Abstract
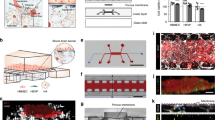
The neurovascular unit, which consists of vascular cells surrounded by astrocytic end-feet and neurons, controls cerebral blood flow and the permeability of the blood–brain barrier (BBB) to maintain homeostasis in the neuronal milieu. Studying how some pathogens and drugs can penetrate the human BBB and disrupt neuronal homeostasis requires in vitro microphysiological models of the neurovascular unit. Here we show that the neurotropism of Cryptococcus neoformans—the most common pathogen causing fungal meningitis—and its ability to penetrate the BBB can be modelled by the co-culture of human neural stem cells, brain microvascular endothelial cells and brain vascular pericytes in a human-neurovascular-unit-on-a-chip maintained by a stepwise gravity-driven unidirectional flow and recapitulating the structural and functional features of the BBB. We found that the pathogen forms clusters of cells that penetrate the BBB without altering tight junctions, suggesting a transcytosis-mediated mechanism. The neurovascular-unit-on-a-chip may facilitate the study of the mechanisms of brain infection by pathogens, and the development of drugs for a range of brain diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The main data supporting the findings of this study are available within the paper and its Supplementary Information. The datasets from the quantitative analyses generated during this study, including source data and the data used to make the figures as well as all images, are too large and numerous to be publicly shared, but they are available for research purposes from the corresponding authors on reasonable request. RNA-seq data have been deposited in the NCBI Gene Expression Omnibus (GEO), with accession number GSE171937.
Code availability
MATLAB was used to analyse results of fluorescence recovery after photobleaching experiments. The codes used for the analysis can be provided on request.
References
Abbott, N. J., Rönnbäck, L. & Hansson, E. Astrocyte–endothelial interactions at the blood–brain barrier. Nat. Rev. Neurosci. 7, 41–53 (2006).
Wolburg, H. & Lippoldt, A. Tight junctions of the blood–brain barrier: development, composition and regulation. Vasc. Pharmacol. 38, 323–337 (2002).
Schinkel, A. H. P-glycoprotein, a gatekeeper in the blood–brain barrier. Adv. Drug Deliv. Rev. 36, 179–194 (1999).
Karssen, A. et al. Multidrug resistance P-glycoprotein hampers the access of cortisol but not of corticosterone to mouse and human brain. Endocrinology 142, 2686–2694 (2001).
Willner, P. Validation criteria for animal models of human mental disorders: learned helplessness as a paradigm case. Prog. Neuropsychopharmacol. Biol. Psychiatry 10, 677–690 (1986).
Dixit, R. & Boelsterli, U. A. Healthy animals and animal models of human disease(s) in safety assessment of human pharmaceuticals, including therapeutic antibodies. Drug Discov. Today 12, 336–342 (2007).
Vu, K., Weksler, B., Romero, I., Couraud, P.-O. & Gelli, A. Immortalized human brain endothelial cell line HCMEC/D3 as a model of the blood–brain barrier facilitates in vitro studies of central nervous system infection by Cryptococcus neoformans. Eukaryot. Cell 8, 1803–1807 (2009).
Vu, K., Eigenheer, R. A., Phinney, B. S. & Gelli, A. Cryptococcus neoformans promotes its transmigration into the CNS by inducing molecular and cellular changes in brain endothelial cells. Infect. Immun. 81, 3139–3147 (2013).
Vu, K. et al. Invasion of the central nervous system by Cryptococcus neoformans requires a secreted fungal metalloprotease. mBio 5, e01101–e01114 (2014).
Cardoso, F. L., Brites, D. & Brito, M. A. Looking at the blood–brain barrier: molecular anatomy and possible investigation approaches. Brain Res. Rev. 64, 328–363 (2010).
Dalvi, S. et al. The blood brain barrier — regulation of fatty acid and drug transport. Neurochemistry (ed. Heinbockel, T.) 1–33 (2014).
