Abstract
Immune checkpoint blockade (ICB) therapy does not benefit the majority of treated patients, and those who respond to the therapy can become resistant to it. Here we report the design and performance of systemically administered protease activity sensors conjugated to anti-programmed cell death protein 1 (αPD1) antibodies for the monitoring of antitumour responses to ICB therapy. The sensors consist of a library of mass-barcoded protease substrates that, when cleaved by tumour proteases and immune proteases, are released into urine, where they can be detected by mass spectrometry. By using syngeneic mouse models of colorectal cancer, we show that random forest classifiers trained on mass spectrometry signatures from a library of αPD1-conjugated mass-barcoded activity sensors for differentially expressed tumour proteases and immune proteases can be used to detect early antitumour responses and discriminate resistance to ICB therapy driven by loss-of-function mutations in either the B2m or Jak1 genes. Biomarkers of protease activity may facilitate the assessment of early responses to ICB therapy and the classification of refractory tumours based on resistance mechanisms.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The main data supporting the results in this study are available within the paper and its Supplementary Information. The sequencing datasets generated from murine tumours and analysed from human samples (Riaz, N. et al.11) have been deposited in NCBI’s Gene Expression Omnibus and are accessible through the GEO Series accession numbers GSE192796 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE192796) and GSE91061 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE91061), respectively. Other data generated and analysed during the study are available from the corresponding author on reasonable request. Source data for tumour burdens are provided with this paper.
Code availability
All codes used in this work are available on request from the corresponding author.
References
Ribas, A. & Wolchok, J. D. Cancer immunotherapy using checkpoint blockade. Science 359, 1350–1355 (2018).
Sharma, P. & Allison, J. P. The future of immune checkpoint therapy. Science 348, 56–61 (2015).
Sharma, P., Hu-Lieskovan, S., Wargo, J. A. & Ribas, A. Primary, adaptive, and acquired resistance to cancer immunotherapy. Cell 168, 707–723 (2017).
Kalbasi, A. & Ribas, A. Tumour-intrinsic resistance to immune checkpoint blockade. Nat. Rev. Immunol. https://doi.org/10.1038/s41577-019-0218-4 (2019).
Nishino, M., Ramaiya, N. H., Hatabu, H. & Hodi, F. S. Monitoring immune-checkpoint blockade: response evaluation and biomarker development. Nat. Rev. Clin. Oncol. 14, 655–668 (2017).
Hodi, F. S. et al. Evaluation of immune-related response criteria and RECIST v1.1 in patients with advanced melanoma treated with pembrolizumab. J. Clin. Oncol. 34, 1510–1517 (2016).
Garon, E. B. et al. Pembrolizumab for the treatment of non–small-cell lung cancer. N. Engl. J. Med. 372, 2018–2028 (2015).
Nishino, M. et al. Immune-related tumor response dynamics in melanoma patients treated with pembrolizumab: identifying markers for clinical outcome and treatment decisions. Clin. Cancer Res. 23, 4671–4679 (2017).
Gerwing, M. et al. The beginning of the end for conventional RECIST — novel therapies require novel imaging approaches. Nat. Rev. Clin. Oncol. 16, 442–458 (2019).
Mandal, R. & Chan, T. A. Personalized oncology meets immunology: the path toward precision immunotherapy. Cancer Discov. 6, 703–713 (2016).
Riaz, N. et al. Tumor and microenvironment evolution during immunotherapy with nivolumab. Cell 171, 934–949.e16 (2017).
Fairfax, B. P. et al. Peripheral CD8+ T cell characteristics associated with durable responses to immune checkpoint blockade in patients with metastatic melanoma. Nat. Med. 26, 193–199 (2020).
Valpione, S. et al. Immune awakening revealed by peripheral T cell dynamics after one cycle of immunotherapy. Nat. Cancer 1, 210–221 (2020).
Goldberg, S. B. et al. Early assessment of lung cancer immunotherapy response via circulating tumor DNA. Clin. Cancer Res. 24, 1872–1880 (2018).
Kessenbrock, K., Plaks, V. & Werb, Z. Matrix metalloproteinases: regulators of the tumor microenvironment. Cell 141, 52–67 (2010).
Dudani, J. S., Warren, A. D. & Bhatia, S. N. Harnessing protease activity to improve cancer care. Annu. Rev. Cancer Biol. 2, 353–376 (2018).
