Abstract
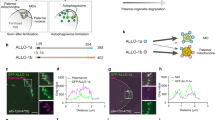
In Caenorhabditis elegans embryos, paternally provided organelles, including mitochondria, are eliminated by a process of selective autophagy called allophagy, the mechanism by which mitochondrial DNA is inherited maternally. However, it remains unclear how paternal organelles are recognized and targeted for autophagy. Here, we identified an autophagy receptor for allophagy, ALLO-1. ALLO-1 is essential for autophagosome formation around paternal organelles and directly binds to the worm LC3 homologue LGG-1 through its LC3-interacting region (LIR) motif. After fertilization, ALLO-1 accumulates on the paternal organelles before autophagosome formation, and this localization depends on the ubiquitin modification of the paternal organelles. We also identified IKKE-1, a worm homologue of the TBK1 and IKKε family kinase, as another critical regulator of allophagy. IKKE-1 interacts with ALLO-1, and the IKKE-1-dependent phosphorylation of ALLO-1 is important for paternal organelle clearance. Thus, we propose that ALLO-1 is the allophagy receptor whose function is regulated by IKKE-1-dependent phosphorylation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Sato, M. & Sato, K. Degradation of paternal mitochondria by fertilization-triggered autophagy in C. elegans embryos. Science 334, 1141–1144 (2011).
Al Rawi, S. et al. Postfertilization autophagy of sperm organelles prevents paternal mitochondrial DNA transmission. Science 334, 1144–1147 (2011).
Sato, M. & Sato, K. Maternal inheritance of mitochondrial DNA: degradation of paternal mitochondria by allogeneic organelle autophagy, allophagy. Autophagy 8, 424–425 (2012).
Al Rawi, S. et al. Allophagy: a macroautophagic process degrading spermatozoid-inherited organelles. Autophagy 8, 421–423 (2012).
Politi, Y. et al. Paternal mitochondrial destruction after fertilization is mediated by a common endocytic and autophagic pathway in Drosophila. Dev. Cell 29, 305–320 (2014).
Rojansky, R., Cha, M. Y. & Chan, D. C. Elimination of paternal mitochondria in mouse embryos occurs through autophagic degradation dependent on PARKIN and MUL1. eLife 5, e17896 (2016).
Zhou, Q. et al. Mitochondrial endonuclease G mediates breakdown of paternal mitochondria upon fertilization. Science 353, 394–399 (2016).
Mizushima, N. & Komatsu, M. Autophagy: renovation of cells and tissues. Cell 147, 728–741 (2011).
Birgisdottir, A. B., Lamark, T. & Johansen, T. The LIR motif–crucial for selective autophagy. J. Cell Sci. 126, 3237–3247 (2013).
Randow, F. & Youle, R. J. Self and nonself: how autophagy targets mitochondria and bacteria. Cell Host Microbe 15, 403–411 (2014).
Rogov, V., Dotsch, V., Johansen, T. & Kirkin, V. Interactions between autophagy receptors and ubiquitin-like proteins form the molecular basis for selective autophagy. Mol. Cell 53, 167–178 (2014).
Okamoto, K. Organellophagy: eliminating cellular building blocks via selective autophagy. J. Cell Biol. 205, 435–445 (2014).
Komatsu, M. et al. Homeostatic levels of p62 control cytoplasmic inclusion body formation in autophagy-deficient mice. Cell 131, 1149–1163 (2007).
Pickrell, A. M. & Youle, R. J. The roles of PINK1, parkin, and mitochondrial fidelity in Parkinson’s disease. Neuron 85, 257–273 (2015).
Yamano, K., Matsuda, N. & Tanaka, K. The ubiquitin signal and autophagy: an orchestrated dance leading to mitochondrial degradation. EMBO Rep. 17, 300–316 (2016).
Thurston, T. L., Ryzhakov, G., Bloor, S., von Muhlinen, N. & Randow, F. The TBK1 adaptor and autophagy receptor NDP52 restricts the proliferation of ubiquitin-coated bacteria. Nat. Immunol. 10, 1215–1221 (2009).
Matsumoto, G., Wada, K., Okuno, M., Kurosawa, M. & Nukina, N. Serine 403 phosphorylation of p62/SQSTM1 regulates selective autophagic clearance of ubiquitinated proteins. Mol. Cell 44, 279–289 (2011).