Booth, R. & Kim, H. Characterization of a microfluidic in vitro model of the blood–brain barrier (μBBB). Lab Chip 12, 1784–1792 (2012).
Prabhakarpandian, B. et al. SyM-BBB: a microfluidic blood brain barrier model. Lab Chip 13, 1093–1101 (2013).
Griep, L. et al. BBB on chip: microfluidic platform to mechanically and biochemically modulate blood–brain barrier function. Biomed. Microdevices 15, 145–150 (2013).
Yeon, J. H. et al. Reliable permeability assay system in a microfluidic device mimicking cerebral vasculatures. Biomed. Microdevices 14, 1141–1148 (2012).
Brown, J. A. et al. Metabolic consequences of inflammatory disruption of the blood–brain barrier in an organ-on-chip model of the human neurovascular unit. J. Neuroinflammation 13, 1–17 (2016).
Odijk, M. et al. Measuring direct current trans-epithelial electrical resistance in organ-on-a-chip microsystems. Lab Chip 15, 745–752 (2015).
Herland, A. et al. Distinct contributions of astrocytes and pericytes to neuroinflammation identified in a 3D human blood–brain barrier on a chip. PLoS ONE 11, e0150360 (2016).
Campisi, M. et al. 3D self-organized microvascular model of the human blood-brain barrier with endothelial cells, pericytes and astrocytes. Biomaterials 180, 117–129 (2018).
Maoz, B. M. et al. A linked organ-on-chip model of the human neurovascular unit reveals the metabolic coupling of endothelial and neuronal cells. Nat. Biotechnol. 36, 865 (2018).
Park, T.-E. et al. Hypoxia-enhanced Blood-Brain Barrier Chip recapitulates human barrier function and shuttling of drugs and antibodies. Nat. Commun. 10, 2621 (2019).
Rajasingham, R. et al. Global burden of disease of HIV-associated cryptococcal meningitis: an updated analysis. Lancet Infect. Dis. 17, 873–881 (2017).
Sabiiti, W. & May, R. C. Mechanisms of infection by the human fungal pathogen Cryptococcus neoformans. Future Microbiol. 7, 1297–1313 (2012).
Xue, C. Cryptococcus and beyond—inositol utilization and its implications for the emergence of fungal virulence. PLoS Pathog. 8, e1002869 (2012).
Yang, K. et al. A microfluidic array for quantitative analysis of human neural stem cell self-renewal and differentiation in three-dimensional hypoxic microenvironment. Biomaterials 34, 6607–6614 (2013).
Shin, Y. et al. Reconstituting vascular microenvironment of neural stem cell niche in three‐dimensional extracellular matrix. Adv. Healthc. Mater. 3, 1457–1464 (2014).
Cucullo, L., Hossain, M., Puvenna, V., Marchi, N. & Janigro, D. The role of shear stress in blood–brain barrier endothelial physiology. BMC Neurosci. 12, 40 (2011).
Shin, J. et al. Tissue adhesive catechol-modified hyaluronic acid hydrogel for effective, minimally invasive cell therapy. Adv. Funct. Mater. 25, 3814–3824 (2015).
Yang, K. et al. Polydopamine-mediated surface modification of scaffold materials for human neural stem cell engineering. Biomaterials 33, 6952–6964 (2012).
Lee, H., Rho, J. & Messersmith, P. B. Facile conjugation of biomolecules onto surfaces via mussel adhesive protein inspired coatings. Adv. Mater. 21, 431–434 (2009).
Park, H.-J. et al. Catechol-functionalized hyaluronic acid hydrogels enhance angiogenesis and osteogenesis of human adipose-derived stem cells in critical tissue defects. Biomacromolecules 17, 1939–1948 (2016).
Shen, Q. et al. Endothelial cells stimulate self-renewal and expand neurogenesis of neural stem cells. Science 304, 1338–1340 (2004).
Mi, H., Haeberle, H. & Barres, B. A. Induction of astrocyte differentiation by endothelial cells. J. Neurosci. 21, 1538–1547 (2001).