Martínez-Lostao, L., Anel, A. & Pardo, J. How do cytotoxic lymphocytes kill cancer cells? Clin. Cancer Res. 21, 5047–5056 (2015).
Hilderbrand, S. A. & Weissleder, R. Near-infrared fluorescence: application to in vivo molecular imaging. Curr. Opin. Chem. Biol. 14, 71–79 (2010).
Sanman, L. E. & Bogyo, M. Activity-based profiling of proteases. Annu. Rev. Biochem. 83, 249–273 (2014).
Savariar, E. N. et al. Real-time in vivo molecular detection of primary tumors and metastases with ratiometric activatable cell-penetrating peptides. Cancer Res. 73, 855–864 (2013).
Larimer, B. M. et al. Granzyme B PET imaging as a predictive biomarker of immunotherapy response. Cancer Res. 77, 2318–2327 (2017).
Kwong, G. A. et al. Synthetic biomarkers: a twenty-first century path to early cancer detection. Nat. Rev. Cancer 21, 655–668 (2021).
Kwong, G. A. et al. Mass-encoded synthetic biomarkers for multiplexed urinary monitoring of disease. Nat. Biotechnol. 31, 63–70 (2013).
Lin, K. Y., Kwong, G. A., Warren, A. D., Wood, D. K. & Bhatia, S. N. Nanoparticles that sense thrombin activity as synthetic urinary biomarkers of thrombosis. ACS Nano 7, 9001–9009 (2013).
Warren, A. D., Kwong, G. A., Wood, D. K., Lin, K. Y. & Bhatia, S. N. Point-of-care diagnostics for noncommunicable diseases using synthetic urinary biomarkers and paper microfluidics. Proc. Natl Acad. Sci. USA 111, 3671–3676 (2014).
Kwong, G. A. et al. Mathematical framework for activity-based cancer biomarkers. Proc. Natl Acad. Sci. USA 112, 12627–12632 (2015).
Mac, Q. D. et al. Non-invasive early detection of acute transplant rejection via nanosensors of granzyme B activity. Nat. Biomed. Eng. 3, 281–291 (2019).
Kirkpatrick, J. D. et al. Urinary detection of lung cancer in mice via noninvasive pulmonary protease profiling. Sci. Transl. Med. 12, eaaw0262 (2020).
Cazanave, S. C. et al. Peptide-based urinary monitoring of fibrotic nonalcoholic steatohepatitis by mass-barcoded activity-based sensors. Sci. Transl. Med. 13, eabe8939 (2021).
Casciola-Rosen, L. et al. Mouse and human granzyme B have distinct tetrapeptide specificities and abilities to recruit the bid pathway. J. Biol. Chem. 282, 4545–4552 (2007).
Harris, J. L., Peterson, E. P., Hudig, D., Thornberry, N. A. & Craik, C. S. Definition and redesign of the extended substrate specificity of granzyme B. J. Biol. Chem. 273, 27364–27373 (1998).
Ruggles, S. W., Fletterick, R. J. & Craik, C. S. Characterization of structural determinants of granzyme B reveals potent mediators of extended substrate specificity. J. Biol. Chem. 279, 30751–30759 (2004).
He, S., Li, J., Lyu, Y., Huang, J. & Pu, K. Near-infrared fluorescent macromolecular reporters for real-time imaging and urinalysis of cancer immunotherapy. J. Am. Chem. Soc. 142, 7075–7082 (2020).
Zhang, Y. et al. Activatable polymeric nanoprobe for near-infrared fluorescence and photoacoustic imaging of T lymphocytes. Angew. Chem. Int. Ed. 133, 5986–5992 (2021).
Efremova, M. et al. Targeting immune checkpoints potentiates immunoediting and changes the dynamics of tumor evolution. Nat. Commun. 9, 32 (2018).
Villanueva, J. et al. Differential exoprotease activities confer tumor-specific serum peptidome patterns. J. Clin. Invest. 116, 271–284 (2006).
Villanueva, J. et al. A sequence-specific exopeptidase activity test (SSEAT) for ‘functional’ biomarker discovery. Mol. Cell. Proteomics 7, 509–518 (2008).
Werle, M. & Bernkop-Schnürch, A. Strategies to improve plasma half life time of peptide and protein drugs. Amino Acids 30, 351–367 (2006).
