Abstract
Consumer wearable devices that continuously measure vital signs have been used to monitor the onset of infectious disease. Here, we show that data from consumer smartwatches can be used for the pre-symptomatic detection of coronavirus disease 2019 (COVID-19). We analysed physiological and activity data from 32 individuals infected with COVID-19, identified from a cohort of nearly 5,300 participants, and found that 26 of them (81%) had alterations in their heart rate, number of daily steps or time asleep. Of the 25 cases of COVID-19 with detected physiological alterations for which we had symptom information, 22 were detected before (or at) symptom onset, with four cases detected at least nine days earlier. Using retrospective smartwatch data, we show that 63% of the COVID-19 cases could have been detected before symptom onset in real time via a two-tiered warning system based on the occurrence of extreme elevations in resting heart rate relative to the individual baseline. Our findings suggest that activity tracking and health monitoring via consumer wearable devices may be used for the large-scale, real-time detection of respiratory infections, often pre-symptomatically.
Similar content being viewed by others
Main
Early detection of infectious disease is important to mitigate the spread of disease by increasing self-isolation and early treatments. Presently, most diagnostic methods involve sampling nasal fluids, saliva or blood, followed by nucleic acid-based tests for detecting active infections or blood-based serological detection for past infections. Although they are highly sensitive, nucleic acid-based diagnostics may require samples gathered several days post-exposure for unambiguous positive detection1. Moreover, they cannot be implemented routinely at low cost and are constrained by emerging shortages in key reagents.
Consumer wearable devices are an accurate and widely deployed technology to establish individual baseline parameters of health, which may be used to detect substantial deviations from baseline physiology at the onset of infection2,3,4. We have previously shown that smartwatches and simple pulse oximeters can be used for the early detection of Lyme disease, and in retrospective studies, heart rate and skin temperature can be used to detect viral respiratory infections, including asymptomatic infections5. Wearable sensors have also been used to detect atrial fibrillation6. Other recent studies have shown that elevated heart rate measurements from smartwatches can be used in epidemiological studies to track the spread of respiratory viruses7,8.
The use of wearable devices has ample potential to mitigate the coronavirus disease 2019 (COVID-19) pandemic. To date, the pandemic has infected tens of millions of individuals and caused over one million deaths worldwide (https://covid19.who.int). There is a substantial need for improved infection tracking, and population-scale technology solutions provide a promising avenue to identify cases in real time for infection detection and tracking9. Active infections are currently identified using PCR assays, which may require up to 3 d after infection for a reliable positive signal1. In addition, PCR tests are not widely used on a daily basis. Moreover, since most infections become apparent only upon symptom onset, the current methods of testing are unlikely to identify pre-symptomatic carriers, which is a considerable challenge for the implementation of early-stage interventions that reduce transmission. It is believed that as many as 50% of individuals with COVID-19 are asymptomatic, facilitating further viral spread10,11. As such, accessible and inexpensive methods for the early detection of COVID-19 in real time are urgently needed.
Smartwatches and other wearable devices are already used by tens of millions of people worldwide and measure many physiological parameters, such as heart rate, skin temperature and sleep12. Here, we investigate the use of wearable devices for the early detection of COVID-19 in a retrospective manner, and also present an approach for using wearable device-detected physiological parameters for real-time health monitoring and surveillance. Using heart rate and steps data from a large cohort of 5,262 individuals, we show that heart rate signals from fitness trackers can be used to retrospectively detect COVID-19 infection well in advance of symptom onset (offline detection). In addition, we developed an online detection algorithm to identify early stages of infection by real-time heart rate monitoring. We also examine the association between symptom type and severity, heart rate signals and the effect of infection on activity and sleep.
Results
Study design and overview
We investigated whether smartwatches could be used to detect COVID-19 at an early, pre-symptomatic stage. We enrolled a cohort of participants who had self-reported COVID-19 or other infections, as well as a wearable device capable of detecting heart rate, steps and other physiological measurements (Fig. 1). We then examined whether physiological deviations from baseline were detected around the period of illness, as well as the detection frequency and timing of onset of the event, and associations with symptoms. Finally, we used retrospectively collected data to develop an online method for potential real-time, early detection of illness onset.
A total of 5,262 participants were recruited, including individuals who were: (1) sick and tested positive for COVID-19 (dark red); (2) sick and tested positive for other illnesses (gold); (3) sick without a confirmed diagnosis (dark grey); and (4) not sick but were at high risk of exposure (light grey). Participants were asked to log daily symptoms and to share their fitness tracker data via the study app, MyPHD. The data types collected included heart rate, steps and sleep over a period of several months. Two infection detection algorithms were developed (RHR-Diff and HROS-AD). The bottom two panels represent derived heart rate metrics from the two algorithms over a period of months in one individual, centred around the onset of symptoms (day 0). The earliest detected abnormal heart rate elevations are marked by red stars. The anomaly periods detected by RHR-Diff are spanned by red arrows. The anomaly time points detected by HROS-AD are marked by red dots. The symptom onset day and diagnosis day are indicated by vertical dashed red and purple lines, respectively.
Under protocol number 55577, approved by the Stanford University Institutional Review Board, we enrolled 5,262 participants who completed surveys of illness, diagnosis and symptom dates, illness severity and symptom type (Figs. 1 and 2a and Supplementary Tables 1–5) through a secure REDCap system. Of these participants, 4,642 reported wearing a smartwatch: 3,325 wore Fitbits, 984 wore Apple watches, 428 wore a Garmin device and the remaining wore other devices (Supplementary Fig. 1). Of these, 114 individuals reported COVID-19 illness with symptom and diagnosis dates, and another 47 individuals reported a different respiratory infection with symptom and diagnosis dates for an identified pathogen. We were not able to acquire wearable device data near the symptom date from many of these. Since most people wore Fitbits, we focused on this group. Thirty-two COVID-19-positive participants (27 confirmed; see Methods) had Fitbit data spanning and adjacent to the COVID-19 disease dates, as well as symptom dates and diagnosis dates. Four of these individuals with Fitbit devices lacked either a reported symptom date or a diagnosis date. Of note, at least five participants in our study had wearable device data but lacked measurements at or shortly after the time of infection, suggesting that some participants do not habitually wear their devices when ill (for example, participant AV2GF3B in Supplementary Fig. 3a).
