Abstract
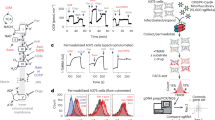
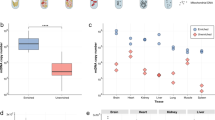
Although mitochondria are ubiquitous organelles, they exhibit tissue-specific morphology, dynamics and function. Here, we describe a robust approach to isolate mitochondria from specific cells of diverse tissue systems in Caenorhabditis elegans. Cell-specific mitochondrial affinity purification (CS-MAP) yields intact and functional mitochondria with exceptional purity and sensitivity (>96% enrichment, >96% purity, and single-cell and single-animal resolution), enabling comparative analyses of protein and nucleic acid composition between organelles isolated from distinct cellular lineages. In animals harbouring a mixture of mutant and wild-type mitochondrial genomes, we use CS-MAP to reveal subtle mosaic patterns of cell-type-specific heteroplasmy across large populations of animals (>10,000 individuals). We demonstrate that the germline is more prone to propagating deleterious mitochondrial genomes than somatic lineages, which we propose is caused by enhanced mtDNA replication in this tissue.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
15 February 2018
In the version of this Technical Report originally published, chromosome representations (indicated by black lines) were missing from Fig. 2a due to a technical error. The corrected version of Fig. 2a is shown below. This has now been amended in all online versions of the Technical Report.
References
West, A. P. & Shadel, G. S. Mitochondrial DNA in innate immune responses and inflammatory pathology. Nat. Rev. Immunol. 17, 363–375 (2017).
Friedman, J. R. & Nunnari, J. Mitochondrial form and function. Nature 505, 335–343 (2014).
Gorman, G. S. et al. Mitochondrial diseases. Nat. Rev. Dis. Prim. 2, 16080 (2016).
Tuppen, H. A., Blakely, E. L., Turnbull, D. M. & Taylor, R. W. Mitochondrial DNA mutations and human disease. Biochim. Biophys. Acta 1797, 113–128 (2010).
Wallace, D. C. A mitochondrial paradigm of metabolic and degenerative diseases, aging, and cancer: a dawn for evolutionary medicine. Annu. Rev. Genet. 39, 359–407 (2005).
Wallace, D. C. & Chalkia, D. Mitochondrial DNA genetics and the heteroplasmy conundrum in evolution and disease. Cold Spring Harb. Perspect. Biol. 5, a021220 (2013).
Frokjaer-Jensen, C. et al. Random and targeted transgene insertion in Caenorhabditis elegans using a modified Mos1 transposon. Nat. Methods 11, 529–534 (2014).
Billing, O., Kao, G. & Naredi, P. Mitochondrial function is required for secretion of DAF-28/insulin in C. elegans. PLoS. ONE 6, e14507 (2011).
Vrablik, T. L., Wang, W. Q., Upadhyay, A. & Hanna-Rose, W. Muscle type-specific responses to NAD+ salvage biosynthesis promote muscle function in Caenorhabditis elegans. Dev. Biol. 349, 387–394 (2011).
Tsang, W. Y. & Lemire, B. D. Stable heteroplasmy but differential inheritance of a large mitochondrial DNA deletion in nematodes. Biochem. Cell. Biol. 80, 645–654 (2002).
Fukushige, T., Hawkins, M. G. & McGhee, J. D. The GATA-factor elt-2 is essential for formation of the Caenorhabditis elegans intestine. Dev. Biol. 198, 286–302 (1998).
Dayama, G., Emery, S. B., Kidd, J. M. & Mills, R. E. The genomic landscape of polymorphic human nuclear mitochondrial insertions. Nucleic Acids Res. 42, 12640–12649 (2014).
Hazkani-Covo, E., Zeller, R. M. & Martin, W. Molecular poltergeists: mitochondrial DNA copies (numts) in sequenced nuclear genomes. PLoS. Genet. 6, e1000834 (2010).
Elson, J. L., Samuels, D. C., Turnbull, D. M. & Chinnery, P. F. Random intracellular drift explains the clonal expansion of mitochondrial DNA mutations with age. Am. J. Human. Genet. 68, 802–806 (2001).
Menzies, R. A. & Gold, P. H. The turnover of mitochondria in a variety of tissues of young adult and aged rats. J. Biol. Chem. 246, 2425–2429 (1971).