Diao, L. & Meibohm, B. Pharmacokinetics and pharmacokinetic-pharmacodynamic correlations of therapeutic peptides. Clin. Pharmacokinet. 52, 855–868 (2013).
Desnoyers, L. R. et al. Tumor-specific activation of an EGFR-targeting probody enhances therapeutic index. Sci. Transl. Med. 5, 207ra144 (2013).
Strohl, W. R. Fusion proteins for half-life extension of biologics as a strategy to make biobetters. BioDrugs 29, 215–239 (2015).
Duraiswamy, J., Kaluza, K. M., Freeman, G. J. & Coukos, G. Dual blockade of PD-1 and CTLA-4 combined with tumor vaccine effectively restores T-cell rejection function in tumors. Cancer Res. 73, 3591–3603 (2013).
Selby, M. J. et al. Preclinical development of ipilimumab and nivolumab combination immunotherapy: mouse tumor models, in vitro functional studies, and cynomolgus macaque toxicology. PLoS ONE 11, e0161779 (2016).
Brinkman, E. K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 42, e168 (2014).
Miller, B. C. et al. Subsets of exhausted CD8+ T cells differentially mediate tumor control and respond to checkpoint blockade. Nat. Immunol. 20, 326–336 (2019).
Im, S. J. et al. Defining CD8+ T cells that provide the proliferative burst after PD-1 therapy. Nature 537, 417–421 (2016).
Jansen, C. S. et al. An intra-tumoral niche maintains and differentiates stem-like CD8 T cells. Nature 576, 465–470 (2019).
Paley, M. A. et al. Progenitor and terminal subsets of CD8+ T cells cooperate to contain chronic viral infection. Science 338, 1220–1225 (2012).
Thommen, D. S. et al. A transcriptionally and functionally distinct PD-1+ CD8+ T cell pool with predictive potential in non-small-cell lung cancer treated with PD-1 blockade. Nat. Med. 24, 994–1004 (2018).
Gupta, P. K. et al. CD39 expression identifies terminally exhausted CD8+ T cells. PLoS Pathog. 11, e1005177 (2015).
Canale, F. P. et al. CD39 expression defines cell exhaustion in tumor-infiltrating CD8+ T cells. Cancer Res. 78, 115–128 (2018).
Horton, B. L., Williams, J. B., Cabanov, A., Spranger, S. & Gajewski, T. F. Intratumoral CD8+ T-cell apoptosis is a major component of T-cell dysfunction and impedes anti-tumor immunity. Cancer Immunol. Res. 6, 14–24 (2018).
Liberzon, A. et al. The molecular signatures database hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Schwartz, L. H. et al. RECIST 1.1 – update and clarification: from the RECIST Committee. Eur. J. Cancer 62, 132–137 (2016).
Arlot, S. & Celisse, A. A survey of cross-validation procedures for model selection. Stat. Surv. 4, 40–79 (2010).
Patel, S. P. & Kurzrock, R. PD-L1 expression as a predictive biomarker in cancer immunotherapy. Mol. Cancer Ther. 14, 847–856 (2015).
Chan, T. A. et al. Development of tumor mutation burden as an immunotherapy biomarker: utility for the oncology clinic. Ann. Oncol. 30, 44–56 (2019).
Cristescu, R. et al. Pan-tumor genomic biomarkers for PD-1 checkpoint blockade–based immunotherapy. Science 362, eaar3593 (2018).
Chang, L., Chang, M., Chang, H. M. & Chang, F. Microsatellite instability: a predictive biomarker for cancer immunotherapy. Appl. Immunohistochem. Mol. Morphol. 26, e15 (2018).
Bratman, S. V. et al. Personalized circulating tumor DNA analysis as a predictive biomarker in solid tumor patients treated with pembrolizumab. Nat. Cancer 1, 873–881 (2020).
Tietze, J. K. et al. The proportion of circulating CD45RO+CD8+ memory T cells is correlated with clinical response in melanoma patients treated with ipilimumab. Eur. J. Cancer 75, 268–279 (2017).
Chen, P.-L. et al. Analysis of immune signatures in longitudinal tumor samples yields insight into biomarkers of response and mechanisms of resistance to immune checkpoint blockade. Cancer Discov. 6, 827–837 (2016).
Tumeh, P. C. et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature 515, 568–571 (2014).
Jiang, P. et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat. Med. 24, 1550–1558 (2018).