a, Summary of data collected from 32 study participants who reported a confirmed diagnosis of COVID-19 with a symptom onset and/or test date. Each row along the y axis represents one participant, with prediction groups labelled to the left. The plot shows sick periods for COVID-19 (between black arrows, with dashed lines for unknown bounds where the symptom onset or recovery day was unclear), COVID-19 test dates (red crosses), sick periods for other illnesses (between orange arrows) and the days for which participants filled in the daily survey (diamonds), with purple diamonds representing days when symptoms were reported and blue diamonds representing days when symptoms were not reported. b, Overlapping bar plots depicting heart rate metrics and timings of infection detection from RHR-Diff and HROS-AD with respect to the infection detection window for COVID-19-positive participants. The plots are manually grouped into three groups: group I (single region; blue); group II (early and multiple regions; red); and group III (other; gold), focusing mainly on the single-region group. One additional participant who reported a diagnosis of influenza B is also shown for comparison (purple). Summary plots for all COVID-19-positive participants from all three groups are shown in Supplementary Fig. 2a, and all participants with other illnesses are shown in Supplementary Fig. 2b. The x axis shows days during the infection detection window and the y axis shows standardized residual values from RHR-Diff (brown bars) and standardized (0 to −1) HROS values from HROS-AD (transparent green bars) in the intervals during which COVID-19 infection was detected by each algorithm. These values are plotted separately for each participant. This window is a period of time centred around symptom onset at day 0 (substituted by the diagnosis day wherever the day of symptom onset was unavailable). The infection detection window spans a period of 15 d before day 0 and 7 d after day 0.
We also analysed data from two classes of control individuals with Fitbit data: (1) 15 individuals with confirmed illness that was not due to severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) for whom wearable device data were available (see Methods) (one case was associated with influenza B, another was associated with rhinovirus and the remainder were of unknown cause; Supplementary Tables 3 and 4); and (2) 73 healthy individuals who did not report any illness or symptoms during the same period when we collected data from the COVID-19-positive individuals.
Abnormal resting heart rate (RHR) and heart rate-to-steps ratio are associated with COVID-19 illness
First, we determined whether abnormal physiological events are associated with SARS-CoV-2 infection and whether these can be detected using a smartwatch at or near the time of infection. Several parameters were investigated: elevated RHR relative to a previous healthy window; an increased heart rate relative to number of steps (that is, the RHR-to-steps ratio); and sleep (see Methods). We focused primarily on early-onset events since participants often take medications or undergo other treatments once symptomatic.
We developed two methods for detecting aberrant physiology. (1) Using the RHR difference (RHR-Diff) method, we detected and identified elevated RHR time intervals based on the standardized residuals (see Methods). The standardized residuals were constructed at 1-h resolution by comparing each interval with the average daily curve using a 28-d sliding window. We applied a non-parametric approach13 to test whether the sequence of standardized residuals was a homogeneous process. Using a significance level of 0.05, the elevated regions were reported as abnormal RHR periods (Supplementary Table 6). (2) Using the heart rate over steps anomaly detection (HROS-AD) method, we created a new feature known as heart rate over steps (HROS) by dividing heart rate by steps data and comparing the HROS value at each hourly interval with the rest of the intervals using Gaussian density estimation14,15. We smoothed and standardized the HROS value (see Methods). Using Gaussian density estimation, we computed an anomaly score for each observation and classified them as normal (1) or an anomaly (−1) with an outlier threshold of 0.1 (see Methods and Supplementary Table 7). Analyses were performed on all of the data available for each individual, including data collected before, during and after the reported illness. A method similar to HROS-AD, the RHR anomaly detector (RHR-AD; see Methods) produced similar results (Supplementary Table 8).
Using dates of symptom onset and diagnosis to define sick periods, we then defined a sickness detection window for each individual based on the symptom onset date wherever available (14 d before to 7 d after), and the diagnosis date when the symptom date was not available (two cases). The timeframe of 14 d was chosen since this has been suggested to cover the duration of the COVID-19 incubation period in most cases16,17. We scored both of our detection methods based on the interval (RHR-Diff) or HROS-AD period that overlapped with the sickness detection window. In all of the 26 individuals detected (100%; groups I and II; see below for details), we found outlying periods near the time of infection using either RHR-Diff or HROS-AD, with 22 identified using both methods (Supplementary Tables 9–12). RHR-Diff detected two high-signal regions not identified by HROS-AD, and HROS-AD detected one high-signal region not identified by RHR-Diff. Interestingly, we observed that neither method detected stable signal regions specifically at COVID-19-infected regions in six individuals, as described below.
The 32 individuals fell into three types of patterns (Supplementary Fig. 2a and Fig. 2b). Group I included 16 individuals for whom we were able to detect disease primarily as a single elevated period or a tight cluster of elevated periods before or overlapping with the disease period. Two examples are shown in Fig. 3a,b in which we detected elevated heart rates starting 15 and 4 d before symptom onset, respectively. In cases where a tight cluster of elevated periods was observed, it is possible that the normal physiological periods reflect times of self-medication or, less likely, disease remission.
a–d, Examples of heart rate metrics during COVID-19 infection for four individual participants, two from group I (a and b) and one each from group II (c) and group III (d). Red and purple vertical dashed lines indicate the days of symptom onset and diagnosis, respectively. Shown are the standardized HROS from the HROS-AD method (bottom plot in each panel; dark blue lines) and the standardized heart rate residuals from the RHR-Diff method (top plot in each panel; black lines). For RHR-Diff, the green dashed line is at 0. Gold solid triangles mark the infection detection window used to score detections as a hit or a miss. Also indicated are time intervals when the heart rate residuals were significantly elevated from RHR-Diff (red arrows in the top plots of each panel) and times when anomalies were detected by HROS-AD (red dots in the bottom plots of each panel).
Group II comprised ten individuals and formed a cluster where we were able to detect a symptom-associated peak as well as an earlier significant heart rate elevation period within 28 d of the symptom onset based on RHR-Diff (−21/+7 d). An example is shown in Fig. 3c. In some cases, this affected our ability to clearly differentiate physiological changes associated with the COVID-19 infection since it merged with the earlier elevations in heart rate. Three of these individuals had a self-reported stress period (either illness or other), raising the possibility that the stress-associated event may have contributed to COVID-19 illness onset.
Group III consisted of the six individuals for whom a single stable elevated period could not be easily discerned at or before symptom onset; these individuals often had many signals distributed across a substantial period of time (Fig. 3d is an example) or no significant signal. Interestingly, two of these individuals had respiratory lung disease and another had severe allergies, and it is likely that these conditions and/or the pharmacological therapies used to treat them interfered with outlier detection. However, not all individuals with respiratory conditions are missed using a Fitbit; three other individuals with these conditions gave high RHR-Diff (Supplementary Fig. 3a) and HROS-AD (Supplementary Fig. 3b) signals associated with illness.
