Abstract
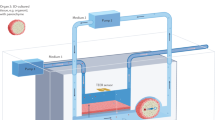
Organ chips can recapitulate organ-level (patho)physiology, yet pharmacokinetic and pharmacodynamic analyses require multi-organ systems linked by vascular perfusion. Here, we describe an ‘interrogator’ that employs liquid-handling robotics, custom software and an integrated mobile microscope for the automated culture, perfusion, medium addition, fluidic linking, sample collection and in situ microscopy imaging of up to ten organ chips inside a standard tissue-culture incubator. The robotic interrogator maintained the viability and organ-specific functions of eight vascularized, two-channel organ chips (intestine, liver, kidney, heart, lung, skin, blood–brain barrier and brain) for 3 weeks in culture when intermittently fluidically coupled via a common blood substitute through their reservoirs of medium and endothelium-lined vascular channels. We used the robotic interrogator and a physiological multicompartmental reduced-order model of the experimental system to quantitatively predict the distribution of an inulin tracer perfused through the multi-organ human-body-on-chips. The automated culture system enables the imaging of cells in the organ chips and the repeated sampling of both the vascular and interstitial compartments without compromising fluidic coupling.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All the data supporting the results in this study are available within the article and its Supplementary Information. Mass spectrometry data of brain-chip metabolites are available as Supplementary Dataset 1. The broad range of raw datasets acquired and analysed (or any subsets of it), which for reuse would require contextual metadata, are available from the corresponding author upon reasonable request.
Code availability
A SolidWorks CAD package of the Interrogator that defines all hardware components and assembly instructions are provided as Supplementary Design Files 1 and 2. The mass spectrometry data are provided as Supplementary Dataset 1. Control software is available at https://gitlab.com/wyss-microengineering/hydra-controller, and video tutorials are available at https://vimeo.com/album/5703210. Organ-chip simulations developed using CoBi Tools are available at http://medicalavatars.cfdrc.com/index.php/cobi-tools.
References
Bhatia, S. N. & Ingber, D. E. Microfluidic organs-on-chips. Nat. Biotechnol. 32, 760–772 (2014).
Lee, S. et al. Microfluidic-based vascularized microphysiological systems. Lab Chip 18, 2686–2709 (2018).
Bhushan, A., Martucci, N. J., Usta, O. B. & Yarmush, M. L. New technologies in drug metabolism and toxicity screening: organ-to-organ interaction. Expert Opin. Drug Metab. Toxicol. 12, 475–477 (2016).
Benigni, R. Predictive toxicology today: the transition from biological knowledge to practicable models. Expert Opin. Drug Metab. Toxicol. 12, 989–992 (2016).
Mak, I. W., Evaniew, N. & Ghert, M. Lost in translation: animal models and clinical trials in cancer treatment. Am. J. Transl. Res. 6, 114–118 (2014).
Ewart, L. et al. Application of microphysiological systems to enhance safety assessment in drug discovery. Annu. Rev. Pharmacol. Toxicol. 58, 65–82 (2018).
Prantil-Baun, R. et al. Physiologically based pharmacokinetic and pharmacodynamic analysis enabled by microfluidically linked organs-on-chips. Annu. Rev. Pharmacol. Toxicol. 58, 37–64 (2018).
Vernetti, L. et al. Functional coupling of human microphysiology systems: intestine, liver, kidney proximal tubule, blood–brain barrier and skeletal muscle. Sci. Rep. 7, 42296 (2017).
Sung, J. H. et al. Using PBPK guided “Body-on-a-Chip” systems to predict mammalian response to drug and chemical exposure. Exp. Biol. Med. 239, 1225–1239 (2014).
Wang, Y. I et al. Self-contained, low-cost Body-on-a-Chip systems for drug development. Exp. Biol. Med. 242, 1701–1713 (2017).
Edington, C. D. et al. Interconnected microphysiological systems for quantitative biology and pharmacology studies. Sci. Rep. 8, 4530 (2018).
Loskill, P., Marcus, S. G., Mathur, A., Reese, W. M. & Healy, K. E. μOrgano: a Lego®-like plug & play system for modular multi-organ-chips. PLoS ONE 10, e0139587 (2015).
Maoz, B. M. et al. A linked organ-on-chip model of the human neurovascular unit reveals the metabolic coupling of endothelial and neuronal cells. Nat. Biotechnol. 36, 865–874 (2018).
Coppeta, J. R. et al. A portable and reconfigurable multi-organ platform for drug development with onboard microfluidic flow control. Lab Chip 17, 134–144 (2016).
Xiao, S. et al. A microfluidic culture model of the human reproductive tract and 28-day menstrual cycle. Nat. Commun. 8, 14584 (2017).