Carvey, P. M., Hendey, B. & Monahan, A. J. The blood–brain barrier in neurodegenerative disease: a rhetorical perspective. J. Neurochem. 111, 291–314 (2009).
Armulik, A. et al. Pericytes regulate the blood–brain barrier. Nature 468, 557 (2010).
Brachvogel, B. et al. Isolated Anxa5+/Sca-1+ perivascular cells from mouse meningeal vasculature retain their perivascular phenotype in vitro and in vivo. Exp. Cell. Res. 313, 2730–2743 (2007).
Baumann, N. & Pham-Dinh, D. Biology of oligodendrocyte and myelin in the mammalian central nervous system. Physiol. Rev. 81, 871–927 (2001).
Brown, J. A. et al. Recreating blood–brain barrier physiology and structure on chip: A novel neurovascular microfluidic bioreactor. Biomicrofluidics 9, 054124 (2015).
Ahn, S. I. et al. Microengineered human blood–brain barrier platform for understanding nanoparticle transport mechanisms. Nat. Commun. 11, 175 (2020).
Essodaigui, M., Broxterman, H. & Garnier-Suillerot, A. Kinetic analysis of calcein and calcein−acetoxymethylester efflux mediated by the multidrug resistance protein and P-glycoprotein. Biochemistry 37, 2243–2250 (1998).
Deli, M. A., Ábrahám, C. S., Kataoka, Y. & Niwa, M. Permeability studies on in vitro blood–brain barrier models: physiology, pathology, and pharmacology. Cell. Mol. Neurobiol. 25, 59–127 (2005).
Mark, K. S., Trickler, W. J. & Miller, D. W. Tumor necrosis factor-α induces cyclooxygenase-2 expression and prostaglandin release in brain microvessel endothelial cells. J. Pharmacol. Exp. Ther. 297, 1051–1058 (2001).
Varatharaj, A. & Galea, I. The blood–brain barrier in systemic inflammation. Brain Behav. Immun. 60, 1–12 (2017).
May, R. C., Stone, N. R., Wiesner, D. L., Bicanic, T. & Nielsen, K. Cryptococcus: from environmental saprophyte to global pathogen. Nat. Rev. Microbiol. 14, 106 (2016).
Kaufman-Francis, K. et al. The early innate immune response to, and phagocyte-dependent entry of, Cryptococcus neoformans map to the perivascular space of cortical post-capillary venules in neurocryptococcosis. Am. J. Pathol. 188, 1653–1665 (2018).
Ngamskulrungroj, P., Chang, Y., Sionov, E. & Kwon-Chung, K. J. The primary target organ of Cryptococcus gattii is different from that of Cryptococcus neoformans in a murine model. mBio 3, e00103–12 (2012).
Bielska, E. & May, R. C. What makes Cryptococcus gattii a pathogen? FEMS Yeast Res. 16, fov106 (2016).
Liu, T.-B. et al. Brain inositol is a novel stimulator for promoting Cryptococcus penetration of the blood-brain barrier. PLoS Pathog. 9, e1003247 (2013).
Jung, K.-W. et al. Systematic functional profiling of transcription factor networks in Cryptococcus neoformans. Nat. Commun. 6, 6757 (2015).
Lee, K.-T. et al. Systematic functional analysis of kinases in the fungal pathogen Cryptococcus neoformans. Nat. Commun. 7, 12766 (2016).
Lee, K.-T. et al. Fungal kinases and transcription factors regulating brain infection in Cryptococcus neoformans. Nat. Commun. 11, 1–15 (2020).
Bott, K. et al. The effect of matrix characteristics on fibroblast proliferation in 3D gels. Biomaterials 31, 8454–8464 (2010).
Rao, R. R., Peterson, A. W., Ceccarelli, J., Putnam, A. J. & Stegemann, J. P. Matrix composition regulates three-dimensional network formation by endothelial cells and mesenchymal stem cells in collagen/fibrin materials. Angiogenesis 15, 253–264 (2012).