Amaria, R. N. et al. Neoadjuvant immune checkpoint blockade in high-risk resectable melanoma. Nat. Med. 24, 1649–1654 (2018).
Grasso, C. S. et al. Conserved interferon-γ signaling drives clinical response to immune checkpoint blockade therapy in melanoma. Cancer Cell 38, 500–515.e3 (2020).
Larimer, B. M. et al. The effectiveness of checkpoint inhibitor combinations and administration timing can be measured by granzyme B PET imaging. Clin. Cancer Res. 25, 1196–1205 (2019).
Nguyen, A. et al. Granzyme B nanoreporter for early monitoring of tumor response to immunotherapy. Sci. Adv. 6, eabc2777 (2020).
Zhao, N. et al. In vivo measurement of granzyme proteolysis from activated immune cells with PET. ACS Cent. Sci. 7, 1638–1649 (2021).
Carter, H. B. & Pearson, J. D. PSA velocity for the diagnosis of early prostate cancer. A new concept. Urol. Clin. North Am. 20, 665–670 (1993).
Vickers, A. J. et al. Prostate-specific antigen velocity for early detection of prostate cancer. Eur. Urol. 56, 753–760 (2009).
La Thangue, N. B. & Kerr, D. J. Predictive biomarkers: a paradigm shift towards personalized cancer medicine. Nat. Rev. Clin. Oncol. 8, 587–596 (2011).
Salgado, R. et al. Steps forward for cancer precision medicine. Nat. Rev. Drug Discov. 17, 1–2 (2018).
Brown, N. A. & Elenitoba-Johnson, K. S. J. Enabling precision oncology through precision diagnostics. Annu. Rev. Pathol. 15, 97–121 (2020).
Ramos-Casals, M. et al. Immune-related adverse events of checkpoint inhibitors. Nat. Rev. Dis. Primers 6, 38 (2020).
Esfahani, K. et al. Moving towards personalized treatments of immune-related adverse events. Nat. Rev. Clin. Oncol. 17, 504–515 (2020).
Postow, M. A., Sidlow, R. & Hellmann, M. D. Immune-related adverse events associated with immune checkpoint blockade. N. Engl. J. Med. https://doi.org/10.1056/NEJMra1703481 (2018).
Johnson, D. B. et al. Fulminant myocarditis with combination immune checkpoint blockade. N. Engl. J. Med. 375, 1749–1755 (2016).
Goldinger, S. M. et al. Cytotoxic cutaneous adverse drug reactions during anti-PD-1 therapy. Clin. Cancer Res. 22, 4023–4029 (2016).
Hua, C. et al. Association of vitiligo with tumor response in patients with metastatic melanoma treated with pembrolizumab. JAMA Dermatol. 152, 45–51 (2016).
Zhang, X. et al. Hepatitis B virus reactivation in cancer patients with positive hepatitis B surface antigen undergoing PD-1 inhibition. J. Immunother. Cancer 7, 322 (2019).
Del Castillo, M. et al. The spectrum of serious infections among patients receiving immune checkpoint blockade for the treatment of melanoma. Clin. Infect. Dis. 63, 1490–1493 (2016).
Fujita, K. et al. Emerging concerns of infectious diseases in lung cancer patients receiving immune checkpoint inhibitor therapy. Respir. Med. 146, 66–70 (2019).
Hutchinson, J. A. et al. Virus-specific memory T cell responses unmasked by immune checkpoint blockade cause hepatitis. Nat. Commun. 12, 1439 (2021).
Beerli, R. R., Hell, T., Merkel, A. S. & Grawunder, U. Sortase enzyme-mediated generation of site-specifically conjugated antibody drug conjugates with high in vitro and in vivo potency. PLoS ONE 10, e0131177 (2015).
Jeger, S. et al. Site-specific and stoichiometric modification of antibodies by bacterial transglutaminase. Angew. Chem. Int. Ed. Engl. 49, 9995–9997 (2010).
Yu, C. et al. Proximity-induced site-specific antibody conjugation. Bioconjug. Chem. 29, 3522–3526 (2018).
Puente, X. S., Sánchez, L. M., Overall, C. M. & López-Otín, C. Human and mouse proteases: a comparative genomic approach. Nat. Rev. Genet. 4, 544–558 (2003).
Kaiserman, D. et al. The major human and mouse granzymes are structurally and functionally divergent. J. Cell Biol. 175, 619–630 (2006).