Wild, P. et al. Phosphorylation of the autophagy receptor optineurin restricts Salmonella growth. Science 333, 228–233 (2011).
Kanki, T. et al. Casein kinase 2 is essential for mitophagy. EMBO Rep. 14, 788–794 (2013).
Tanaka, C. et al. Hrr25 triggers selective autophagy-related pathways by phosphorylating receptor proteins. J. Cell Biol. 207, 91–105 (2014).
Heo, J. M., Ordureau, A., Paulo, J. A., Rinehart, J. & Harper, J. W. The PINK1-PARKIN mitochondrial ubiquitylation pathway drives a program of OPTN/NDP52 recruitment and TBK1 activation to promote mitophagy. Mol. Cell 60, 7–20 (2015).
Lazarou, M. et al. The ubiquitin kinase PINK1 recruits autophagy receptors to induce mitophagy. Nature 524, 309–314 (2015).
Soulat, D. et al. The DEAD-box helicase DDX3X is a critical component of the TANK-binding kinase 1-dependent innate immune response. EMBO J. 27, 2135–2146 (2008).
Meissner, B. et al. Determining the sub cellular localization of proteins within Caenorhabditis elegans body wall muscle. PLoS ONE 6, e19937 (2011).
Lin, L., Yang, P., Huang, X., Zhang, H. & Lu, Q. The scaffold protein EPG-7 links cargo-receptor complexes with the autophagic assembly machinery. J. Cell Biol. 201, 113–129 (2013).
Kirkin, V., McEwan, D. G., Novak, I. & Dikic, I. A role for ubiquitin in selective autophagy. Mol. Cell 34, 259–269 (2009).
Zhang, Y. et al. SEPA-1 mediates the specific recognition and degradation of P granule components by autophagy in C. elegans. Cell 136, 308–321 (2009).
Sato, M., Konuma, R., Sato, K., Tomura, K. & Sato, K. Fertilization-induced K63-linked ubiquitylation mediates clearance of maternal membrane proteins. Development 141, 1324–1331 (2014).
Wei, Y., Chiang, W. C., Sumpter, R. Jr, Mishra, P. & Levine, B. Prohibitin 2 is an inner mitochondrial membrane mitophagy receptor. Cell 168, 224–238 (2017).
Pilli, M. et al. TBK-1 promotes autophagy-mediated antimicrobial defense by controlling autophagosome maturation. Immunity 37, 223–234 (2012).
Matsumoto, G., Shimogori, T., Hattori, N. & Nukina, N. TBK1 controls autophagosomal engulfment of polyubiquitinated mitochondria through p62/SQSTM1 phosphorylation. Hum. Mol. Genet. 24, 4429–4442 (2015).
Richter, B. et al. Phosphorylation of OPTN by TBK1 enhances its binding to Ub chains and promotes selective autophagy of damaged mitochondria. Proc. Nat. Acad. Sci. USA 113, 4039–4044 (2016).
Thurston, T. L. et al. Recruitment of TBK1 to cytosol-invading Salmonella induces WIPI2-dependent antibacterial autophagy. EMBO J. 35, 1779–1792 (2016).
Clement, J. F., Meloche, S. & Servant, M. J. The IKK-related kinases: from innate immunity to oncogenesis. Cell Res. 18, 889–899 (2008).
Helgason, E., Phung, Q. T. & Dueber, E. C. Recent insights into the complexity of Tank-binding kinase 1 signaling networks: the emerging role of cellular localization in the activation and substrate specificity of TBK1. FEBS Lett. 587, 1230–1237 (2013).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Kamath, R. S. et al. Systematic functional analysis of the Caenorhabditis elegans genome using RNAi. Nature 421, 231–237 (2003).
Plowman, G. D., Sudarsanam, S., Bingham, J., Whyte, D. & Hunter, T. The protein kinases of Caenorhabditis elegans: a model for signal transduction in multicellular organisms. Proc. Natl Acad. Sci. USA 96, 13603–13610 (1999).
Pellettieri, J., Reinke, V., Kim, S. K. & Seydoux, G. Coordinate activation of maternal protein degradation during the egg-to-embryo transition in C. elegans. Dev. Cell 5, 451–462 (2003).