For groups I and II, the number of days between the beginning of the aberrant signal and the date of symptom onset (when available), as well as the date of diagnosis, were distributed as shown in Fig. 4a,b (Supplementary Table 13). Of the 26 detected cases, 24 had both a symptom onset date and a diagnosis date, and the remaining two cases had either one, but not both dates. In total, 88% (22 out of 25) and 100% (25 out of 25) of individuals with a symptom onset date (n = 25 cases) or a diagnosis date (n = 25 cases) showed elevated signals in advance or at the time of symptom onset or diagnosis, respectively. Signals were detected several days ahead of symptom onset and diagnosis, with median values of 4 and 7 d in advance, respectively. The median heart rate increase in the first period of onset was 7 beats per minute, with a broad range (Fig. 4d and Supplementary Table 14). Overall, these results indicate that altered physiology is associated with COVID-19 illness, often in advance of symptoms, and that this can be detected with a wearable device.
a,b, Histograms summarizing the distribution of early-detected COVID-19 events compared with the first day of self-reported symptoms (a) and the reported diagnosis day (b). If multiple RHR-Diff intervals existed within or intersecting the COVID-19 event (−14/+7 d versus the symptom day), the first day of the closest interval ahead of this event was used. If no interval before the event was observed, the closest interval after the event was used. If the symptom day was not available or there was no interval detected within the 21 d surrounding the symptom onset, the closest interval within 28 d of the diagnosis onset (−21/+7 d) was used. The colours represent the number of individuals in each group (group I: blue; group II: red; group III: gold). The purple line shows the kernel probability density estimate. Also shown are individuals for whom the algorithm missed detecting COVID-19 infection (separated from the quantitative part of the graph by grey dashed lines). c, As in a, but for participants with other illnesses. d, Boxplots summarizing the hourly ΔRHR of the detected COVID-19 or other illness interval compared with the baseline RHR of the same individual. These boxplots exclude individuals for whom the RHR-Diff algorithm missed detecting infection. Central lines represent median values, box limits represent upper (third) and lower (first) quartiles, whiskers represent 1.5× the interquartile range above and below the upper and lower quartiles, respectively, and red crosses represent outliers. The number above each boxplot represents the median value of ΔRHR for the indicated individual.
To determine whether the increased RHR signal was specific for COVID-19, we also analysed the 15 cases where individuals reported non-COVID-19 illness. For 14 cases in which the first symptom was reported, increased RHR was evident near symptom onset in nine instances; moreover, in these nine cases, increased RHR was apparent before or at the time of symptom onset (Supplementary Figs. 2b and 3a,b and Supplementary Tables 6 and 15–17). One example for an influenza B infection is shown in Fig. 2b. The median time of signal onset relative to symptoms was 2 d (Fig. 4c). These results indicate that the elevated heart rates that occur before disease also provide utility as a general signal of respiratory illness, consistent with our previous results5.
Sleep and activity alterations associated with COVID-19 illness
Having established aberrant physiological signals associated with COVID-19 illness, we investigated whether COVID-19 also affected behaviour, specifically steps and sleep duration (Fig. 5a–d and Supplementary Tables 18–23; see Methods). Although Fitbit devices are not considered gold standards for many sleep parameters, they most accurately measure sleep duration and are widely used18,19. They are less accurate for the sleep stages (for example, rapid eye movement (REM) and deep sleep) and this was not pursued. We examined the parameters reported by the manufacturer (see Methods) and found that steps (with missing data imputation and without) significantly decreased at the onset of the outlying RHR-Diff signal associated with COVID-19 illness (linear mixed model (LMM); P = 8.71 × 10−33; Fig. 5a,b and Supplementary Fig. 4a). Sleep duration significantly increased after the onset of the outlying RHR-Diff signal, but only with missing data imputation (LMM; P = 0.003; Fig. 5c,d and Supplementary Fig. 4b). These results indicate that COVID-19 illness alters steps and sleep patterns, which can be tracked using a wearable device.
a, Heatmap showing standardized daily steps per participant (that is, z scores of daily steps) for the 22 participants we have steps data for, and for whom RHR-Diff detected a change of RHR any day between 14 d before the symptom onset date and 2 d after. Tile colours indicate the z score and asterisks represent the first day of detection for each participant. b, Boxplot showing the change in daily steps between days before and after the detection start date in a window of −21 d before to +7 d after the symptom onset date. c, Heatmap showing the standardized sleep duration per participant (that is, z scores of total sleep duration) for the 13 participants we have sleep data for and for whom RHR-Diff detected a change of RHR any day between 14 d before the symptom onset date and 2 d after. d, Boxplot showing the change in total sleep duration between days before and after the detection start date in a window of −21 d before to +7 d after the date of symptom onset. For the heatmaps in a and c, black rectangles highlight the period after symptom onset. The boxplots in b and d include data with imputation (see Methods). Data without imputation are shown in Supplementary Figs. 4a,b. For both b and d, central lines represent median values, box limits represent upper (third) and lower (first) quartiles and whiskers represent 1.5× the interquartile range above and below the upper and lower quartiles, respectively.
Association between heart rate signals and symptoms
A subset of participants filled out daily logs before or during their COVID-19 illness, providing a detailed time course of symptom severity, progression and relapse (Fig. 6a–d), while others filled out detailed past illness surveys that summarized their symptoms over the entire illness period (Fig. 6e). The first individual, APGIB2T, (Fig. 6a) had an early RHR-Diff signal 1 week before symptom onset. The disease progressed quickly into severe diarrhoea, fatigue, headaches, elevated temperature and positive COVID-19, peaking in severity and then declining over 2 weeks. In total, 18 d of initial illness were followed by 12 d when the participant felt recovered, before a relapse characterized by elevated temperature, fatigue, diarrhoea and elevated heart metric signals. A second participant, AQC0L71, (Fig. 6b) began daily logs when symptoms developed, reporting a 22-d period of mild-to-moderate coughing, fatigue and aches and pains that was anticipated by both the RHR-Diff and HROS-AD heart rate metrics. Symptoms then deteriorated rapidly, coupled with abnormal physiological signals suggested by both algorithms, elevated temperature and a positive COVID-19 test. The participant was admitted to hospital 5 days later and the time from symptom onset to recovery was 41 d. A third participant, A0VFT1N, reported COVID-19 illness lasting 13 d (Fig. 6c) that was led by an RHR-Diff alarm, followed by ongoing symptoms of fatigue and occasional chest pains. Heart rate metrics alarms accompanied the return of shortness of breath, for which the participant was hospitalized 35 d after initial symptom onset. Daily logs began at symptom onset for A1K5DRI, the fourth participant (Fig. 6d), 3 d after an RHR-Diff alarm. Illness progressed over 23 d, with a rapid rise in temperature and HROS-AD alarms accompanied by severe fatigue, aches and pains and slow recovery.