Esch, M. B., Ueno, H., Applegate, D. R. & Shuler, M. L. Modular, pumpless body-on-a-chip platform for the co-culture of GI tract epithelium and 3D primary liver tissue. Lab Chip 16, 2719–2729 (2016).
Oleaga, C et al. Multi-organ toxicity demonstration in a functional human in vitro system composed of four organs. Sci. Rep. 6, 20030 (2016).
Sances, S. et al. Human iPSC-derived endothelial cells and microengineered organ-chip enhance neuronal development. Stem Cell Rep. 10, 1222–1236 (2018).
Maschmeyer, I. et al. A four-organ-chip for interconnected long-term co-culture of human intestine, liver, skin and kidney equivalents. Lab Chip 15, 2688–2699 (2015).
Bauer, S. et al. Functional coupling of human pancreatic islets and liver spheroids on-a-chip: towards a novel human ex vivo type 2 diabetes model. Sci. Rep. 7, 14620 (2017).
Yu, F., Selva Kumar, N. D., Choudhury, D., Foo, L. C. & Ng, S. H. Microfluidic platforms for modeling biological barriers in the circulatory system. Drug Discov. Today 23, 815–829 (2018).
Poisson, J. et al. Liver sinusoidal endothelial cells: physiology and role in liver diseases. J. Hepatol. 66, 212–227 (2017).
Huh, D. et al. Microfabrication of human organs-on-chips. Nat. Protoc. 8, 2135–2157 (2013).
Novak, R et al. Scalable fabrication of stretchable, dual channel, microfluidic organ chips. J. Vis. Exp. 140, e58151 (2018).
Ewart, L. et al. Navigating tissue chips from development to dissemination: a pharmaceutical industry perspective. Exp. Biol. Med. 242, 1579–1585 (2017).
Brown, J. A. et al. Recreating blood–brain barrier physiology and structure on chip: a novel neurovascular microfluidic bioreactor. Biomicrofluidics 9, 054124 (2015).
Griep, L. M. et al. BBB on chip: microfluidic platform to mechanically and biochemically modulate blood–brain barrier function. Biomed. Microdevices 15, 145–150 (2013).
Huh, D. et al. Reconstituting organ-level lung functions on a chip. Science 328, 1662–1668 (2010).
Kim, H. J., Huh, D., Hamilton, G. & Ingber, D. E. Human gut-on-a-chip inhabited by microbial flora that experiences intestinal peristalsis-like motions and flow. Lab Chip 12, 2165–2174 (2012).
Kim, H. J. & Ingber, D. E. Gut-on-a-Chip microenvironment induces human intestinal cells to undergo villus differentiation. Integr. Biol. 5, 1130–1140 (2013).
Jang, K.-J. et al. Human kidney proximal tubule-on-a-chip for drug transport and nephrotoxicity assessment. Integr. Biol. 5, 1119–1129 (2013).
Imura, Y., Asano, Y., Sato, K. & Yoshimura, E. A microfluidic system to evaluate intestinal absorption. Anal. Sci. 25, 1403–1407 (2009).
Leclerc, E., Sakai, Y. & Fujii, T. Microfluidic PDMS (polydimethylsiloxane) bioreactor for large-scale culture of hepatocytes. Biotechnol. Prog. 20, 750–755 (2004).
Kim, H. J., Li, H., Collins, J. J. & Ingber, D. E. Contributions of microbiome and mechanical deformation to intestinal bacterial overgrowth and inflammation in a human gut-on-a-chip. Proc. Natl Acad. Sci. USA 113, E7–E15 (2016).
Huh, D. et al. A human disease model of drug toxicity-induced pulmonary edema in a lung-on-a-chip microdevice. Sci. Transl Med. 4, 159ra147 (2012).
Herland, A. et al. Quantitative prediction of human drug pharmacokinetic responses enabled by fluidically coupled vascularized organ chips. Nat. Biomed. Eng. https://doi.org/10.1038/s41551-019-0498-9 (2020).
Toto, R. D. Conventional measurement of renal function utilizing serum creatinine, creatinine clearance, inulin and para-aminohippuric acid clearance. Curr. Opin. Nephrol. Hypertens. 4, 505–509 (1995); discussion 4, 503–504 (1995).
Rahn, K. H., Heidenreich, S. & Brückner, D. How to assess glomerular function and damage in humans. J. Hypertens. 17, 309–317 (1999).
Rose, G. A. Measurement of glomerular filtration rate by inulin clearance without urine collection. BMJ 2, 91–93 (1969).
Musah, S. et al. Mature induced-pluripotent-stem-cell-derived human podocytes reconstitute kidney glomerular-capillary-wall function on a chip. Nat. Biomed. Eng. 1, 0069 (2017).