Levental, I., Georges, P. C. & Janmey, P. A. Soft biological materials and their impact on cell function. Soft Matter 3, 299–306 (2007).
Lu, Y.-B. et al. Viscoelastic properties of individual glial cells and neurons in the CNS. Proc. Natl Acad. Sci. USA 103, 17759–17764 (2006).
Bignami, A., Hosley, M. & Dahl, D. Hyaluronic acid and hyaluronic acid-binding proteins in brain extracellular matrix. Brain Struct. Funct. 188, 419–433 (1993).
Jin, Y. et al. Three-dimensional brain-like microenvironments facilitate the direct reprogramming of fibroblasts into therapeutic neurons. Nat. Biomed. Eng. 2, 522 (2018).
Cho, A.-N. et al. Aligned brain extracellular matrix promotes differentiation and myelination of human-induced pluripotent stem cell-derived oligodendrocytes. ACS Appl. Mater. Interfaces 11, 15344–15353 (2019).
Maeda, N. Proteoglycans and neuronal migration in the cerebral cortex during development and disease. Front. Neurosci. 9, 98 (2015).
Brakebusch, C. et al. Brevican-deficient mice display impaired hippocampal CA1 long-term potentiation but show no obvious deficits in learning and memory. Mol. Cell. Biol. 22, 7417–7427 (2002).
Margolis, R. K., Rauch, U., Maurel, P. & Margolis, R. U. Neurocan and phosphacan: two major nervous tissue-specific chondroitin sulfate proteoglycans. Perspect. Dev. Neurobiol. 3, 273–290 (1996).
Wurmser, A. E., Palmer, T. D. & Gage, F. H. Cellular interactions in the stem cell niche. Science 304, 1253–1255 (2004).
Wang, J. D., Khafagy, E.-S., Khanafer, K., Takayama, S. & ElSayed, M. E. Organization of endothelial cells, pericytes, and astrocytes into a 3D microfluidic in vitro model of the blood–brain barrier. Mol. Pharm. 13, 895–906 (2016).
Adriani, G., Ma, D., Pavesi, A., Kamm, R. D. & Goh, E. L. A 3D neurovascular microfluidic model consisting of neurons, astrocytes and cerebral endothelial cells as a blood–brain barrier. Lab Chip 17, 448–459 (2017).
Seo, J. H. et al. Oligodendrocyte precursors induce early blood-brain barrier opening after white matter injury. J. Clin. Invest. 123, 782–6 (2013).
Banerjee, S. & Bhat, M. A. Neuron-glial interactions in blood–brain barrier formation. Annu. Rev. Neurosci. 30, 235–258 (2007).
Dohgu, S. et al. Autocrine and paracrine up-regulation of blood–brain barrier function by plasminogen activator inhibitor-1. Microvasc. Res. 81, 103–107 (2011).
Shindo, A. et al. Astrocyte-derived pentraxin 3 supports blood–brain barrier integrity under acute phase of stroke. Stroke 47, 1094–1100 (2016).
Durieu‐Trautmann, O. et al. Nitric oxide and endothelin secretion by brain microvessel endothelial cells: regulation by cyclic nucleotides. J. Cell. Physiol. 155, 104–111 (1993).
Gadea, A., Schinelli, S. & Gallo, V. Endothelin-1 regulates astrocyte proliferation and reactive gliosis via a JNK/c-Jun signaling pathway. J. Neurosci. 28, 2394–2408 (2008).
Bauer, B., Hartz, A. M. & Miller, D. S. Tumor necrosis factor α and endothelin-1 increase P-glycoprotein expression and transport activity at the blood–brain barrier. Mol. Pharmacol. 71, 667–675 (2007).
Aplin, A. C., Zhu, W.-H., Fogel, E. & Nicosia, R. F. Vascular regression and survival are differentially regulated by MT1-MMP and TIMPs in the aortic ring model of angiogenesis. Am. J. Physiol. Cell Physiol. 297, C471–C480 (2009).