Aguilera, T. A., Olson, E. S., Timmers, M. M., Jiang, T. & Tsien, R. Y. Systemic in vivo distribution of activatable cell penetrating peptides is superior to that of cell penetrating peptides. Integr. Biol. 1, 371–381 (2009).
Whitley, M. J. et al. A mouse-human phase 1 co-clinical trial of a protease-activated fluorescent probe for imaging cancer. Sci. Transl. Med. 8, 320ra4 (2016).
Timmer, J. C. & Salvesen, G. S. Caspase substrates. Cell Death Differ. 14, 66–72 (2007).
Poreba, M. et al. Unnatural amino acids increase sensitivity and provide for the design of highly selective caspase substrates. Cell Death Differ. 21, 1482–1492 (2014).
Rut, W. et al. Recent advances and concepts in substrate specificity determination of proteases using tailored libraries of fluorogenic substrates with unnatural amino acids. Biol. Chem. 396, 329–337 (2015).
Miller, M. A. et al. Proteolytic activity matrix analysis (PrAMA) for simultaneous determination of multiple protease activities. Integr. Biol. 3, 422–438 (2011).
Zhuang, Q., Holt, B. A., Kwong, G. A. & Qiu, P. Deconvolving multiplexed protease signatures with substrate reduction and activity clustering. PLoS Comput. Biol. 15, e1006909 (2019).
Austin, R. J. et al. TriTACs, a novel class of T-cell–engaging protein constructs designed for the treatment of solid tumors. Mol. Cancer Ther. 20, 109–120 (2021).
Triplett, T. A. et al. Reversal of indoleamine 2,3-dioxygenase–mediated cancer immune suppression by systemic kynurenine depletion with a therapeutic enzyme. Nat. Biotechnol. 36, 758–764 (2018).
Clark, M. F., Lister, R. M. & Bar-Joseph, M. ELISA techniques. Methods Enzymol 118, 742–766 (1986).
Acknowledgements
This work was funded by the NIH Director’s New Innovator Award DP2HD091793 and the National Cancer Institute R01 grant 5R01CA237210. Q.D.M. and A.S. were supported by the NSF Graduate Research Fellowships Program (Grant No. DGE-1650044). G.A.K. holds a Career Award at the Scientific Interface from the Burroughs Wellcome Fund. P.Q. is an ISAC Marylou Ingram Scholar and a Carol Ann and David D. Flanagan Faculty Fellow. This work was performed in part at the Georgia Tech Institute for Electronics and Nanotechnology, a member of the National Nanotechnology Coordinated Infrastructure, which is supported by the National Science Foundation (Grant ECCS-1542174). This content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. We thank the staff at Georgia Tech mass spectrometry core, flow cytometry analysis core and the animal facility for their assistance in performing our studies.
Author information
Authors and Affiliations
Contributions
Q.D.M., J.R.B. and G.A.K. conceived the idea. Q.D.M., A.S., H.P., C.X., J.R.B, F.-Y.S., S.Z.S., P.Q. and G.A.K. designed experiments and interpreted results. Q.D.M., A.S., H.P., C.X., J.R.B, F.-Y.S., S.Z.S., H.S., A.M.H. and T.T.L. carried out the experiments. Q.D.M., A.S. and G.A.K. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
G.A.K. is co-founder of and serves as consultant to Glympse Bio and Satellite Bio. This study could affect his personal financial status. The terms of this arrangement have been reviewed and approved by Georgia Tech in accordance with its conflict-of-interest policies. Q.D.M., J.R.B. and G.A.K are listed as inventors on a patent application (PCT/US2019/050530) pertaining to the results of the paper. The patent applicant is the Georgia Tech Research Corporation. The names of the inventors are Quoc Mac, James Bowen and Gabriel Kwong. The patent is currently pending/published (publication number WO2020055952A1). The mass-barcoded antibody–sensor conjugates and related applications are covered in this patent.
Peer review
Peer review information
Nature Biomedical Engineering thanks Charles Craik, Matt Trau and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Synthesis strategy for antibody-sensor conjugates for biodistribution of antibody and GzmB cleavage.
a, Synthetic scheme of antibody-sensor conjugate labelled with near infrared fluorophore VT750 to determine antibody biodistribution. b, Synthetic scheme of NIR activatable GzmB probe, consisting of fluorescently quenched GzmB substrate conjugated to αPD1 antibody.