Kujala, V. J., Pasqualini, F. S., Goss, J. A., Nawroth, J. C. & Parker, K. K. Laminar ventricular myocardium on a microelectrode array-based chip. J. Mater. Chem. B 4, 3534–3543 (2016).
Géraud, C. et al. Unique cell type-specific junctional complexes in vascular endothelium of human and rat liver sinusoids. PLoS ONE 7, e34206 (2012).
Wikswo, J. P. et al. Scaling and systems biology for integrating multiple organs-on-a-chip. Lab Chip 13, 3496–3511 (2013).
Low, L. A. & Tagle, D. A. Organs-on-chips: progress, challenges, and future directions. Exp. Biol. Med. 242, 1573–1578 (2017).
Benam, K. H. et al. Small airway-on-a-chip enables analysis of human lung inflammation and drug responses in vitro. Nat. Methods 13, 151–157 (2016).
Benam, K. H. et al. Matched-comparative modeling of normal and diseased human airway responses using a microengineered breathing lung chip. Cell Syst. 3, 456–466.e4 (2016).
Bein, A. et al. Microfluidic organ-on-a-chip models of human intestine. Cell. Mol. Gastroenterol. Hepatol. 5, 659–668 (2018).
Jalili-Firoozinezhad, S. et al. Modeling radiation injury-induced cell death and countermeasure drug responses in a human Gut-on-a-Chip. Cell Death Dis. 9, 223 (2018).
Agarwal, A., Goss, J. A., Cho, A., McCain, M. L. & Parker, K. K. Microfluidic heart on a chip for higher throughput pharmacological studies. Lab Chip 13, 3599–3608 (2013).
Hubatsch, I., Ragnarsson, E. G. E. & Artursson, P. Determination of drug permeability and prediction of drug absorption in Caco-2 monolayers. Nat. Protoc. 2, 2111–2119 (2007).
Lehr, C.-M. Cell Culture Models of Biological Barriers: In vitro Test Systems for Drug Absorption and Delivery (CRC Press, 2003).
Roberts, M. S. & Rowland, M. A dispersion model of hepatic elimination. 1. Formulation of the model and bolus considerations. J. Pharmacokinet. Biopharm. 14, 227–260 (1986).
Dong, J. & Park, M. S. Discussions on the hepatic well-stirred model: re-derivation from the dispersion model and re-analysis of the lidocaine data. Eur. J. Pharm. Sci. 124, 46–60 (2018).
Somayaji, M. R., Das, D. & Przekwas, A. Computational approaches for modeling and analysis of human-on-chip systems for drug testing and characterization. Drug Discov. Today 21, 1859–1862 (2016).
Robinson, D. E., Balter, N. J. & Schwartz, S. L. A physiologically based pharmacokinetic model for nicotine and cotinine in man. J. Pharmacokinet. Biopharm. 20, 591–609 (1992).
Miller, P. G. & Shuler, M. L. Design and demonstration of a pumpless 14 compartment microphysiological system. Biotechnol. Bioeng. 113, 2213–2227 (2016).
Jalili-Firoozinezhad, S et al. A complex human gut microbiome cultured in an anaerobic intestine-on-a-chip. Nat. Biomed. Eng. 3, 520–531 (2019).
Phan, D. T. T. et al. A vascularized and perfused organ-on-a-chip platform for large-scale drug screening applications. Lab Chip 17, 511–520 (2017).
Kasendra, M. et al. Development of a primary human Small Intestine-on-a-Chip using biopsy-derived organoids. Sci. Rep. 8, 2871 (2018).
Benam, K. H. et al. Human lung small airway-on-a-chip protocol. Methods Mol. Biol. 1612, 345–365 (2017).
Jang, K.-J. et al. Reproducing human and cross-species drug toxicities using a Liver-Chip. Sci. Transl. Med. 11, eaax5516 (2019).
Tran, T. T. et al. Exact kinetic analysis of passive transport across a polarized confluent MDCK cell monolayer modeled as a single barrier. J. Pharm. Sci. 93, 2108–2123 (2004).
Maddocks, O. D. K. et al. Serine starvation induces stress and p53-dependent metabolic remodelling in cancer cells. Nature 493, 542–546 (2013).
Przekwas, A., Friend, T., Teixeira, R., Chen, Z. J. & Wilkerson, P. Spatial Modeling Tools for Cell Biology (CFD Research Corporation, 2006).
Wang, G. et al. Modeling the mitochondrial cardiomyopathy of Barth syndrome with iPSC and heart-on-chip technologies. Nat. Med. 20, 616–623 (2014).