Lawler, J. The functions of thrombospondin-1 and-2. Curr. Opin. Cell Biol. 12, 634–640 (2000).
Gao, Q., Chen, K., Gao, L., Zheng, Y. & Yang, Y.-G. Thrombospondin-1 signaling through CD47 inhibits cell cycle progression and induces senescence in endothelial cells. Cell Death Dis. 7, e2368 (2016).
Cunningham, K. S. & Gotlieb, A. I. The role of shear stress in the pathogenesis of atherosclerosis. Lab. Investig. 85, 9 (2005).
Helmke, B. P., Goldman, R. D. & Davies, P. F. Rapid displacement of vimentin intermediate filaments in living endothelial cells exposed to flow. Circ. Res. 86, 745–752 (2000).
Lee, T. Y. J. & Gotlieb, A. I. Microfilaments and microtubules maintain endothelial integrity. Microsc. Res. Tech. 60, 115–127 (2003).
Akimoto, S., Mitsumata, M., Sasaguri, T. & Yoshida, Y. Laminar shear stress inhibits vascular endothelial cell proliferation by inducing cyclin-dependent kinase inhibitor p21Sdi1/Cip1/Waf1. Circ. Res. 86, 185–190 (2000).
Shao, J. et al. Integrated microfluidic chip for endothelial cells culture and analysis exposed to a pulsatile and oscillatory shear stress. Lab Chip 9, 3118–3125 (2009).
Stone, P. H. et al. Effect of endothelial shear stress on the progression of coronary artery disease, vascular remodeling, and in-stent restenosis in humans. Circulation 108, 438–444 (2003).
Pan, W. & Kastin, A. J. TNFα transport across the blood–brain barrier is abolished in receptor knockout mice. Exp. Neurol. 174, 193–200 (2002).
Brumble, L. et al. Fungal infections of the central nervous system: clinical, radiographic and laboratory manifestations. J. Microbiol. Exp. 5, 00167 (2017).
Shi, M. & Mody, C. H. Fungal infection in the brain: what we learned from intravital imaging. Front. Immunol. 7, 292 (2016).
Stie, J., Bruni, G. & Fox, D. Surface-associated plasminogen binding of Cryptococcus neoformans promotes extracellular matrix invasion. PLoS ONE 4, e5780 (2009).
Shi, M. et al. Real-time imaging of trapping and urease-dependent transmigration of Cryptococcus neoformans in mouse brain. J. Clin. Investig. 120, 1683–1693 (2010).
Charlier, C. et al. Evidence of a role for monocytes in dissemination and brain invasion by Cryptococcus neoformans. Infect. Immun. 77, 120–127 (2009).
Santiago-Tirado, F. H., Onken, M. D., Cooper, J. A., Klein, R. S. & Doering, T. L. Trojan horse transit contributes to blood–brain barrier crossing of a eukaryotic pathogen. mBio 8, e02183–02116 (2017).
Chaka, W. et al. Cytokine profiles in cerebrospinal fluid of human immunodeficiency virus—infected patients with cryptococcal meningitis: no leukocytosis despite high interleukin-8 levels. J. Infect. Dis. 176, 1633–1636 (1997).
Einsiedel, L., Gordon, D. L. & Dyer, J. R. Paradoxical inflammatory reaction during treatment of Cryptococcus neoformans var. gattii meningitis in an HIV-seronegative woman. Clin. Infect. Dis. 39, e78–e82 (2004).
Porte, R. et al. The long pentraxin PTX3 as a humoral innate immunity functional player and biomarker of infections and sepsis. Front. Immunol. 10, 794 (2019).
Martin-Manso, G. et al. Endogenous thrombospondin-1 regulates leukocyte recruitment and activation and accelerates death from systemic candidiasis. PLoS ONE 7, e48775 (2012).
Haris, M., Cai, K., Singh, A., Hariharan, H. & Reddy, R. In vivo mapping of brain myo-inositol. Neuroimage 54, 2079–2085 (2011).