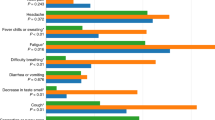
a–d, Plots of four individual participants (APGIB2T (a), AQC0L71 (b), A0VFT1N (c) and A1K5DRI (d)) over the course of COVID-19 infection. Vertical columns along the x axes each represent a single day of symptoms (from early illness (leftmost) to late illness) and are aligned with the heart rate metrics below. Columns showing symptoms are only present for the days when the daily survey was completed, while heart rate metrics progress continually below. ‘Overall feeling’ indicates how the participants reported feeling on a particular day, with a bar plot above indicating the measured temperature if reported, and specific symptoms highlighted below as a heatmap depicting the severity. Black vertical lines below the symptoms and descending into the heart rate metrics are labelled to highlight significant days during the illness course, and align with the symptoms above. The RHR-Diff plots show standardized heart rate residuals from RHR-Diff (black lines) and time intervals when the heart rate residuals were significantly elevated (red lines with arrowheads). The bottom plots in each panel show standardized HROS using the HROS-AD method (black line), and each detected anomaly is indicated by a red oval. e, Summary of symptoms data for individuals who provided surveys on a past COVID-19 illness. Each column represents a study participant, as labelled below. Shown are (from top to bottom): a bar plot of average temperatures in °C reported during illness; overall feelings (see legend above); total duration between reported symptom onset and recovery (if provided); a boxplot (showing numerical median values) of ΔRHR in beats per minute when heart rate residual alarms were raised; and a plot of individual symptoms (where black boxes indicate reported symptoms and white boxes indicate no reported symptoms). For the boxplot, central lines represent median values, box limits represent the upper (third) and lower (first) quartiles, the whiskers represent 1.5× the interquartile range above and below the upper and lower quartiles, respectively, and red crosses are outliers.
Lastly, in addition to the daily survey, we also examined symptoms reported post-illness (that is, retrospectively). In this limited sample, we did not detect any obvious association between the magnitude of RHR differences during alarm periods and symptom type or number, illness length or temperature (Fig. 6e). Overall, at the individual level, COVID-19 progression and severity were generally concordant with heart rate metrics, but these cases highlight temporal and individual variation more widely observed with the illness11,20.
An approach to detect early COVID-19 onset in real time
The ability to detect altered physiology in advance of symptoms raises the possibility that an online method can be developed to detect early stages of COVID-19 illness in advance of symptoms using a smartwatch. To test this possibility, we developed an online detection method called CuSum (see Methods). This detection was based on cumulative statistics21,22,23 that cumulate the deviations of the elevated residual RHRs. The test statistics from the previous 28 d of baseline records built an empirical null distribution.
We report a warning alarm as the first time we observed a test statistic more extreme compared with the null distribution, with a P value generated from comparing the current test statistics with the baseline measurements. To reduce the number of alarms, a two-tiered warning system was developed. The first time the P value was less than 0.01 (usually in the first few hours), an initial warning alarm (yellow alert) signalled. Monitoring continued, and if it remained elevated over 24 h, it signalled a positive event (red alert; see Methods).
We tested this method initially on four individuals for whom we had collected >6 months of wearable device data (Fig. 7a,b, Supplementary Fig. 5 and Supplementary Table 24). In addition to the annotated COVID-19 infection, other strong elevated signals were identified, as well as smaller signals. Some of these corresponded to annotated infections. Others were not annotated but occurred at periods that might be associated with increased heart rate. For example, three of the four individuals had high heart rates in the November to December holiday period (‘holiday bump’, Fig. 7b and Supplementary Fig. 5), which is commonly associated with air travel, alcohol and stress, as well as illness. A number of alarms of lower duration or signal were also observed.
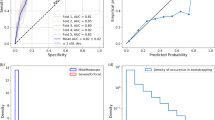
a–d, Examples of online prediction performance during COVID-19 infection for two participants with long-term data (a and b), one example of other (non-COVID-19) illness (c) and one example from the healthy group (d; note the smaller scale compared with a–c). For each plot, the x axis is the number of days pre- or post-symptom onset. The red and purple vertical dashed lines indicate days of symptom onset and diagnosis, respectively, and the blue dashed vertical lines indicate the alarming time from online detection (see Methods). e, Alarm counts per 30 d for participants with COVID-19. The blue and red bars indicate the alarm counts before and after the COVID-19 event. Average alarm counts are 0.29 versus 1.35 before and after infection, respectively. f, Early detection comparison between offline detection (RHR-Diff) and online detection (CuSum). Detection days are compared with the symptom day. Each red circle indicates one participant. The black dashed line is the identity line and the blue dashed lines surrounding it are at a distance of ±1 d from the identity line. The grey dotted lines separate the quantitative part of the graph from the missed cases. g, Comparison of the total alarm duration across the COVID-19 positive group, other illness group and potentially healthy group. h, Comparison of the alarm peak height across the different groups described in g. For each sickness case in g and h, the alarms are further assigned to three categories: pre-sickness, during sickness and post-sickness. Only P values with a significance of <0.05 are shown. In addition, a slight increase in alarm frequency was observed, but it did not achieve significance (Supplementary Fig. 7 and Supplementary Table 29). For the boxplots in g and h, central lines represent median values, box limits represent the upper (third) and lower (first) quartiles, whiskers represent 1.5× the interquartile range above and below the upper and lower quartiles, respectively and black dots represent outliers.
We also examined 24 individuals who had at least 28 d of data ahead of symptom onset (Supplementary Fig. 3a and Supplementary Table 25). In total, 62.5% (15/24) had an alarm on or before COVID-19 symptom onset using the CuSum alarm model. Four more people had an alarm 1 d after self-reported symptoms. Of the remainder, two individuals had previously been missed in offline detection and three had respiratory illnesses that had been difficult to detect in our initial retrospective study. As shown in Fig. 7e, 11 individuals had non-COVID-19 alarms before COVID-19 infections ranging from 0.14 to 1 alarms per month. Interestingly, the number of alarms increased considerably post-COVID-19 infection, suggesting the possibility of lingering physiological sequelae of COVID-19 illness (Fig. 7e and Supplementary Table 26).
