replicate #1; red lines, replicate #2) (e) and biological replicates (red lines, QELS; blue lines, UV; black (replicate #1) and green dots (replicate #2), hydrodynamic radius; Differences in UV and QELS signal intensity is due to the different amount of input samples for two replicates) (f), kept at either 4 oC or -80 oC for one week (red lines, QELS; blue lines, UV; black (fresh) and green dots (frozen), hydrodynamic radius) (g), cells of different passage numbers (blue and red lines, UV of cells at passage 10 and 18, respectively; black dots, hydrodynamic radius) (h), and under hypoxic versus normoxic conditions for 48 h (blue and red lines, UV for samples cultured with 20% and 1% O2, respectively; black dots, hydrodynamic radius) (i). Experiments were repeated independently 3 times for (e-g) and twice for (h) with similar results. For (i), the experiment was repeated with 3 different cell lines independently with similar results. (j) AF4 and (k) TEM analysis of nanoparticles isolated in parallel from the blank media control and CM of 3-day cultures of B16-F10 and MDA-MB-4175. This experiment was done once. (Red and Blue lines, UV; black dots, hydrodynamic radius; Scale bar, 200 nm.) Statistical source data are provided in Supplementary Table 8.
Supplementary Figure 2 Identification of exomeres and exosome subpopulations released by multiple cancer cell lines.
Shown are AF4 profiles (a) and representative TEM images (b) of unfractionated input samples and pooled fractions of exomeres, Exo- S and Exo-L that derived from various cancer cell lines, including AsPC-1, Pan02, MDA-MB-231-4175, and 4T1. Multiple independent experiments were conducted with similar results for (a) (repeated times: AsPC-1, 9x; Pan02, 16x; 4175, 17x; 4T1, 10x) and (b) (AsPC-1, 3x; Pan02, 2x; 4175, 1x; 4T1, 4x). Scale bar, 200 nm. x- axis, time (min); y-axis (scale) and black dots, hydrodynamic radius (nm); Red and blue lines illustrate the QELS (DLS) intensity and UV absorbance (shown on a relative scale), respectively.
Supplementary Figure 3 Detection of exomeres, Exo-S and Exo-L in samples isolated from the tissue explant cultures.
(a) AF4 profile of exosomes isolated from explant culture of fresh human melanoma tissues. Red and blue lines illustrate the QELS (DLS) intensity and UV absorbance, respectively. This experiment was repeated with 4 independent specimens with similar results. (b) TEM images of exosome samples isolated from the explant culture of normal mouse mammary fat pad and lung tissues. This experiment was repeated independently 2 times with similar results. Scale bar, 500 nm.
Supplementary Figure 4 Proteomic profiling of exomeres and exosome subpopulations derived from multiple cancer cell lines.
(a) Principal component analysis of normalized proteomic mass spectrometry data of exomeres, Exo-S and Exo-L derived from multiple cell lines, including MDA-MB-231-4175, AsPC-1, 4T1, B16-F10 and Pan02. Two independent biological replicates were analyzed for each nanoparticle sample. (b) Heat map illustration of the relative abundance of the Rab family proteins in exomeres, Exo-S and Exo-L. Scale shown is intensity (area) subtracted by mean and divided by row standard deviation (i.e. Δ (area-mean)/SD). (c) Evaluation of the presence of lipoprotein-particle associated proteins (listed in Supplementary Table 5) among the total proteins detected in the exomere, Exo-S and Exo-L derived from different cell lines. Results shown are mean of 2 biologically independent experiments. Statistical source data are provided in Supplementary Table 8. (d) TEM imaging analysis of HDL, LDL and VLDL. Scale bar, 200 nm. This experiment was done once with multiple images showing similar results. (e) Identification of specific association of signaling pathways including hypoxia (FDR, q value = 0.004), microtubule (FDR, q value = 0.002) and coagulation (FDR, q value = 0.013) with exomeres by GSEA (left panel) and the heat map illustration of the expression level of related proteins in different subsets of nanoparticles (right panel). A total of n=30 samples (3 nanoparticle subtypes derived from 5 different cell lines; and two independent biological replicates for each nanoparticle samples) were subjected for Kolmogorov-Smirnov statistical analysis.