Jenuth, J. P., Peterson, A. C. & Shoubridge, E. A. Tissue-specific selection for different mtDNA genotypes in heteroplasmic mice. Nat. Genet. 16, 93–95 (1997).
Gitschlag, B. L. et al. Homeostatic responses regulate selfish mitochondrial genome dynamics in C. elegans. Cell. Metab. 24, 91–103 (2016).
Lemire, B. in WormBook (eds. The C. elegans Research Community) https://doi.org/10.1895/wormbook.1.25.1 (2005).
Narendra, D. P. et al. PINK1 is selectively stabilized on impaired mitochondria to activate parkin. PLoS. Biol. 8, e1000298 (2010).
Lin, Y. F. et al. Maintenance and propagation of a deleterious mitochondrial genome by the mitochondrial unfolded protein response. Nature 533, 416–419 (2016).
Diaz, F. et al. Human mitochondrial DNA with large deletions repopulates organelles faster than full-length genomes under relaxed copy number control. Nucleic Acids Res. 30, 4626–4633 (2002).
Bergstrom, C. T. & Pritchard, J. Germline bottlenecks and the evolutionary maintenance of mitochondrial genomes. Genetics 149, 2135–2146 (1998).
Stewart, J. B., Freyer, C., Elson, J. L. & Larsson, N. G. Purifying selection of mtDNA and its implications for understanding evolution and mitochondrial disease. Nat. Rev. Genet. 9, 657–662 (2008).
Soong, N. W., Hinton, D. R., Cortopassi, G. & Arnheim, N. Mosaicism for a specific somatic mitochondrial-DNA mutation in adult human brain. Nat. Genet. 2, 318–323 (1992).
Chen, Z. et al. Genetic mosaic analysis of a deleterious mitochondrial DNA mutation in Drosophila reveals novel aspects of mitochondrial regulation and function. Mol. Biol. Cell. 26, 674–684 (2015).
Wachsmuth, M., Hubner, A., Li, M., Madea, B. & Stoneking, M. Age-related and heteroplasmy-related variation in human mtDNA copy number. PLoS. Genet. 12, e1005939 (2016).
Chen, W. W., Freinkman, E., Wang, T., Birsoy, K. & Sabatini, D. M. Absolute quantification of matrix metabolites reveals the dynamics of mitochondrial metabolism. Cell 166, 1324–1337 (2016).
Hornig-Do, H. T. et al. Isolation of functional pure mitochondria by superparamagnetic microbeads. Anal. Biochem. 389, 1–5 (2009).
Kayo, S., Bahnemann, J., Klauser, M., Portner, R. & Zeng, A. P. A microfluidic device for immuno-affinity-based separation of mitochondria from cell culture. Lab. Chip 13, 4467–4475 (2013).
Cao, L. Q. et al. The mitochondrial bottleneck occurs without reduction of mtDNA content in female mouse germ cells. Nat. Genet. 39, 386–390 (2007).
Cree, L. M. et al. A reduction of mitochondrial DNA molecules during embryogenesis explains the rapid segregation of genotypes. Nat. Genet. 40, 249–254 (2008).
Daniele, J. R., Heydari, K., Arriaga, E. A. & Dillin, A. Identification and characterization of mitochondrial subtypes in Caenorhabditis elegans via analysis of individual mitochondria by flow cytometry. Anal. Chem. 88, 6309–6316 (2016).
Mattiasson, G. Flow cytometric analysis of isolated liver mitochondria to detect changes relevant to cell death. Cytom. A 60, 145–154 (2004).
Saunders, J. E., Beeson, C. C. & Schnellmann, R. G. Characterization of functionally distinct mitochondrial subpopulations. J. Bioenerg. Biomembr. 45, 87–99 (2013).
Wang, H. et al. cGAL, a temperature-robust GAL4-UAS system for Caenorhabditis elegans. Nat. Methods 14, 145–148 (2017).
Hasegawa, E., Karashima, T., Sumiyoshi, E. & Yamamoto, M. C. elegans CPB-3 interacts with DAZ-1 and functions in multiple steps of germline development. Dev. Biol. 295, 689–699 (2006).
White, J. G., Southgate, E., Thomson, J. N. & Brenner, S. The structure of the nervous system of the nematode Caenorhabditis elegans. Phil. Trans. R. Soc. Lond. B 314, 1–340 (1986).