Extended Data Fig. 2 Biodistribution of antibody-sensor conjugates and GzmB cleavage.
a, Schematic of antibody-sensor conjugate labelled with near infrared fluorophore VT750 to determine antibody biodistribution. b, Representative whole organ images and c, quantification of distribution of fluorescent signal in tumour and major organs (dLNs, tumour draining lymph nodes) (two-way ANOVA with Sidak’s post-test and correction for multiple comparisons, ns = not significant, n = 5 biological replicates, error bars depict s.e.m.). d, Schematic and in vitro cleavage of NIR activatable GzmB probe labelled with quenched near infrared fluorophore 800CW to localize site of probe activation (FC, fold change; ****P < 0.0001, n = 3 technically independent wells, error bars depict s.e.m.). e, Representative whole organ images and f, quantification of reporter fluorescence after treatment with Iso-GS or αPD1-GS (two-way ANOVA with Sidak’s post-test and correction for multiple comparisons, *P = 0.040, n = 4-5 biological replicates, error bars depict s.e.m.).
Extended Data Fig. 3 Diagnostic performance of αPD1-GS in ICB nonresponsive models.
a, Average and b, individual tumour growth curves of CT26 tumour bearing mice treated with either αCTLA4-GS or matched IgG2 isotype control (Iso-GS) (two-way ANOVA with Sidak’s post-test and correction for multiple comparisons, ns = not significant, n = 10-11 biological replicates, error bars depict s.e.m.). Black arrows denote the treatment time points. c, Normalized urine fluorescence of mice with CT26 tumours after each administration of αCTLA4-GS or Iso-GS (two-way ANOVA with Sidak’s post-test and correction for multiple comparisons, ns = not significant, n = 10-11 biological replicates). d, Average and e, individual tumour growth curves of CT26 tumour bearing mice treated with αPD1-GS or matched IgG1 isotype control (Iso-GS) (two-way ANOVA with Sidak’s post-test and correction for multiple comparisons, ns = not significant, n = 6 biological replicates, error bars depict s.e.m.). Black arrows denote the treatment time points. f, Normalized urine fluorescence of mice with CT26 tumours after each administration of αPD1-GS or Iso-GS (two-way ANOVA with Sidak’s post-test and correction for multiple comparisons, ns = not significant, n = 6 biological replicates).
Extended Data Fig. 4 Differential expression analysis of proteases in ICB response and resistance.
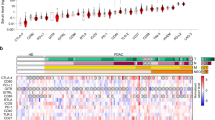
a, Heatmaps showing row-normalized expression (FPKM) of proteases differentially expressed between αPD1-treated WT tumours and IgG1-treated controls (n = 5 biological replicates). b, (Left) Volcano plots summarizing differentially expressed proteases between αPD1-treated B2m−/− and Jak1−/− MC38 tumours (n = 5 biological replicates). The threshold for differentially expressed genes (opaque dots) was defined as P value ≤ 0.05 and |log2(fold change)| ≥ 1. (Right) Heatmaps showing row-normalized expression (FPKM) of proteases differentially expressed between B2m−/− and Jak1−/− MC38 tumours (n = 5 biological replicates). c, (Left) Volcano plots summarizing differentially expressed proteases between human tumours from responders (CR + PR) and non-responders (PD) (n = 5 independent patient samples). The threshold for differentially expressed genes was defined as P value ≤ 0.01 and |log2(fold change)| ≥ 1. (Right) Heatmaps showing row-normalized expression (FPKM) of proteases differentially expressed between human tumours from responders (CR + PR) and non-responders (PD) (n = 5 independent patient samples).
Supplementary information
Supplementary Information
Supplementary figures and tables.
Source data
Source Data Fig. 1
Source data for tumour burden.
Source Data Fig. 3
Source data for tumour burden.
Source Data Fig. 4
Source data for tumour burden.
Source Data Extended Data Fig. 3
Source data for tumour burden.
Rights and permissions
About this article
Cite this article
Mac, Q.D., Sivakumar, A., Phuengkham, H. et al. Urinary detection of early responses to checkpoint blockade and of resistance to it via protease-cleaved antibody-conjugated sensors. Nat. Biomed. Eng 6, 310–324 (2022). https://doi.org/10.1038/s41551-022-00852-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-022-00852-y
This article is cited by
-
Artificial urinary biomarker probes for diagnosis
Nature Reviews Bioengineering (2024)