We compared our online detection results with those observed using the RHR-Diff approach and found overall good agreement (either less than 1 d difference or both missed) in 13 of the cases (Fig. 7f and Supplementary Table 27). However, one case was only detected using the online CuSum approach and another was only detected by the RHR-Diff approach (this individual had a pre-existing chronic respiratory condition. Nine cases were detected 2–6 d later than offline (see Supplementary Fig. 3a for examples). RHR-Diff is more sensitive since it detects significant intervals based on the global dataset, whereas the detection by CuSum was solely based on the data received in advance of infection.
As controls, we used the CuSum online detection method to examine wearable device data for: (1) the 73 individuals who did not report illness during the same period as the COVID-19-positive individuals; and (2) the 13 individuals with 15 non-COVID-19 illnesses (Supplementary Table 25; examples in Fig. 7c,d). The healthy individuals also had alarms (Fig. 7d and Supplementary Fig. 6), although the alarm durations and peaks were generally shorter and smaller, respectively, than those of the COVID-19 and other illnesses (Fig. 7g,h and Supplementary Table 28). There were also signals that were similar to those for infections, possibly representing asymptomatic illnesses. The holiday bump was also observed in one of three other individuals whose data covered that period. The 15 individuals with other illnesses gave a signal at or before the illness in nine of 15 cases (Supplementary Fig. 3a). The presence of alarms was expected during healthy periods for all individuals, since the alarming method was set to identify signals that lay near the end of the normal distribution; these will have occurred by statistical chance, as well as by triggers other than the identified illness.
Discussion
From a sizable cohort, we identified a number of individuals who tested positive for COVID-19 and other illnesses and who wore a smartwatch. Using these data, we developed algorithms that detected elevated RHRs and outlying heart rate/steps measurements, usually in advance of symptoms. The early times of detection were generally consistent with the latent period of pre-symptomatic illness reported previously10. In two group I individuals, the signal was observed nine or more days preceding symptoms. Because the actual timing of infection in these cases was not known, it is possible that these and other events represented early stress events that merged into the COVID-19 illness (for example, Fig. 6b). Indeed, in ten group II individuals, a discrete early event was observed, and in three individuals, this was associated with a self-reported illness or family stress event. It is possible that early stress events increased individuals’ vulnerability to COVID-19, resulting in illness.
We used the information learned from the retrospective analysis to design a prototype approach for the real-time, early detection of COVID-19 illness (CuSum). In addition to detection of the COVID-19 events, other events were identified, of which some probably reflect illnesses, including asymptomatic cases, as we observed previously5. Many of the other events could reflect situations that stimulate sustained increased heart rate, such as medication, alcohol, travel and emotional or other stress inducers. Indeed, four of seven cases with data covering the December holidays showed significant elevations of long duration. Those events short in duration (for example, due to watching a scary movie) will probably go off after a brief period of time. Thus, using our proposed two-tiered continuous alarm system, early events can be acted on by self-isolation and, if an increased signal ensues, can be escalated to physician consultation and/or direct viral diagnostics. The alarming parameters can also be adjusted to increase or decrease sensitivity with a concomitant increase or decrease in the number of alarms. This adjustment may be valuable depending on the person’s preference or risk. In the version presented here, we were able to detect 63% of known COVID-19 infections with an alarming frequency of 0.66 per month in the healthy individuals; 63% is likely to be an overestimate for COVID-19 as asymptomatic cases are not accounted for, but an underestimate for all infections as many signals may represent such illnesses (both unreported symptomatic and asymptomatic).
It should be noted that the wearable devices used in our study have not yet been approved by the US Food and Drug Administration for early illness detection and our study is still modest in size. Another limitation we observed is that some individuals do not wear their devices (or let their charge expire) when symptomatic, which may affect monitoring patterns. Patterns of non-use were not Fitbit specific, and we expect that devices requiring daily charging will also have more missing data. Nonetheless, devices whose charge lasts for several days should be powerful enough for early detection before loss of device function.
Our approach is a general detection method and presently cannot distinguish infections with SARS-COV-2 from those caused by other viruses (other than pre-symptomatic duration), since increased RHR is common to many respiratory infections. Regardless, any illness onset information is valuable, especially during a pandemic, and can be followed up with appropriate testing. It is also likely that other types of physiological measurements that are obtainable from wearable devices (for example, heart rate variability, respiration rate, skin temperature, blood oxygen saturation and electrocardiogram readings) will be valuable for distinguishing illnesses caused by different infectious agents and could be used to increase diagnostic sensitivity and perhaps even predict illness severity and symptoms24,25,26,27. Data on reported respiratory rates and blood oxygen saturation are expected to be particularly useful in COVID-19 prediction28, although the disease is quite heterogeneous in its physiological presentation29,30, as observed in our study. At the time of writing, such data were not available to us; however, these data, especially when combined with machine learning approaches, as well as an increased number of study participants, will greatly improve diagnostics. Regardless, this continuous monitoring approach is expected to be powerful for early infectious illness detection and offers many advantages that may help increase disease detection during the current global pandemic. Specifically, wearable device-based disease detection does not require testing infrastructure, materials or personnel that can be overburdened by global supply chain shortages. In addition, real-time monitoring by smartwatches is a passive form of testing that does not burden patient schedules and can serve as a high-resolution continuous screening to inform follow-up testing and self-isolation. We hope that ongoing screening for COVID-19 risk using wearable devices can provide a scalable solution to help overcome current barriers with testing, and inform early diagnosis and treatment to mitigate the spread of the disease. Such information will inform patients for self-isolation, diagnosis confirmation and early treatment.
Methods
Participant recruitment
We recruited 5,262 adult individuals for this study under protocol number 55577 approved by the Stanford University Institutional Review Board. Participants were recruited using REDCap and informed electronic consent was obtained from all participants31. Recruitment was done through social media, word of mouth, COVID-19 registries and presentations, as well as via referrals from Stanford Health Care. We recruited participants with a confirmed or suspected COVID-19 infection, as well as those at high risk of exposure to COVID-19 (for example, via family members or relevant occupation), individuals with unknown respiratory illness and individuals who did not report any illness. Participants were asked to wear their fitness tracker daily, as much as possible, and to download a study app called MyPHD (see ‘MyPHD app for wearable device data collection’ below) with which to share their wearable device data. In addition, long-term wearable device data collected during periods before the COVID-19 pandemic (2019 or before) from seven individuals enrolled in our iPOP study32 were also extracted and analysed. The iPOP study was approved by the Stanford University Institutional Review Board under protocol number 23602.