Varkey, J. P., Muhlrad, P. J., Minniti, A. N., Do, B. & Ward, S. The Caenorhabditis elegans Spe-26 gene is necessary to form spermatids and encodes a protein similar to the actin-associated proteins Kelch and Scruin. Genes. Dev. 9, 1074–1086 (1995).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Zeiser, E., Frokjaer-Jensen, C., Jorgensen, E. & Ahringer, J. MosSCI and gateway compatible plasmid toolkit for constitutive and inducible expression of transgenes in the C. elegans germline. PLoS. ONE 6, e20082 (2011).
Ahier, A. & Jarriault, S. Simultaneous expression of multiple proteins under a single promoter in Caenorhabditis elegans via a versatile 2A-based toolkit. Genetics 196, 605–613 (2014).
Nichols, A. L. et al. The apoptotic engulfment machinery regulates axonal degeneration in C. elegans neurons. Cell. Rep. 14, 1673–1683 (2016).
Ahier, A. & Zuryn, S. Cell-specific mitochondrial affinity purification (CS-MAP) protocol. Protoc. Exch. https://doi.org/10.1038/protex.2017.152 (2017).
Kagias, K., Ahier, A., Fischer, N. & Jarriault, S. Members of the NODE (Nanog and Oct4-associated deacetylase) complex and SOX-2 promote the initiation of a natural cellular reprogramming event in vivo. Proc. Natl. Acad. Sci. USA 109, 6596–6601 (2012).
Restif, C. et al. CeleST: computer vision software for quantitative analysis of C. elegans swim behavior reveals novel features of locomotion. PLoS. Comput. Biol. 10, e1003702 (2014).
Miller, M. A. Sperm and oocyte isolation methods for biochemical and proteomic analysis. Methods Mol. Biol. 351, 193–201 (2006).
Biasini, M. et al. SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res. 42, W252–258 (2014).
Ashkenazy, H. et al. ConSurf 2016: an improved methodology to estimate and visualize evolutionary conservation in macromolecules. Nucleic Acids Res. 44, W344–350 (2016).
Acknowledgements
The authors thank R. Tweedale and L. Richards for comments on the manuscript, M. Hilliard and members of the Zuryn laboratory for discussions and comments, M. Hilliard and T. Bredy for sharing reagents and equipment, R. Amor and L. Hammond for support with microscopy, and A. Ho for support with statistics. Some strains were provided by the CGC, which is funded by the NIH Office of Research Infrastructure Programs (P40 OD010440). This work was supported by NHMRC Project Grant 1128381, a University of Queensland Early Career Researcher Grant (UQECR1608181) and a Stafford Fox Senior Research Fellowship to S.Z., and a University of Queensland International Scholarship to C.Y.D.
Author information
Authors and Affiliations
Contributions
A.A. carried out most experiments. C.-Y.D., A.T., A.B.-G., I.K. and S.Z. contributed some experiments. A.A. and S.Z. designed and interpreted experiments and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
A correction to this article is available online at https://doi.org/10.1038/s41556-018-0055-x.
Supplementary information
Supplementary Information
Supplementary Figures 1–7 and Supplementary References.
Supplementary Table 1
Statistical source data for Figure 6.
Supplementary Table 2
List of primers used in this study.
Supplementary Table 3
List of generated transgenic strains used in this study.
Supplementary Table 4
Antibodies and the working dilutions used in this study.
Rights and permissions
About this article
Cite this article
Ahier, A., Dai, CY., Tweedie, A. et al. Affinity purification of cell-specific mitochondria from whole animals resolves patterns of genetic mosaicism. Nat Cell Biol 20, 352–360 (2018). https://doi.org/10.1038/s41556-017-0023-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-017-0023-x
This article is cited by
-
ATFS-1 counteracts mitochondrial DNA damage by promoting repair over transcription
Nature Cell Biology (2023)
-
Mutant C. elegans mitofusin leads to selective removal of mtDNA heteroplasmic deletions across generations to maintain fitness
BMC Biology (2022)
-
LONP-1 aids propagation of deleterious mtDNA
Nature Cell Biology (2022)
-
LONP-1 and ATFS-1 sustain deleterious heteroplasmy by promoting mtDNA replication in dysfunctional mitochondria
Nature Cell Biology (2022)
-
An energetics perspective on geroscience: mitochondrial protonmotive force and aging
GeroScience (2021)