Supplementary Figure 5 Mass spectrometric analysis of N-glycans enriched in exomeres, Exo-S and Exo-L derived from AsPC1 and MDA-MB-231-4175.
(a) Total protein profile content of the isolated exomere, Exo-S and Exo-L subpopulations derived from AsPC-1, MDA-MB-231-4175 and B16-F10 assessed by silver staining. This experiment was repeated independently twice for B16-F10 and 4175 with similar results and done once for AsPC-1. (b) The N-glycan mass spectra of particles derived from AsPC-1 (left panel) and MDA-MB-231-4175 (right panel), respectively. This experiment was done once with 3 analytic replicates with similar results. (c) and (d) Quantification of the top six most abundant N-glycan structures identified in the study of AsPC-1 and MDA-MB-231-4175 derived particles. Data shown were quantified and normalized to the most abundant structure in the sample. Results are represented as mean of 2 and 3 independent analytical measurements for AsPC-1 for MDA-MB-231-4175, respectively. NanoHPLC-PGC-HRMS extracted ion chromatograms (EIC) and CID-MS/MS spectra for (e-g) the ion at m/z 1007.38(2-), corresponding to a core-fucosylated complex type N-glycan, characteristic of exomere, and (h-j) the ion at m/z 1111.39(2-), corresponding to a disialylated complex-type N-glycan found in all fractions of B16F10. Fragmentation analysis for extracted ion chromatogram m/z 1007.38 (2-) confirming the structure of this N-glycan in exomeres (e and g) and demonstrating the absence of this N-glycan in Exo-S (f). According to the relative retention time on the PGC column, exomeres contain both α2,3-linked and α2,6-linked sialoglycoforms of the ions at m/z 1111.39(2-) (h). The N-glycan m/z 1111.39(2-) from Exo-S showed N-glycans displaying exclusively α2,3-linked sialic acids based on PGC-LC relative retention time (i). This experiment (e-j) was done once. Statistical source data are provided in Supplementary Table 8
Supplementary Figure 6 Bioanalyzer analysis of the size distribution of DNA associated with exomere, Exo-S and Exo-L derived from B16-F10 (top) and MDA-MB-231-4175 (bottom).
This experiment was repeated twice independently with similar results
Supplementary information
Supplementary Information
Supplementary Figures 1–7 and Supplementary Table legends.
Supplementary Table 1
Types of cancer and corresponding cell lines examined for exosome subpopulations by AF4.
Supplementary Table 2
Complete list of proteins identified by proteomic mass spectrometry analysis of exomeres, Exo-S and Exo-L subpopulations derived from various cancer cell lines.
Supplementary Table 3
Subcellular localization of proteins associated with exomeres, Exo-S and Exo-L revealed by GSEA.
Supplementary Table 4
Lists of proteins that are uniquely associated with or among the top 50 most abundant proteins in exomere, Exo-S and Exo-L derived from different cancer cell lines.
Supplementary Table 5
Proteomic analysis of proteins associated with HDL, LDL and VLDL.
Supplementary Table 6
GSEA of proteins associated with exomeres, Exo-S and Exo-L derived from various cancer cell lines.
Supplementary Table 7
Lipid classes identified in exomeres and exosome subsets derived from different cell lines.
Supplementary Table 8
Statistics source data.
Rights and permissions
About this article
Cite this article
Zhang, H., Freitas, D., Kim, H.S. et al. Identification of distinct nanoparticles and subsets of extracellular vesicles by asymmetric flow field-flow fractionation. Nat Cell Biol 20, 332–343 (2018). https://doi.org/10.1038/s41556-018-0040-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0040-4
This article is cited by
-
Protein cargo in extracellular vesicles as the key mediator in the progression of cancer
Cell Communication and Signaling (2024)
-
Versatile extracellular vesicle-mediated information transfer: intercellular synchronization of differentiation and of cellular phenotypes, and future perspectives
Inflammation and Regeneration (2024)
-
A state-of-the-art review of the recent advances in exosome isolation and detection methods in viral infection
Virology Journal (2024)
-
An in vitro CRISPR screen of cell-free DNA identifies apoptosis as the primary mediator of cell-free DNA release
Communications Biology (2024)
-
Analysis of the longitudinal stability of human plasma miRNAs and implications for disease biomarkers
Scientific Reports (2024)