Hicks, J. K., D’Souza, C. A., Cox, G. M. & Heitman, J. Cyclic AMP-dependent protein kinase catalytic subunits have divergent roles in virulence factor production in two varieties of the fungal pathogen Cryptococcus neoformans. Eukaryot. Cell 3, 14–26 (2004).
Ward, L. D. & Bussemaker, H. J. Predicting functional transcription factor binding through alignment-free and affinity-based analysis of orthologous promoter sequences. Bioinformatics 24, i165–i171 (2008).
Klingelhoefer, L. & Reichmann, H. Pathogenesis of Parkinson disease—the gut–brain axis and environmental factors. Nat. Rev. Neurol. 11, 625 (2015).
Ghaisas, S., Maher, J. & Kanthasamy, A. Gut microbiome in health and disease: Linking the microbiome–gut–brain axis and environmental factors in the pathogenesis of systemic and neurodegenerative diseases. Pharmacol. Ther. 158, 52–62 (2016).
Acknowledgements
This work was supported by grants (2017M3C7A1047659, 2016R1E1A1A01943365, 2021R1A2C3004262 and 2018R1A5A1025077) from the National Research Foundation of Korea funded by the Ministry of Science and ICT, Republic of Korea. This work was also supported by the Korea Evaluation Institute of Industrial Technology grant funded by the Korea government (MSIT) (no. 20009125) and the Institute for Basic Science (IBS-R026-D1), and the Strategic Initiative for Microbiomes in Agriculture and Food funded by Ministry of Agriculture, Food and Rural Affairs (916006-2). We thank J. Cheon (Yonsei University and IBS) for his support with facilities and equipment for analyses.
Author information
Authors and Affiliations
Contributions
J.K. and K.-T.L. contributed equally to this work. J.K. and K.-T.L. designed, led the experiments, performed data analysis and wrote the paper. J.K. led the overall work on human cell culture, analysis and microfluidic systems. K.-T.L. performed overall work on fungal experiments and transcriptome analysis. J.S.L. and K.Y. cultured NSCs, J.S. prepared and characterized hydrogels and B.C. fabricated microfluidic chips. Y.S.C. prepared and performed proteomic analysis of BEM. N.C. and S.H.L. provided materials and advised on the data analyses. J.-H.L. advised on the data analyses and wrote the manuscript. Y.-S.B. and S.-W.C. designed and supervised the experiments and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
J.K., K.-T.L., Y.-S.B. and S.-W.C. are co-inventors on patent applications (Korean Patent 10-2019-0115272, US patent 17/277,640 and EP patent 19 863 762.1) related to the hNVU chip developed in the manuscript.
Additional information
Peer review information Nature Biomedical Engineering thanks John Wikswo and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary methods, references, figures, notes,video captions, and tables.
Supplementary Video 1
Live imaging of the fungal infection model. The video clip of C. neoformans wild-type cells stacking and penetrating the BBB in the hNVU chip.
Supplementary Video 2
3D confocal image of the C. neoformans penetration site. The 3D image was processed by IMARIS (Bitplane; http://www.bitplane.com).
Rights and permissions
About this article
Cite this article
Kim, J., Lee, KT., Lee, J.S. et al. Fungal brain infection modelled in a human-neurovascular-unit-on-a-chip with a functional blood–brain barrier. Nat Biomed Eng 5, 830–846 (2021). https://doi.org/10.1038/s41551-021-00743-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-021-00743-8
This article is cited by
-
Organ-on-chip models for infectious disease research
Nature Microbiology (2024)
-
Vascularized organoid-on-a-chip: design, imaging, and analysis
Angiogenesis (2024)
-
Human disease models in drug development
Nature Reviews Bioengineering (2023)
-
Modelling host–microbiome interactions in organ-on-a-chip platforms
Nature Reviews Bioengineering (2023)
-
Design and engineering of organ-on-a-chip
Biomedical Engineering Letters (2023)