Metadata collection and surveys
Study metadata, such as demographic information, reports of past illnesses, daily symptom tracking and so on, were collected via REDCap. At enrollment, participants were asked to provide: (1) demographic information, such as age, sex, ethnicity, height and weight; (2) medical history, including chronic illnesses, routinely taken medications and so on; and (3) COVID-19 illness status (that is, whether they had a confirmed or suspected COVID-19 infection and, if tested, the test date, results and symptom onset date).
In addition, all participants were asked to complete a daily symptom tracking survey, which tracked the symptoms experienced and their severity on a scale of 1–5 (mild, mild to moderate, moderate, severe or worst possible), body temperature (if recorded), new tests or diagnoses of COVID-19 or other respiratory illnesses, test results, recovery dates and so on. Finally, participants were also asked to fill out a one-time past illness survey, where they could report past sickness periods (up to five illnesses in total) since 1 November 2019. The past illnesses survey recorded the length of the sickness period and other elements similar to the daily survey: diagnoses (if any) of COVID-19 or other respiratory illnesses, any symptoms they reported experiencing during this period, as well as body temperature and symptom severity on a scale of 1–5.
For this study, we restricted our analyses to a dataset of 32 individuals who reported a positive COVID-19 diagnosis, a diagnosis date and/or symptom onset date (usually both; n = 28) and wearable device data appropriate for the analyses. Five of these individuals also reported other non-COVID-19 respiratory infections, including four individuals who reported two other illnesses since October 2019. Nearly all (n = 27) provided diagnosis confirmation: 23 of the participants provided written documentation of their test result and four others provided verbal confirmation. We also analysed data for 15 non-COVID-19 illness events from 13 other participants; one was diagnosed with influenza B, another with a rhinovirus infection and four with non-COVID-19 infections (type unknown). Long-term (>1 year) data during periods before the COVID-19 pandemic (2019 or before) from seven additional participants from the iPOP study with a total of nine infections were also analysed. Illness was confirmed for six of these events by elevated C-reactive protein levels (as determined by high-sensitivity C-reactive protein test; Supplementary Table 30). Data from February 2020 until June 2020 were also analysed from 73 healthy participants who did not report any illness.
MyPHD app for wearable device data collection
After participants enrolled on REDCap, they were directed to download MyPHD, a smartphone app developed by our study team, to collect their wearable device data in a de-identified and encrypted manner. The MyPHD app was made available to study participants for both Android and iOS platforms. For Fitbit watches, the data were accessed through the Fitbit application programming interfaces, and for wearable devices with Apple HealthKit integration, we obtained the data via HealthKit. Data transfer was done from the source to a Health Insurance Portability and Accountability Act-compliant Google Cloud Platform project in an encrypted form, then the data were decrypted for pre-processing and analysis in a controlled-access, secure environment.
Wearable devices and data types collected
Participants wore Fitbit smartwatches, including different versions, such as Fitbit Ionic, Charge 4 and Charge 3. The data types collected included heart rate, steps and sleep. Raw heart rate, steps and sleep data were collected in JavaScript Object Notation format. Heart rate data were retrieved at 15-s resolution, steps values at a resolution of 1 min and sleep data as sleep stage intervals (wake, light, deep and REM).
Wearable device data pre-processing
The retrieved raw heart rate, sleep and steps data from Fitbit were processed and integrated using a systematic workflow to produce a uniform format among different retrieval protocols. First, heart rate outliers (heart rate > 200 and heart rate < 30) were removed, as were all duplicates in the heart rate, steps and sleep data. Time stamps were unified to a standard time zone to be able to match different types of wearable device data with metadata. Heart rate features were extracted, such as median heart rate per minute, average heart rate per minute, night-time RHR and so on. Additionally, daily steps were calculated. For sleep features, total sleep duration per night, as well as wake, light, deep and REM stage durations and their corresponding percentages for each night, were calculated.
Symptoms and other metadata processing
Participant metadata and symptom surveys were downloaded and processed using a custom R and Python script. A total of 136 participants reported a positive COVID-19 diagnosis, but many were lacking a clear diagnosis or symptom date or appropriate wearable device data for the analyses. Height was converted to centimetres (Supplementary Tables 1 and 2), weight was converted to kilograms (Supplementary Tables 1 and 2) and reported temperature was converted to Celsius (Fig. 6) for all participants.
RHR-Diff offline anomaly detection
The RHRs were obtained using the same approach as in Li et al.5. For each person, the RHRs were then standardized in 1-h resolution based on the average of daily curves from a 28-d sliding window. The missing values in the RHRs were imputed as zeroes before the detection. We applied anomaly time-interval detection based on rank scans from the work of Arias-Castro et al.13 on the standardized residuals. Under a significance level of 0.05, the detected elevated time intervals were reported. To reduce possible false positives, short detected intervals of <24 h were removed. If there was a gap of two days or less near the symptoms onset date, it was treated as a single signal.
HROS-AD offline anomaly detection
HROS-AD is an unsupervised anomaly detection model consisting of two major steps:
-
1.
In the data pre-processing step, we combined heart rate and step data from each user to compute a new feature known as HROS. HROSi is a feature of a user i’s HROS (a value of 1 is added to all steps to avoid the zero-division problem). Next, we used moving averages (mean = 400 h) and down-sampling (mean = 1 h) to smoothen the time-series data and standardized further with a Z score transformation.
-
2.
In the anomaly detection step, when a HROS data point deviated markedly from others in a sample, it was called an anomaly or outlier. Any other expected observation was labelled as an inlier.
We used the covariance.EllipticEnvelope class from the scikit-learn package in Python14,15,33 to fit a Gaussian distribution of the data, pointing out the anomalies that might be contaminating our dataset because they are extreme points in the general distribution of the dataset. For simplicity, we call this method HROS-AD when the input data are HROS. Within the HROS-AD method, EllipticEnvelope is a function that calculates the distance of each HROS observation with respect to the grand mean that takes into account all of the observations in the data and detects both univariate and multivariate outliers.