Sato, M. et al. Caenorhabditis elegans RME-6 is a novel regulator of RAB-5 at the clathrin-coated pit. Nat. Cell Biol. 7, 559–569 (2005).
Sato, M. et al. Regulation of endocytic recycling by C. elegans Rab35 and its regulator RME-4, a coated-pit protein. EMBO J. 27, 1183–1196 (2008).
Praitis, V., Casey, E., Collar, D. & Austin, J. Creation of low-copy integrated transgenic lines in Caenorhabditis elegans. Genetics 157, 1217–1226 (2001).
Okamoto, H. & Thomson, J. N. Monoclonal antibodies which distinguish certain classes of neuronal and supporting cells in the nervous tissue of the nematode Caenorhabditis elegans. J. Neurosci. 5, 643–653 (1985).
Kanda, Y. Investigation of the freely available easy-to-use software ‘EZR’ for medical statistics. Bone Marrow Transplant. 48, 452–458 (2013).
Okuda, S. et al. jPOSTrepo: an international standard data repository for proteomes. Nucleic Acids Res. 45, D1107–D1111 (2017).
Acknowledgements
We thank S. Tomizawa and the members of the Sato laboratory for technical assistance and discussions, N. Mizushima (The University of Tokyo, Japan) for discussions, S. Mitani (Tokyo Women’s Medical University, Japan) and the Caenorhabditis Genetic Center for supplying the C. elegans strains, N. Matsuda (Tokyo Metropolitan Institute of Medical Science, Japan) and K. Honma (Maebashi Institute of Technology, Japan) for technical advice and Y. Kohara (National Institute of Genetics, Japan), A. Audhya (University of Wisconsin-Madison, USA) and B. Grant (Rutgers University, USA) for the plasmid and antibody. This research was supported by the MEXT KAKENHI (grant numbers 26111503 and 16H01191), The Cell Science Research Foundation, the Uehara Memorial Foundation, Takeda Science Foundation and Joint Usage and Joint Research Programs of the Institute of Advanced Medical Sciences, Tokushima University (to M.S.), and by the JSPS KAKENHI (grant numbers 26291036, 17K19377 and 17H03669), Sumitomo Foundation, Naito Foundation and Ono Medical Research Foundation (to Ken S.). This work was also supported by the Joint Research Program of the Institute for Molecular and Cellular Regulation at Gunma University.
Author information
Authors and Affiliations
Contributions
M.S. and Ken S. designed the experiments and analysed the data. M.S., Katsuya S. and K.T. performed the experiments. H.K. performed LC-MS/MS analysis. M.S., Ken S. and H.K. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–6, Supplementary Table Legends.
Supplementary Table 1
Identification of proteins copurified with GFP-ALLO-1 by LC-MS/MS analysis. List of top 20 peptides copurified with GFP or GFP-ALLO-1. Number of peptides found in LC-MS/MS analysis is shown. Experiment was repeated three times with similar results.
Supplementary Table 2
List of kinases screened in this study. The genes predicted to encode kinases were selected from the genome-wide RNAi library (360 genes).
Supplementary Table 3
List of antibodies used in this study.
Supplementary Table 4
C. elegans strain list used in this study.
Supplementary Table 5
Statistics source data supporting Figures 6b, 7f and 8g.
Rights and permissions
About this article
Cite this article
Sato, M., Sato, K., Tomura, K. et al. The autophagy receptor ALLO-1 and the IKKE-1 kinase control clearance of paternal mitochondria in Caenorhabditis elegans . Nat Cell Biol 20, 81–91 (2018). https://doi.org/10.1038/s41556-017-0008-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-017-0008-9
This article is cited by
-
ALLO-1- and IKKE-1-dependent positive feedback mechanism promotes the initiation of paternal mitochondrial autophagy
Nature Communications (2024)
-
MARC-3, a membrane-associated ubiquitin ligase, is required for fast polyspermy block in Caenorhabditis elegans
Nature Communications (2024)
-
The relationship between autophagy and respiratory viruses
Archives of Microbiology (2024)
-
Semi-in vitro detection of Mg2+-dependent DNase that specifically digest mitochondrial nucleoids in the zygote of Physarum polycephalum
Scientific Reports (2022)
-
Ancestral function of Inhibitors-of-kappaB regulates Caenorhabditis elegans development
Scientific Reports (2020)