Acknowledgements
This research was sponsored by the Wyss Institute for Biologically Inspired Engineering at Harvard University and the Defense Advanced Research Projects Agency under Cooperative Agreement number W911NF-12-2-0036. The views and conclusions contained in this document are those of the authors and should not be interpreted as representing the official policies, either expressed or implied, of the Defense Advanced Research Projects Agency or the US government. This work was performed in part at the Center for Nanoscale Systems (CNS), a member of the National Nanotechnology Coordinated Infrastructure Network (NNCI), which is supported by the National Science Foundation under NSF award number 1541959. The CNS is part of Harvard University, the Harvard Materials Research Science and Engineering Center (DMR-1420570). The authors thank J. Caramanica and P. Machado for their machining expertise, M. Rosnach for his artwork, B. Fountaine and S. Kroll for their help with photography, M. Rousseau for help with videography, C. Vidoudez for mass spectrometry analysis, and J. Wikswo for helpful input at the start of this project.
Author information
Authors and Affiliations
Contributions
R.N., A.H., B.M.M. and R.P.-B. led the data analyses for the generation of figures and, with D.E.I., prepared the manuscript. A.H., B.M.M., A. Chalkiadaki, D.B.C, M.C., T.H., M.B., S.D., E.A.F., S.S.F.J., T.G., S.J.-F., V.K., L.L., R.M., Y.M., J.A.N., B.O., T.-E.P., H.S., B.S., G.J.T., Z.T., T.H.-I., K.-J.J., A.S.-P. and M.Y. planned and performed biological experiments, with G.A.H., O.L., A.B., R.N., R.P.-B., K.K.P. and D.E.I. supervising the work. A.Cho., E.C., Y.C., J.F., R.F., C.F.N., R.N., G.A.H., J.A.G., N.W., K.K.P. and D.E.I. were responsible for the development and fabrication of the chip. J.S., G.T.II, C.H., J.F.-A., J.A.G. and D.L. conceptualized the robotics-based organ chip linking and developed early versions of the Interrogator instrument and software with an integrated robotic sampler. M.I., Y.C., S.M., A.D., T.D., T.F., O.H. B.A.N., and R.N were responsible for the software and hardware engineering and were involved in the development of the final instrument and sensors used in this study. D.D., M.R.S. and A.P. were responsible for MCRO model development and data analyses, working closely with A.H., B.M.M., R.P.-B. and R.N. Finally, R.N., R.P.-B., O.L., G.A.H., D.L., K.K.P. and D.E.I. were responsible for overseeing and orchestrating the entire effort.
Corresponding author
Ethics declarations
Competing interests
D.E.I. is a founder and holds equity in Emulate, Inc., and chairs its scientific advisory board. K.K.P. is a consultant and a member of the scientific advisory board of Emulate, Inc. S.S.F.J., J.F.-A., G.A.H., C.H., K.-J.J., V.K., L.L., D.L., J.N., J.S., G.T.II and N.W. are employees of and hold equity in Emulate, Inc. A.B., Y.C., M.C., S.D., J.F.-A., T.F., E.A.F., J.A.G., G.A.H., T.H.-I., O.H., A.H., C.H., D.E.I., M.I., K.-J.J., V.K., L.L., D.L., O.L., B.M.M., Y.M., J.N., R.N., T.-E.P., K.K.P., J.S., A.S.-P., G.T.II and N.W. are inventors on intellectual property licensed to Emulate, Inc.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary methods, figures, tables and video captions.
Supplementary Design Files 1
SolidWorks CAD package for the interrogator device.
Supplementary Design Files 2
SolidWorks CAD package for the fluorescence module.
Supplementary Dataset 1
Mass spectrometry data of brain-chip metabolites.
Supplementary Video 1
Overview of the Interrogator-device components and organ-chip linking.
Supplementary Video 2
Gut-chip stretching, visualized via the microscope module.
Supplementary Video 3
Heart chip beating after being linked for 3 weeks on the Interrogator device.
Supplementary Video 4
High‐fidelity diffusion model of the heart chip.
Supplementary Video 5
CoBi Q‐3D model of gut–liver chip linking.
Rights and permissions
About this article
Cite this article
Novak, R., Ingram, M., Marquez, S. et al. Robotic fluidic coupling and interrogation of multiple vascularized organ chips. Nat Biomed Eng 4, 407–420 (2020). https://doi.org/10.1038/s41551-019-0497-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-019-0497-x
This article is cited by
-
Microfluidic high-throughput 3D cell culture
Nature Reviews Bioengineering (2024)
-
Engineering native biological complexity from the inside–out and outside–in
Nature Chemical Engineering (2024)
-
Bioengineered skin organoids: from development to applications
Military Medical Research (2023)
-
A human lung alveolus-on-a-chip model of acute radiation-induced lung injury
Nature Communications (2023)
-
Human disease models in drug development
Nature Reviews Bioengineering (2023)