HROS-AD uses a key parameter called contamination that provides information about the proportion of the HROS outliers present in each dataset and can take a value up to 0.5 (Supplementary Table 31). We start with a value of 0.01 because 0.01 is the percentage of observations that should fall over the absolute value 3 in the Z score distance from the mean in a standardized Gaussian distribution. If we do not detect any anomalies, we gradually increase the contamination value from 0.01 until we find an anomaly. If we find too many anomalies with a 0.01 contamination score, we gradually decrease the contamination value. The predictions contain a vector of values between 1 and −1 (1 being normal and −1 being anomalous).
We deleted the predictions if they were overlapping daytime (6:00–00:00) or missing steps in the alert window of 21 d before symptom onset and 7 d post-symptom onset. There were five participants who had missing step data for at least one day in the alert window, and three of the participants had at least one prediction overlapping daytime or missing steps in the alert window.
We also used resting heart rate (RHR) instead of HROS to check the model performance. We call this method RHR-AD. It uses the same pipeline as HROS-AD except the input is RHR, which is the heart rate at a given time point where the step count for the previous 12 min was 0. Overall, the results between HROS-AD and RHR-AD were very similar (Supplementary Tables 7, 8, and 15).
Activity and sleep analysis
In our analysis, we only considered individuals who had detectable changes in RHR between −14 d before symptom onset and 2 d after, using our RHR-Diff algorithm. We also removed individuals with more than 50% of steps or sleep (each individually) data missing in a window of 21 d before symptom onset and 7 d after. This resulted in 22 individuals for the steps analysis, and 13 individuals for the sleep analysis. Following this filtration criteria, missing values were imputed using the last observation carried forward (LOCF) method. Afterward, daily steps and total sleep duration were Z score normalized for each person independently.
Since wearable device data tend to have missing values (especially sleep, since some participants do not wear the watch every night), we evaluated the change in daily steps and total sleep duration without imputing the missing values. In a separate analysis (Supplementary Fig. 4), we compared daily steps and total sleep pre- and post-detection without the imputation process. We only considered 7 d pre-detection and 7 d post-detection.
LMMs were conducted for daily steps and total sleep duration using the nlme package (version 3.1-142) in R. In our model, we included day annotation as a fixed effect and subject ID as a random effect. An analysis of variance test was applied on the fitted model to retrieve a P value for the tested hypothesis.
CuSum online detection
CuSum statistics were calculated based on the work of Levin and Kline13,23. The values of CuSum statistics in the previous baseline days were used to construct a null distribution for each hour, and a sliding 1 h interval was then interrogated for residuals compared with the baseline distribution for that hour. The baseline window was set to 28 d. The threshold parameter in the CuSum statistic was set as half of the 90% quantile of the baseline residuals for the short-term data and half of the 99% quantile for the long-term data. Under the significance level 0.01, an alarm candidate was recorded the first time that the CuSum statistic was significantly higher than the values from the null distribution. We tracked the records of CuSum statistics for 48 h. To reduce possible false positives, we started monitoring the statistic when it rose above the threshold in the second hour. In cases where the CuSum statistic stopped increasing within 24 h or returned to zero within 48 h, the initial alarm was removed.
Visualization methods
We used ggplot2, Matplotlib and MATLAB to plot most of the figures34,35.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The de-identified raw heart rate, steps and sleep data used in this study can be downloaded from the study data repository (https://storage.googleapis.com/gbsc-gcp-project-ipop_public/COVID-19/COVID-19-Wearables.zip). Processed data, including algorithm outputs and the data used for plotting the figures are provided as Supplementary Data 1.
Code availability
Code for the algorithms used in this manuscript is available at https://github.com/mwgrassgreen/WearableDetection (RHR-Diff and CuSum) and https://github.com/gireeshkbogu/AnomalyDetect (HROS-AD).
References
Sethuraman, N., Jeremiah, S. S. & Ryo, A. Interpreting diagnostic tests for SARS-CoV-2. J. Am. Med. Assoc. https://doi.org/10.1001/jama.2020.8259 (2020).
Marinsek, N. et al. Measuring COVID-19 and influenza in the real world via person-generated health data. Preprint at medRxiv https://doi.org/10.1101/2020.05.28.20115964 (2020).
Dunn, J., Runge, R. & Snyder, M. Wearables and the medical revolution. Per. Med. 15, 429–448 (2018).
Kellogg, R. A., Dunn, J. & Snyder, M. P. Personal omics for precision health. Circ. Res. 122, 1169–1171 (2018).
Li, X. et al. Digital health: tracking physiomes and activity using wearable biosensors reveals useful health-related information. PLoS Biol. 15, e2001402 (2017).
Perez, M. V. et al. Large-scale assessment of a smartwatch to identify atrial fibrillation. N. Engl. J. Med. 381, 1909–1917 (2019).
Radin, J. M., Wineinger, N. E., Topol, E. J. & Steinhubl, S. R. Harnessing wearable device data to improve state-level real-time surveillance of influenza-like illness in the USA: a population-based study. Lancet Digital Health 2, e85–e93 (2020).
Zhu, G. et al. Learning from large-scale wearable device data for predicting epidemics trend of COVID-19. Discrete Dyn. Nat. Soc. 2020, 6152041 (2020).
Seshadri, D. R.et al. Wearable sensors for COVID-19: a call to action to harness our digital infrastructure for remote patient monitoring and virtual assessments. Front. Digital Health https://doi.org/10.3389/fdgth.2020.00008 (2020).
Arons, M. M. et al. Presymptomatic SARS-CoV-2 infections and transmission in a skilled nursing facility. N. Engl. J. Med. 382, 2081–2090 (2020).
He, X. et al. Temporal dynamics in viral shedding and transmissibility of COVID-19. Nat. Med. 26, 672–675 (2020).
Witt, D., Kellogg, R., Snyder, M. & Dunn, J. Windows into human health through wearables data analytics. Curr. Opin. Biomed. Eng. 9, 28–46 (2019).
Arias-Castro, E., Castro, R. M., Tánczos, E. & Wang, M. Distribution-free detection of structured anomalies: permutation and rank-based scans. J. Am. Stat. Assoc. 113, 789–801 (2018).
Rousseeuw, P. J. & Van Driessen, K. A. Fast algorithm for the minimum covariance determinant estimator. Technometrics 41, 212–223 (1999).
Garreta, R. & Moncecchi, G. Learning scikit-learn: Machine Learning in Python (Packt Publishing, 2013).
Backer, J. A., Klinkenberg, D. & Wallinga, J. Incubation period of 2019 novel coronavirus (2019-nCoV) infections among travellers from Wuhan, China, 20–28 January 2020. Eurosurveillance 25, 2000062 (2020).
Lauer, S. A. et al. The incubation period of coronavirus disease 2019 (COVID-19) from publicly reported confirmed cases: estimation and application. Ann. Intern. Med. 172, 577–582 (2020).
Haghayegh, S., Khoshnevis, S., Smolensky, M. H., Diller, K. R. & Castriotta, R. J. Accuracy of wristband Fitbit models in assessing sleep: systematic review and meta-analysis. J. Med. Internet Res. 21, e16273 (2019).
Liang, Z. & Chapa-Martell, M. A. Accuracy of Fitbit wristbands in measuring sleep stage transitions and the effect of user-specific factors. JMIR Mhealth Uhealth 7, e13384 (2019).
Bi, Q. et al. Epidemiology and transmission of COVID-19 in 391 cases and 1286 of their close contacts in Shenzhen, China: a retrospective cohort study. Lancet Infect. Dis. https://doi.org/10.1016/s1473-3099(20)30287-5 (2020).
Arias-Castro, E. & Wang, M. Distribution-free tests for sparse heterogeneous mixtures. Test 26, 71–94 (2017).
Page, E. S. Cumulative sum charts. Technometrics 3, 1–9 (1961).
Levin, B. & Kline, J. The CuSum test of homogeneity with an application in spontaneous abortion epidemiology. Stat. Med. 4, 469–488 (1985).
Xie, J. et al. Association between hypoxemia and mortality in patients with COVID-19. Mayo Clin. Proc. 95, 1138–1147 (2020).
Jouffroy, R., Jost, D. & Prunet, B.Prehospital pulse oximetry: a red flag for early detection of silent hypoxemia in COVID-19 patients. Critical Care 24, 313 (2020).
Lorente-Ros, A. et al. Myocardial injury determination improves risk stratification and predicts mortality in COVID-19 patients. Cardiol. J. https://doi.org/10.5603/CJ.a2020.0089 (2020).
Chen, R. et al. Personal omics profiling reveals dynamic molecular and medical phenotypes. Cell 148, 1293–1307 (2012).
Miller, D. J. et al. Analyzing changes in respiratory rate to predict the risk of COVID-19 infection. Preprint at medRxiv https://doi.org/10.1101/2020.06.18.20131417 (2020).
Mathew, D. et al. Deep immune profiling of COVID-19 patients reveals distinct immunotypes with therapeutic implications. Science 369, eabc8511 (2020).
Siordia, J. A. Epidemiology and clinical features of COVID-19: a review of current literature. J. Clin. Virol. 127, 104357 (2020).
Harris, P. A. et al. Research electronic data capture (REDCap)—a metadata-driven methodology and workflow process for providing translational research informatics support. J. Biomed. Inform. 42, 377–381 (2009).
Zhou, W. et al. Longitudinal multi-omics of host–microbe dynamics in prediabetes. Nature 569, 663–671 (2019).
Boschetti, A. & Massaron, L. Python Data Science Essentials (Packt Publishing, 2015).
Wickham, H. Ggplot2: Elegant Graphics for Data Analysis (Springer, 2016).
Hunter, J. D. Matplotlib: a 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Acknowledgements
This work was supported by NIH grants and gifts from the Flu Lab, as well as departmental funding from the Stanford Genetics department. We thank D. Berrent from Survivor Corps for assistance with recruitment, and A. McDonough and T. Helgren from Fitbit Inc. for help with accessing the Fitbit data. We thank Fitbit Inc. for promoting this study and for the donation of devices. The Stanford Healthcare Innovation Lab gratefully acknowledges the support of A. Duisberg. The Google Cloud Platform costs were covered by Google for Education academic research and COVID-19 grant awards. This work was also supported by Case Western Reserve University through departmental start-up funding and the High-Performance Computing award for COVID-19 Research, both awarded to X.L.
Author information
Authors and Affiliations
Contributions
M.P.S., T.M., X.L. and A.B. conceived of and designed the study. T.M., X.L., A.B. and M.P.S. supervised the project. T.M. performed the project administration. A.C., E.H., E.M.H., O.D.-R., T.M., B.F., S.K., B.R. and M.P.S. performed the Institutional Review Board review, recruited participants and coordinated the study. T.M., A.W.B., A.C., E.H., O.D.-R., B.F., S.K., M.G. and R.K. collected the survey data. A.A.M., A.B. and A.A. collected and processed the wearable device data. A.B., A.A., A.A.M., A.W.B. and T.M. coordinated and submitted the data. A.B. and A.A. developed the software (the MyPHD app). G.K.B., M.W. and X.L. developed the algorithm. T.M., M.W., A.A.M., G.K.B., A.W.B. and X.L. analysed the data. T.M., M.W., A.A.M., G.K.B., A.W.B., X.L. and M.P.S. prepared the manuscript. All co-authors reviewed and edited the manuscript. A.B.G., B.R., A.B., X.L. and M.P.S. sourced the funding.
Corresponding authors
Ethics declarations
Competing interests
M.P.S. is cofounder and a member of the scientific advisory board of Personalis, Qbio, January, SensOmics, Protos, Mirvie and Oralome. He is on the scientific advisory board of Danaher, GenapSys and Jupiter.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary figures and a list of the supplementary tables in Supplementary Data 1.
Supplementary Data 1
Miscellaneous datasets, as described in the Supplementary Information.
Rights and permissions
About this article
Cite this article
Mishra, T., Wang, M., Metwally, A.A. et al. Pre-symptomatic detection of COVID-19 from smartwatch data. Nat Biomed Eng 4, 1208–1220 (2020). https://doi.org/10.1038/s41551-020-00640-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-020-00640-6
This article is cited by
-
Use of artificial intelligence to develop predictive algorithms of cough and PCR-confirmed COVID-19 infections based on inputs from clinical-grade wearable sensors
Scientific Reports (2024)
-
PIF dataset: a comprehensive dataset of physiological and inertial features for recognition of human activities
Multimedia Tools and Applications (2024)
-
Smartwatches in healthcare medicine: assistance and monitoring; a scoping review
BMC Medical Informatics and Decision Making (2023)
-
Descriptive characteristics of continuous oximetry measurement in moderate to severe covid-19 patients
Scientific Reports (2023)
-
Feasibility of using intermittent active monitoring of vital signs by smartphone users to predict SARS-CoV-2 PCR positivity
Scientific Reports (2023)